Detailed results of SSR10_NMR_em_bcr3 by PSVS
Output from PDBStat
Constraints analysis
table of NOE constraints
# -------------- SUMMARY OF RESTRAINTS ---------------
# TOTAL NUMBER OF NOE RESTRAINTS : 1661
# INTRA-RESIDUE RESTRAINTS (I=J) : 542
# SEQUENTIAL RESTRAINTS (I-J)=1 : 444
# BACKBONE-BACKBONE : 131
# BACKBONE-SIDE CHAIN : 18
# SIDE CHAIN-SIDE CHAIN : 295
# MEDIUM RANGE RESTRAINTS 1<(I-J)<5 : 362
# BACKBONE-BACKBONE : 144
# BACKBONE-SIDE CHAIN : 129
# SIDE CHAIN-SIDE CHAIN : 89
# LONG RANGE RESTRAINTS (I-J)>=5 : 313
# TOTAL HYDROGEN BOND RESTRAINTS : 0
# LONG RANGE H-BOND RESTR. (I-J)>=5 : 0
# DISULFIDE RESTRAINTS : 0
# INTRA-CHAIN RESTRAINTS : 1661
# INTER-CHAIN RESTRAINTS : 0
# AMBIGUOUS RESTRAINTS : 0
# -----------------------------------------------------
# -----------------------------------------------------
# -----------------------------------------------------
# RES # INTRA INTER seq med lng InterChain
MET 1 0 0.0 0.0 0.0 0.0 0.0
SER 2 0 0.0 0.0 0.0 0.0 0.0
ILE 3 4 4.0 3.5 0.5 0.0 0.0
SER 4 0 6.5 5.5 0.0 1.0 0.0
THR 5 1 10.5 4.0 4.5 2.0 0.0
SER 6 3 7.0 3.5 3.5 0.0 0.0
ALA 7 0 12.0 2.0 3.0 7.0 0.0
GLU 8 6 10.0 2.0 4.0 4.0 0.0
VAL 9 2 10.5 4.5 5.0 1.0 0.0
TYR 10 2 22.0 4.5 5.5 12.0 0.0
TYR 11 2 16.5 4.5 3.0 9.0 0.0
GLU 12 3 9.5 4.5 4.5 0.5 0.0
GLU 13 9 8.5 5.0 3.5 0.0 0.0
ALA 14 0 14.5 4.0 4.0 6.5 0.0
GLU 15 7 6.0 1.5 4.5 0.0 0.0
GLU 16 8 7.5 4.0 3.5 0.0 0.0
PHE 17 2 17.0 5.0 5.0 7.0 0.0
LEU 18 7 14.0 5.5 5.0 3.5 0.0
SER 19 0 5.5 4.0 1.5 0.0 0.0
LYS 20 5 7.5 2.5 5.0 0.0 0.0
GLY 21 0 4.0 2.5 1.5 0.0 0.0
ASP 22 3 8.0 2.5 3.5 2.0 0.0
LEU 23 7 15.5 5.0 2.5 8.0 0.0
VAL 24 4 11.0 7.0 3.0 1.0 0.0
GLN 25 13 8.0 6.0 1.0 1.0 0.0
ALA 26 0 15.0 4.0 4.0 7.0 0.0
CYS 27 3 8.0 4.0 3.5 0.5 0.0
GLU 28 3 4.5 2.5 2.0 0.0 0.0
LYS 29 0 8.5 2.5 1.5 4.5 0.0
TYR 30 0 8.5 2.5 3.5 2.5 0.0
TYR 31 3 11.5 2.0 5.5 4.0 0.0
LYS 32 10 8.0 3.5 1.5 3.0 0.0
ALA 33 0 17.5 3.0 4.0 10.5 0.0
ALA 34 0 11.5 1.5 3.0 7.0 0.0
GLU 35 9 13.0 2.5 4.5 6.0 0.0
GLU 36 6 13.5 4.5 2.0 7.0 0.0
ALA 37 0 14.0 3.0 3.5 7.5 0.0
ILE 38 9 13.0 2.0 3.0 8.0 0.0
LYS 39 0 5.5 2.5 2.5 0.5 0.0
LEU 40 10 9.5 3.0 4.0 2.5 0.0
LEU 41 8 17.5 4.5 8.0 5.0 0.0
VAL 42 2 18.0 5.0 2.0 11.0 0.0
ILE 43 8 9.0 4.5 2.5 2.0 0.0
GLU 44 9 4.5 4.5 0.0 0.0 0.0
ASN 45 3 11.0 3.0 6.5 1.5 0.0
ASN 46 6 2.0 1.5 0.5 0.0 0.0
LEU 47 7 10.0 3.0 1.0 6.0 0.0
LYS 48 9 9.0 4.0 5.0 0.0 0.0
GLU 49 10 9.5 4.5 3.5 1.5 0.0
ILE 50 7 18.0 5.5 3.0 9.5 0.0
THR 51 3 7.0 4.0 2.5 0.5 0.0
ASN 52 0 8.5 1.5 7.0 0.0 0.0
ASN 53 4 5.0 1.5 3.0 0.5 0.0
VAL 54 2 9.5 3.5 3.0 3.0 0.0
LYS 55 7 6.5 6.0 0.5 0.0 0.0
ASN 56 3 7.0 6.0 1.0 0.0 0.0
LYS 57 10 5.0 5.0 0.0 0.0 0.0
GLY 58 0 4.0 4.0 0.0 0.0 0.0
ARG 59 4 4.5 4.5 0.0 0.0 0.0
TRP 60 4 18.0 5.0 0.0 13.0 0.0
LYS 61 6 5.5 4.0 1.5 0.0 0.0
SER 62 5 3.0 2.5 0.5 0.0 0.0
GLU 63 3 5.5 3.5 2.0 0.0 0.0
ASN 64 6 10.0 5.5 3.5 1.0 0.0
LEU 65 7 14.0 4.0 2.0 8.0 0.0
PHE 66 4 4.5 3.5 1.0 0.0 0.0
LYS 67 9 11.0 3.5 4.5 3.0 0.0
ALA 68 0 8.0 2.0 2.5 3.5 0.0
SER 69 3 8.5 2.0 2.0 4.5 0.0
LYS 70 9 6.0 4.5 1.5 0.0 0.0
LEU 71 8 9.0 5.5 2.0 1.5 0.0
LEU 72 6 10.0 3.0 2.0 5.0 0.0
ARG 73 10 5.5 3.0 2.5 0.0 0.0
SER 74 0 3.5 3.0 0.5 0.0 0.0
ASN 75 6 4.0 2.0 2.0 0.0 0.0
ASN 76 0 6.0 3.0 3.0 0.0 0.0
THR 77 3 5.5 3.0 2.5 0.0 0.0
GLU 78 9 9.5 3.0 6.5 0.0 0.0
ILE 79 4 14.0 4.5 6.0 3.5 0.0
PRO 80 0 7.5 4.5 1.5 1.5 0.0
ILE 81 10 9.5 4.0 5.5 0.0 0.0
LEU 82 8 23.0 6.0 7.0 10.0 0.0
TRP 83 2 19.0 4.0 4.0 11.0 0.0
LYS 84 8 6.5 2.0 4.5 0.0 0.0
SER 85 3 6.5 3.5 2.0 1.0 0.0
ALA 86 0 11.0 2.5 3.0 5.5 0.0
TRP 87 4 6.5 1.5 4.0 1.0 0.0
THR 88 1 7.5 2.5 4.5 0.5 0.0
LEU 89 4 9.0 3.0 2.0 4.0 0.0
HIS 90 4 5.0 4.0 1.0 0.0 0.0
VAL 91 2 8.5 4.5 4.0 0.0 0.0
GLU 92 2 3.5 2.5 1.0 0.0 0.0
GLY 93 0 1.0 1.0 0.0 0.0 0.0
PHE 94 2 3.0 3.0 0.0 0.0 0.0
HIS 95 3 3.0 3.0 0.0 0.0 0.0
GLU 96 0 1.0 1.0 0.0 0.0 0.0
LEU 97 11 4.5 4.0 0.5 0.0 0.0
SER 98 2 6.0 5.0 0.0 1.0 0.0
LEU 99 7 6.0 5.0 0.5 0.5 0.0
ASN 100 2 8.0 5.5 2.0 0.5 0.0
GLU 101 4 15.5 4.5 4.5 6.5 0.0
LYS 102 11 7.0 4.5 2.5 0.0 0.0
GLU 103 3 8.5 5.5 3.0 0.0 0.0
VAL 104 3 11.5 5.5 5.0 1.0 0.0
LYS 105 10 8.5 3.5 5.0 0.0 0.0
LYS 106 5 5.0 2.5 2.5 0.0 0.0
LEU 107 6 9.5 2.5 4.5 2.5 0.0
LYS 108 5 7.5 3.0 4.5 0.0 0.0
GLU 109 3 7.5 2.5 5.0 0.0 0.0
ASP 110 2 9.0 2.5 4.0 2.5 0.0
VAL 111 2 19.0 3.5 5.5 10.0 0.0
ARG 112 4 11.5 3.0 6.5 2.0 0.0
LYS 113 9 9.0 2.0 2.5 4.5 0.0
LEU 114 9 17.5 4.5 4.5 8.5 0.0
VAL 115 3 19.0 4.5 5.5 9.0 0.0
ILE 116 10 11.5 5.0 6.5 0.0 0.0
PHE 117 2 13.5 6.0 4.5 3.0 0.0
ALA 118 0 12.0 2.5 4.5 5.0 0.0
VAL 119 4 8.0 3.0 3.5 1.5 0.0
ASN 120 3 9.0 4.5 4.5 0.0 0.0
SER 121 2 7.5 5.0 2.5 0.0 0.0
LEU 122 4 7.5 5.0 1.5 1.0 0.0
GLU 123 5 3.5 3.0 0.5 0.0 0.0
HIS 124 5 1.0 1.0 0.0 0.0 0.0
HIS 125 5 0.5 0.5 0.0 0.0 0.0
HIS 126 3 0.5 0.5 0.0 0.0 0.0
HIS 127 0 0.0 0.0 0.0 0.0 0.0
HIS 128 0 0.0 0.0 0.0 0.0 0.0
HIS 129 0 0.0 0.0 0.0 0.0 0.0
# TOTAL 542 1119.0 444.0 362.0 313.0 0.0
# TOTAL NUMBER OF RESTRAINTS (CHECKING): 1661.0
List of conformationally-resticting NOE constraints
assign ((resid 4 and name HN )) ( (resid 5 and name HN )) 2.66 0.86 0.86
assign ((resid 6 and name HB2 )) ( (resid 7 and name HN )) 3.43 1.63 1.63
assign ((resid 6 and name HB1 )) ( (resid 7 and name HN )) 3.43 1.63 1.63
assign ((resid 7 and name HN )) ( (resid 8 and name HN )) 2.82 1.02 1.02
assign ((resid 9 and name HB )) ( (resid 10 and name HN )) 2.42 0.63 0.63
assign ((resid 10 and name HN )) ( (resid 11 and name HN )) 2.82 1.02 1.02
assign ((resid 10 and name HB1 )) ( (resid 11 and name HN )) 3.02 1.22 1.22
assign ((resid 10 and name HB2 )) ( (resid 11 and name HN )) 2.94 1.15 1.15
assign ((resid 11 and name HN )) ( (resid 12 and name HN )) 2.60 0.81 0.81
assign ((resid 11 and name HB2 )) ( (resid 12 and name HN )) 3.00 1.20 1.20
assign ((resid 11 and name HB1 )) ( (resid 12 and name HN )) 2.71 0.91 0.91
assign ((resid 12 and name HB2 )) ( (resid 13 and name HN )) 2.94 1.15 1.15
assign ((resid 12 and name HB1 )) ( (resid 13 and name HN )) 2.94 1.15 1.15
assign ((resid 13 and name HN )) ( (resid 14 and name HN )) 2.75 0.95 0.95
assign ((resid 13 and name HB2 )) ( (resid 14 and name HN )) 2.78 0.99 0.99
assign ((resid 13 and name HB1 )) ( (resid 14 and name HN )) 2.78 0.99 0.99
assign ((resid 14 and name HN )) ( (resid 15 and name HN )) 2.62 0.82 0.82
assign ((resid 16 and name HA )) ( (resid 17 and name HN )) 2.62 0.82 0.82
assign ((resid 17 and name HB2 )) ( (resid 18 and name HN )) 2.89 1.09 1.09
assign ((resid 17 and name HB1 )) ( (resid 18 and name HN )) 3.05 1.26 1.26
assign ((resid 18 and name HN )) ( (resid 19 and name HN )) 2.62 0.82 0.82
assign ((resid 18 and name HB2 )) ( (resid 19 and name HN )) 2.91 1.11 1.11
assign ((resid 18 and name HB1 )) ( (resid 19 and name HN )) 2.91 1.11 1.11
assign ((resid 19 and name HN )) ( (resid 20 and name HN )) 2.57 0.77 0.77
assign ((resid 20 and name HB2 )) ( (resid 21 and name HN )) 3.52 1.72 1.72
assign ((resid 20 and name HB1 )) ( (resid 21 and name HN )) 3.52 1.72 1.72
assign ((resid 22 and name HB2 )) ( (resid 23 and name HN )) 3.90 2.10 2.10
assign ((resid 22 and name HB1 )) ( (resid 23 and name HN )) 3.90 2.10 2.10
assign ((resid 23 and name HB2 )) ( (resid 24 and name HN )) 2.82 1.02 1.02
assign ((resid 24 and name HN )) ( (resid 25 and name HN )) 2.55 0.75 0.75
assign ((resid 25 and name HN )) ( (resid 26 and name HN )) 2.53 0.73 0.73
assign ((resid 25 and name HB1 )) ( (resid 26 and name HN )) 3.03 1.24 1.24
assign ((resid 25 and name HB2 )) ( (resid 26 and name HN )) 2.76 0.97 0.97
assign ((resid 27 and name HN )) ( (resid 28 and name HN )) 2.75 0.95 0.95
assign ((resid 27 and name HB2 )) ( (resid 28 and name HN )) 3.07 1.27 1.27
assign ((resid 27 and name HB1 )) ( (resid 28 and name HN )) 3.07 1.27 1.27
assign ((resid 28 and name HN )) ( (resid 29 and name HN )) 2.66 0.86 0.86
assign ((resid 29 and name HN )) ( (resid 30 and name HN )) 2.66 0.86 0.86
assign ((resid 29 and name HB2 )) ( (resid 30 and name HN )) 3.20 1.40 1.40
assign ((resid 29 and name HB1 )) ( (resid 30 and name HN )) 3.20 1.40 1.40
assign ((resid 32 and name HN )) ( (resid 33 and name HN )) 2.66 0.86 0.86
assign ((resid 32 and name HB2 )) ( (resid 33 and name HN )) 2.96 1.17 1.17
assign ((resid 35 and name HN )) ( (resid 36 and name HN )) 2.66 0.86 0.86
assign ((resid 35 and name HB1 )) ( (resid 36 and name HN )) 2.85 1.06 1.06
assign ((resid 36 and name HN )) ( (resid 36 and name HB1 )) 2.84 1.04 1.04
assign ((resid 36 and name HB2 )) ( (resid 37 and name HN )) 3.27 1.47 1.47
assign ((resid 36 and name HB1 )) ( (resid 37 and name HN )) 3.27 1.47 1.47
assign ((resid 37 and name HN )) ( (resid 38 and name HN )) 2.64 0.84 0.84
assign ((resid 38 and name HN )) ( (resid 39 and name HN )) 2.62 0.82 0.82
assign ((resid 41 and name HB2 )) ( (resid 42 and name HN )) 3.07 1.27 1.27
assign ((resid 41 and name HB1 )) ( (resid 42 and name HN )) 3.07 1.27 1.27
assign ((resid 42 and name HN )) ( (resid 43 and name HN )) 2.67 0.88 0.88
assign ((resid 43 and name HN )) ( (resid 44 and name HN )) 2.66 0.86 0.86
assign ((resid 44 and name HN )) ( (resid 45 and name HN )) 2.75 0.95 0.95
assign ((resid 44 and name HB2 )) ( (resid 45 and name HN )) 3.05 1.26 1.26
assign ((resid 44 and name HB1 )) ( (resid 45 and name HN )) 3.05 1.26 1.26
assign ((resid 46 and name HN )) ( (resid 47 and name HN )) 2.71 0.91 0.91
assign ((resid 47 and name HA )) ( (resid 48 and name HN )) 2.37 0.57 0.57
assign ((resid 47 and name HB2 )) ( (resid 48 and name HN )) 3.09 1.29 1.29
assign ((resid 49 and name HB2 )) ( (resid 50 and name HN )) 3.29 1.49 1.49
assign ((resid 49 and name HB1 )) ( (resid 50 and name HN )) 3.29 1.49 1.49
assign ((resid 50 and name HN )) ( (resid 51 and name HN )) 2.48 0.68 0.68
assign ((resid 51 and name HN )) ( (resid 52 and name HN )) 2.64 0.84 0.84
assign ((resid 51 and name HB )) ( (resid 52 and name HN )) 2.94 1.15 1.15
assign ((resid 53 and name HB2 )) ( (resid 54 and name HN )) 2.94 1.15 1.15
assign ((resid 53 and name HB1 )) ( (resid 54 and name HN )) 2.94 1.15 1.15
assign ((resid 54 and name HN )) ( (resid 55 and name HN )) 2.66 0.86 0.86
assign ((resid 55 and name HB2 )) ( (resid 56 and name HN )) 2.84 1.04 1.04
assign ((resid 55 and name HB1 )) ( (resid 56 and name HN )) 2.84 1.04 1.04
assign ((resid 56 and name HN )) ( (resid 57 and name HN )) 2.62 0.82 0.82
assign ((resid 57 and name HB2 )) ( (resid 58 and name HN )) 3.16 1.36 1.36
assign ((resid 57 and name HB1 )) ( (resid 58 and name HN )) 3.16 1.36 1.36
assign ((resid 60 and name HN )) ( (resid 61 and name HN )) 3.18 1.38 1.38
assign ((resid 60 and name HB2 )) ( (resid 61 and name HN )) 3.00 1.20 1.20
assign ((resid 60 and name HB1 )) ( (resid 61 and name HN )) 3.12 1.33 1.33
assign ((resid 61 and name HA )) ( (resid 62 and name HN )) 2.60 0.81 0.81
assign ((resid 61 and name HB2 )) ( (resid 62 and name HN )) 3.29 1.49 1.49
assign ((resid 61 and name HB1 )) ( (resid 62 and name HN )) 3.29 1.49 1.49
assign ((resid 63 and name HN )) ( (resid 64 and name HN )) 2.82 1.02 1.02
assign ((resid 64 and name HB2 )) ( (resid 65 and name HN )) 3.14 1.35 1.35
assign ((resid 64 and name HB1 )) ( (resid 65 and name HN )) 3.14 1.35 1.35
assign ((resid 65 and name HN )) ( (resid 66 and name HN )) 2.69 0.90 0.90
assign ((resid 65 and name HB2 )) ( (resid 66 and name HN )) 2.89 1.09 1.09
assign ((resid 65 and name HB1 )) ( (resid 66 and name HN )) 3.05 1.26 1.26
assign ((resid 66 and name HN )) ( (resid 67 and name HN )) 2.69 0.90 0.90
assign ((resid 66 and name HB2 )) ( (resid 67 and name HN )) 3.03 1.24 1.24
assign ((resid 66 and name HB1 )) ( (resid 67 and name HN )) 3.03 1.24 1.24
assign ((resid 68 and name HN )) ( (resid 69 and name HN )) 2.66 0.86 0.86
assign ((resid 71 and name HB2 )) ( (resid 72 and name HN )) 2.96 1.17 1.17
assign ((resid 72 and name HN )) ( (resid 73 and name HN )) 2.69 0.90 0.90
assign ((resid 73 and name HN )) ( (resid 74 and name HN )) 2.62 0.82 0.82
assign ((resid 73 and name HB2 )) ( (resid 74 and name HN )) 3.12 1.33 1.33
assign ((resid 75 and name HN )) ( (resid 76 and name HN )) 2.39 0.59 0.59
assign ((resid 77 and name HN )) ( (resid 78 and name HN )) 2.67 0.88 0.88
assign ((resid 78 and name HN )) ( (resid 79 and name HN )) 2.58 0.79 0.79
assign ((resid 78 and name HB2 )) ( (resid 79 and name HN )) 2.80 1.00 1.00
assign ((resid 78 and name HB1 )) ( (resid 79 and name HN )) 2.94 1.15 1.15
assign ((resid 82 and name HB2 )) ( (resid 83 and name HN )) 3.09 1.29 1.29
assign ((resid 82 and name HB1 )) ( (resid 83 and name HN )) 3.09 1.29 1.29
assign ((resid 83 and name HN )) ( (resid 84 and name HN )) 2.64 0.84 0.84
assign ((resid 85 and name HB2 )) ( (resid 86 and name HN )) 2.98 1.18 1.18
assign ((resid 85 and name HB1 )) ( (resid 86 and name HN )) 2.98 1.18 1.18
assign ((resid 87 and name HN )) ( (resid 88 and name HN )) 2.75 0.95 0.95
assign ((resid 87 and name HB2 )) ( (resid 88 and name HN )) 3.25 1.45 1.45
assign ((resid 88 and name HB )) ( (resid 89 and name HN )) 2.78 0.99 0.99
assign ((resid 89 and name HN )) ( (resid 90 and name HN )) 2.73 0.93 0.93
assign ((resid 89 and name HB2 )) ( (resid 90 and name HN )) 2.91 1.11 1.11
assign ((resid 89 and name HB1 )) ( (resid 90 and name HN )) 3.21 1.42 1.42
assign ((resid 90 and name HB2 )) ( (resid 91 and name HN )) 3.20 1.40 1.40
assign ((resid 90 and name HB1 )) ( (resid 91 and name HN )) 3.20 1.40 1.40
assign ((resid 92 and name HN )) ( (resid 93 and name HN )) 2.75 0.95 0.95
assign ((resid 94 and name HN )) ( (resid 95 and name HN )) 2.64 0.84 0.84
assign ((resid 97 and name HB2 )) ( (resid 98 and name HN )) 3.34 1.54 1.54
assign ((resid 97 and name HB1 )) ( (resid 98 and name HN )) 3.34 1.54 1.54
assign ((resid 98 and name HB2 )) ( (resid 99 and name HN )) 3.25 1.45 1.45
assign ((resid 98 and name HB1 )) ( (resid 99 and name HN )) 3.25 1.45 1.45
assign ((resid 99 and name HN )) ( (resid 100 and name HN )) 3.57 1.78 1.78
assign ((resid 99 and name HB2 )) ( (resid 100 and name HN )) 3.34 1.54 1.54
assign ((resid 99 and name HB1 )) ( (resid 100 and name HN )) 3.09 1.29 1.29
assign ((resid 100 and name HA )) ( (resid 101 and name HN )) 2.53 0.73 0.73
assign ((resid 101 and name HN )) ( (resid 102 and name HN )) 2.69 0.90 0.90
assign ((resid 102 and name HN )) ( (resid 103 and name HN )) 2.51 0.72 0.72
assign ((resid 104 and name HN )) ( (resid 105 and name HN )) 2.62 0.82 0.82
assign ((resid 104 and name HB )) ( (resid 105 and name HN )) 2.60 0.81 0.81
assign ((resid 105 and name HN )) ( (resid 106 and name HN )) 2.58 0.79 0.79
assign ((resid 105 and name HB* )) ( (resid 106 and name HN )) 3.10 1.30 1.30
assign ((resid 106 and name HN )) ( (resid 107 and name HN )) 2.62 0.82 0.82
assign ((resid 110 and name HB2 )) ( (resid 111 and name HN )) 3.11 1.31 1.31
assign ((resid 110 and name HB1 )) ( (resid 111 and name HN )) 3.41 1.62 1.62
assign ((resid 111 and name HN )) ( (resid 112 and name HN )) 2.66 0.86 0.86
assign ((resid 111 and name HB )) ( (resid 112 and name HN )) 2.71 0.91 0.91
assign ((resid 114 and name HN )) ( (resid 115 and name HN )) 2.55 0.75 0.75
assign ((resid 113 and name HB2 )) ( (resid 114 and name HN )) 2.57 0.77 0.77
assign ((resid 113 and name HB1 )) ( (resid 114 and name HN )) 2.85 1.06 1.06
assign ((resid 114 and name HB1 )) ( (resid 115 and name HN )) 2.89 1.09 1.09
assign ((resid 115 and name HB )) ( (resid 116 and name HN )) 2.60 0.81 0.81
assign ((resid 116 and name HN )) ( (resid 117 and name HN )) 2.53 0.73 0.73
assign ((resid 116 and name HB )) ( (resid 117 and name HN )) 2.37 0.57 0.57
assign ((resid 119 and name HB )) ( (resid 120 and name HN )) 2.62 0.82 0.82
assign ((resid 120 and name HN )) ( (resid 121 and name HN )) 2.57 0.77 0.77
assign ((resid 120 and name HB2 )) ( (resid 121 and name HN )) 2.93 1.13 1.13
assign ((resid 120 and name HB1 )) ( (resid 121 and name HN )) 2.93 1.13 1.13
assign ((resid 121 and name HB2 )) ( (resid 122 and name HN )) 3.11 1.31 1.31
assign ((resid 121 and name HB1 )) ( (resid 122 and name HN )) 3.11 1.31 1.31
assign ((resid 123 and name HN )) ( (resid 124 and name HN )) 2.58 0.79 0.79
assign ((resid 123 and name HA )) ( (resid 124 and name HN )) 2.55 0.75 0.75
assign ((resid 125 and name HN )) ( (resid 126 and name HN )) 2.75 0.95 0.95
assign ((resid 10 and name HN )) ( (resid 10 and name HB1 )) 2.82 1.02 1.02
assign ((resid 10 and name HN )) ( (resid 10 and name HB2 )) 2.66 0.86 0.86
assign ((resid 11 and name HN )) ( (resid 11 and name HB2 )) 2.58 0.79 0.79
assign ((resid 11 and name HN )) ( (resid 11 and name HB1 )) 2.66 0.86 0.86
assign ((resid 13 and name HN )) ( (resid 13 and name HB2 )) 2.82 1.02 1.02
assign ((resid 13 and name HN )) ( (resid 13 and name HB1 )) 2.82 1.02 1.02
assign ((resid 15 and name HN )) ( (resid 15 and name HB2 )) 2.80 1.00 1.00
assign ((resid 17 and name HN )) ( (resid 17 and name HB2 )) 2.58 0.79 0.79
assign ((resid 17 and name HN )) ( (resid 17 and name HB1 )) 2.73 0.93 0.93
assign ((resid 18 and name HN )) ( (resid 18 and name HB2 )) 2.73 0.93 0.93
assign ((resid 18 and name HN )) ( (resid 18 and name HB1 )) 2.73 0.93 0.93
assign ((resid 20 and name HN )) ( (resid 20 and name HB1 )) 2.64 0.84 0.84
assign ((resid 22 and name HN )) ( (resid 22 and name HB2 )) 2.60 0.81 0.81
assign ((resid 23 and name HN )) ( (resid 23 and name HB2 )) 2.55 0.75 0.75
assign ((resid 25 and name HN )) ( (resid 25 and name HB1 )) 2.76 0.97 0.97
assign ((resid 25 and name HN )) ( (resid 25 and name HB2 )) 2.58 0.79 0.79
assign ((resid 27 and name HN )) ( (resid 27 and name HB2 )) 2.80 1.00 1.00
assign ((resid 31 and name HN )) ( (resid 31 and name HB2 )) 2.64 0.84 0.84
assign ((resid 31 and name HN )) ( (resid 31 and name HB1 )) 2.64 0.84 0.84
assign ((resid 32 and name HN )) ( (resid 32 and name HB2 )) 2.76 0.97 0.97
assign ((resid 35 and name HN )) ( (resid 35 and name HB2 )) 2.87 1.08 1.08
assign ((resid 36 and name HN )) ( (resid 36 and name HB2 )) 2.84 1.04 1.04
assign ((resid 40 and name HN )) ( (resid 40 and name HB2 )) 2.49 0.70 0.70
assign ((resid 41 and name HN )) ( (resid 41 and name HB2 )) 2.93 1.13 1.13
assign ((resid 44 and name HN )) ( (resid 44 and name HB1 )) 2.82 1.02 1.02
assign ((resid 45 and name HN )) ( (resid 45 and name HB1 )) 2.67 0.88 0.88
assign ((resid 47 and name HN )) ( (resid 47 and name HB2 )) 2.55 0.75 0.75
assign ((resid 47 and name HN )) ( (resid 47 and name HB1 )) 2.67 0.88 0.88
assign ((resid 55 and name HN )) ( (resid 55 and name HB1 )) 2.53 0.73 0.73
assign ((resid 57 and name HN )) ( (resid 57 and name HB1 )) 2.51 0.72 0.72
assign ((resid 59 and name HN )) ( (resid 59 and name HB2 )) 2.64 0.84 0.84
assign ((resid 60 and name HN )) ( (resid 60 and name HB2 )) 2.78 0.99 0.99
assign ((resid 60 and name HN )) ( (resid 60 and name HB1 )) 2.84 1.04 1.04
assign ((resid 65 and name HN )) ( (resid 65 and name HB2 )) 2.67 0.88 0.88
assign ((resid 65 and name HN )) ( (resid 65 and name HB1 )) 2.87 1.08 1.08
assign ((resid 66 and name HN )) ( (resid 66 and name HB1 )) 2.84 1.04 1.04
assign ((resid 67 and name HN )) ( (resid 67 and name HB2 )) 2.48 0.68 0.68
assign ((resid 69 and name HN )) ( (resid 69 and name HB2 )) 2.84 1.04 1.04
assign ((resid 69 and name HN )) ( (resid 69 and name HB1 )) 2.84 1.04 1.04
assign ((resid 79 and name HN )) ( (resid 79 and name HB )) 2.58 0.79 0.79
assign ((resid 82 and name HN )) ( (resid 82 and name HB2 )) 2.93 1.13 1.13
assign ((resid 82 and name HN )) ( (resid 82 and name HB1 )) 2.93 1.13 1.13
assign ((resid 85 and name HN )) ( (resid 85 and name HB2 )) 2.93 1.13 1.13
assign ((resid 85 and name HN )) ( (resid 85 and name HB1 )) 2.93 1.13 1.13
assign ((resid 87 and name HN )) ( (resid 87 and name HB2 )) 2.67 0.88 0.88
assign ((resid 87 and name HN )) ( (resid 87 and name HB1 )) 2.64 0.84 0.84
assign ((resid 88 and name HN )) ( (resid 88 and name HB )) 2.58 0.79 0.79
assign ((resid 89 and name HN )) ( (resid 89 and name HB2 )) 2.69 0.90 0.90
assign ((resid 89 and name HN )) ( (resid 89 and name HB1 )) 2.82 1.02 1.02
assign ((resid 90 and name HN )) ( (resid 90 and name HB2 )) 2.69 0.90 0.90
assign ((resid 90 and name HN )) ( (resid 90 and name HB1 )) 2.69 0.90 0.90
assign ((resid 95 and name HN )) ( (resid 95 and name HB2 )) 2.93 1.13 1.13
assign ((resid 95 and name HN )) ( (resid 95 and name HB1 )) 2.93 1.13 1.13
assign ((resid 98 and name HN )) ( (resid 98 and name HB2 )) 2.96 1.17 1.17
assign ((resid 98 and name HN )) ( (resid 98 and name HB1 )) 2.96 1.17 1.17
assign ((resid 99 and name HN )) ( (resid 99 and name HB2 )) 2.64 0.84 0.84
assign ((resid 100 and name HN )) ( (resid 100 and name HB2 )) 2.87 1.08 1.08
assign ((resid 100 and name HN )) ( (resid 100 and name HB1 )) 2.87 1.08 1.08
assign ((resid 102 and name HN )) ( (resid 102 and name HB1 )) 2.46 0.66 0.66
assign ((resid 104 and name HN )) ( (resid 104 and name HB )) 2.48 0.68 0.68
assign ((resid 107 and name HN )) ( (resid 107 and name HB1 )) 2.84 1.04 1.04
assign ((resid 108 and name HN )) ( (resid 108 and name HB2 )) 3.00 1.20 1.20
assign ((resid 108 and name HN )) ( (resid 108 and name HB1 )) 3.00 1.20 1.20
assign ((resid 110 and name HN )) ( (resid 110 and name HB2 )) 2.64 0.84 0.84
assign ((resid 110 and name HN )) ( (resid 110 and name HB1 )) 2.78 0.99 0.99
assign ((resid 112 and name HN )) ( (resid 112 and name HB2 )) 2.60 0.81 0.81
assign ((resid 112 and name HN )) ( (resid 112 and name HB1 )) 2.60 0.81 0.81
assign ((resid 113 and name HN )) ( (resid 113 and name HB1 )) 2.64 0.84 0.84
assign ((resid 119 and name HN )) ( (resid 119 and name HB )) 2.49 0.70 0.70
assign ((resid 121 and name HN )) ( (resid 121 and name HB2 )) 2.62 0.82 0.82
assign ((resid 124 and name HN )) ( (resid 124 and name HB1 )) 2.73 0.93 0.93
assign ((resid 125 and name HN )) ( (resid 125 and name HB2 )) 2.73 0.93 0.93
assign ((resid 125 and name HN )) ( (resid 125 and name HB1 )) 2.73 0.93 0.93
assign ((resid 126 and name HN )) ( (resid 126 and name HB2 )) 2.89 1.09 1.09
assign ((resid 126 and name HN )) ( (resid 126 and name HB1 )) 2.89 1.09 1.09
assign ((resid 5 and name HN )) ( (resid 6 and name HN )) 3.30 1.51 1.51
assign ((resid 6 and name HN )) ( (resid 7 and name HN )) 2.78 0.99 0.99
assign ((resid 8 and name HN )) ( (resid 9 and name HN )) 2.58 0.79 0.79
assign ((resid 9 and name HN )) ( (resid 10 and name HN )) 2.64 0.84 0.84
assign ((resid 15 and name HN )) ( (resid 16 and name HN )) 2.64 0.84 0.84
assign ((resid 16 and name HN )) ( (resid 17 and name HN )) 2.57 0.77 0.77
assign ((resid 17 and name HN )) ( (resid 18 and name HN )) 2.62 0.82 0.82
assign ((resid 20 and name HN )) ( (resid 21 and name HN )) 2.49 0.70 0.70
assign ((resid 21 and name HN )) ( (resid 22 and name HN )) 2.55 0.75 0.75
assign ((resid 22 and name HN )) ( (resid 23 and name HN )) 3.09 1.29 1.29
assign ((resid 23 and name HN )) ( (resid 24 and name HN )) 2.55 0.75 0.75
assign ((resid 26 and name HN )) ( (resid 27 and name HN )) 2.69 0.90 0.90
assign ((resid 30 and name HN )) ( (resid 31 and name HN )) 2.71 0.91 0.91
assign ((resid 31 and name HN )) ( (resid 32 and name HN )) 2.80 1.00 1.00
assign ((resid 33 and name HN )) ( (resid 34 and name HN )) 2.64 0.84 0.84
assign ((resid 34 and name HN )) ( (resid 35 and name HN )) 2.64 0.84 0.84
assign ((resid 36 and name HN )) ( (resid 37 and name HN )) 2.78 0.99 0.99
assign ((resid 39 and name HN )) ( (resid 40 and name HN )) 2.76 0.97 0.97
assign ((resid 40 and name HN )) ( (resid 41 and name HN )) 2.51 0.72 0.72
assign ((resid 41 and name HN )) ( (resid 42 and name HN )) 2.64 0.84 0.84
assign ((resid 45 and name HN )) ( (resid 46 and name HN )) 2.55 0.75 0.75
assign ((resid 47 and name HN )) ( (resid 48 and name HN )) 2.98 1.18 1.18
assign ((resid 48 and name HN )) ( (resid 49 and name HN )) 2.69 0.90 0.90
assign ((resid 49 and name HN )) ( (resid 50 and name HN )) 2.67 0.88 0.88
assign ((resid 52 and name HN )) ( (resid 53 and name HN )) 2.57 0.77 0.77
assign ((resid 58 and name HN )) ( (resid 59 and name HN )) 2.57 0.77 0.77
assign ((resid 59 and name HN )) ( (resid 60 and name HN )) 3.03 1.24 1.24
assign ((resid 62 and name HN )) ( (resid 63 and name HN )) 3.14 1.35 1.35
assign ((resid 64 and name HN )) ( (resid 65 and name HN )) 2.66 0.86 0.86
assign ((resid 67 and name HN )) ( (resid 68 and name HN )) 2.60 0.81 0.81
assign ((resid 69 and name HN )) ( (resid 70 and name HN )) 2.75 0.95 0.95
assign ((resid 70 and name HN )) ( (resid 71 and name HN )) 2.55 0.75 0.75
assign ((resid 74 and name HN )) ( (resid 75 and name HN )) 2.62 0.82 0.82
assign ((resid 76 and name HN )) ( (resid 77 and name HN )) 3.09 1.29 1.29
assign ((resid 81 and name HN )) ( (resid 82 and name HN )) 2.51 0.72 0.72
assign ((resid 82 and name HN )) ( (resid 83 and name HN )) 2.73 0.93 0.93
assign ((resid 84 and name HN )) ( (resid 85 and name HN )) 2.62 0.82 0.82
assign ((resid 85 and name HN )) ( (resid 86 and name HN )) 2.62 0.82 0.82
assign ((resid 88 and name HN )) ( (resid 89 and name HN )) 2.78 0.99 0.99
assign ((resid 90 and name HN )) ( (resid 91 and name HN )) 2.75 0.95 0.95
assign ((resid 91 and name HN )) ( (resid 92 and name HN )) 2.73 0.93 0.93
assign ((resid 93 and name HN )) ( (resid 94 and name HN )) 2.85 1.06 1.06
assign ((resid 95 and name HN )) ( (resid 96 and name HN )) 2.80 1.00 1.00
assign ((resid 97 and name HN )) ( (resid 98 and name HN )) 2.91 1.11 1.11
assign ((resid 100 and name HN )) ( (resid 101 and name HN )) 3.48 1.69 1.69
assign ((resid 103 and name HN )) ( (resid 104 and name HN )) 2.58 0.79 0.79
assign ((resid 107 and name HN )) ( (resid 108 and name HN )) 2.66 0.86 0.86
assign ((resid 108 and name HN )) ( (resid 109 and name HN )) 2.71 0.91 0.91
assign ((resid 110 and name HN )) ( (resid 111 and name HN )) 2.60 0.81 0.81
assign ((resid 112 and name HN )) ( (resid 113 and name HN )) 2.73 0.93 0.93
assign ((resid 117 and name HN )) ( (resid 118 and name HN )) 2.58 0.79 0.79
assign ((resid 118 and name HN )) ( (resid 119 and name HN )) 2.60 0.81 0.81
assign ((resid 119 and name HN )) ( (resid 120 and name HN )) 2.67 0.88 0.88
assign ((resid 121 and name HN )) ( (resid 122 and name HN )) 2.58 0.79 0.79
assign ((resid 122 and name HN )) ( (resid 123 and name HN )) 2.57 0.77 0.77
assign ((resid 6 and name HA )) ( (resid 9 and name HN )) 2.93 1.13 1.13
assign ((resid 8 and name HA )) ( (resid 10 and name HN )) 3.50 1.71 1.71
assign ((resid 7 and name HA )) ( (resid 10 and name HN )) 3.07 1.27 1.27
assign ((resid 6 and name HA )) ( (resid 10 and name HN )) 3.52 1.72 1.72
assign ((resid 10 and name HN )) ( (resid 12 and name HN )) 3.34 1.54 1.54
assign ((resid 11 and name HN )) ( (resid 33 and name HA )) 3.30 1.51 1.51
assign ((resid 10 and name HA )) ( (resid 12 and name HN )) 3.43 1.63 1.63
assign ((resid 10 and name HA )) ( (resid 13 and name HN )) 3.05 1.26 1.26
assign ((resid 9 and name HA )) ( (resid 13 and name HN )) 3.39 1.60 1.60
assign ((resid 10 and name HA )) ( (resid 14 and name HN )) 3.32 1.53 1.53
assign ((resid 14 and name HN )) ( (resid 16 and name HN )) 3.18 1.38 1.38
assign ((resid 13 and name HA )) ( (resid 16 and name HN )) 2.98 1.18 1.18
assign ((resid 16 and name HN )) ( (resid 18 and name HN )) 3.30 1.51 1.51
assign ((resid 14 and name HA )) ( (resid 17 and name HN )) 2.91 1.11 1.11
assign ((resid 13 and name HA )) ( (resid 17 and name HN )) 2.89 1.09 1.09
assign ((resid 16 and name HA )) ( (resid 18 and name HN )) 3.16 1.36 1.36
assign ((resid 14 and name HA )) ( (resid 18 and name HN )) 3.50 1.71 1.71
assign ((resid 24 and name HN )) ( (resid 26 and name HN )) 3.00 1.20 1.20
assign ((resid 25 and name HN )) ( (resid 27 and name HN )) 3.38 1.58 1.58
assign ((resid 24 and name HA )) ( (resid 26 and name HN )) 3.56 1.76 1.76
assign ((resid 23 and name HA )) ( (resid 26 and name HN )) 2.89 1.09 1.09
assign ((resid 24 and name HA )) ( (resid 27 and name HN )) 3.16 1.36 1.36
assign ((resid 23 and name HA )) ( (resid 27 and name HN )) 3.30 1.51 1.51
assign ((resid 25 and name HA )) ( (resid 28 and name HN )) 3.00 1.20 1.20
assign ((resid 24 and name HA )) ( (resid 28 and name HN )) 3.75 1.96 1.96
assign ((resid 28 and name HN )) ( (resid 30 and name HN )) 3.72 1.92 1.92
assign ((resid 26 and name HA )) ( (resid 29 and name HN )) 3.07 1.27 1.27
assign ((resid 28 and name HA )) ( (resid 31 and name HN )) 2.71 0.91 0.91
assign ((resid 31 and name HA )) ( (resid 33 and name HN )) 3.43 1.63 1.63
assign ((resid 29 and name HA )) ( (resid 33 and name HN )) 3.38 1.58 1.58
assign ((resid 33 and name HN )) ( (resid 35 and name HN )) 3.59 1.80 1.80
assign ((resid 32 and name HA )) ( (resid 35 and name HN )) 2.94 1.15 1.15
assign ((resid 33 and name HA )) ( (resid 36 and name HN )) 3.03 1.24 1.24
assign ((resid 32 and name HA )) ( (resid 36 and name HN )) 3.36 1.56 1.56
assign ((resid 34 and name HA )) ( (resid 37 and name HN )) 2.91 1.11 1.11
assign ((resid 7 and name HA )) ( (resid 37 and name HN )) 3.48 1.69 1.69
assign ((resid 35 and name HA )) ( (resid 38 and name HN )) 3.27 1.47 1.47
assign ((resid 36 and name HA )) ( (resid 39 and name HN )) 2.94 1.15 1.15
assign ((resid 38 and name HA )) ( (resid 41 and name HN )) 2.67 0.88 0.88
assign ((resid 39 and name HA )) ( (resid 42 and name HN )) 3.11 1.31 1.31
assign ((resid 40 and name HA )) ( (resid 43 and name HN )) 3.02 1.22 1.22
assign ((resid 39 and name HA )) ( (resid 43 and name HN )) 3.50 1.71 1.71
assign ((resid 63 and name HA )) ( (resid 65 and name HN )) 3.02 1.22 1.22
assign ((resid 65 and name HN )) ( (resid 67 and name HN )) 3.66 1.87 1.87
assign ((resid 63 and name HA )) ( (resid 66 and name HN )) 2.87 1.08 1.08
assign ((resid 64 and name HA )) ( (resid 67 and name HN )) 2.91 1.11 1.11
assign ((resid 63 and name HA )) ( (resid 67 and name HN )) 3.65 1.85 1.85
assign ((resid 64 and name HA )) ( (resid 68 and name HN )) 3.16 1.36 1.36
assign ((resid 67 and name HA )) ( (resid 69 and name HN )) 3.83 2.03 2.03
assign ((resid 69 and name HN )) ( (resid 71 and name HN )) 3.21 1.42 1.42
assign ((resid 67 and name HA )) ( (resid 70 and name HN )) 3.30 1.51 1.51
assign ((resid 68 and name HA )) ( (resid 71 and name HN )) 3.09 1.29 1.29
assign ((resid 79 and name HA )) ( (resid 81 and name HN )) 3.59 1.80 1.80
assign ((resid 81 and name HN )) ( (resid 83 and name HN )) 3.30 1.51 1.51
assign ((resid 79 and name HA )) ( (resid 82 and name HN )) 3.07 1.27 1.27
assign ((resid 81 and name HA )) ( (resid 83 and name HN )) 3.61 1.81 1.81
assign ((resid 80 and name HA )) ( (resid 83 and name HN )) 3.07 1.27 1.27
assign ((resid 79 and name HA )) ( (resid 83 and name HN )) 3.29 1.49 1.49
assign ((resid 82 and name HA )) ( (resid 84 and name HN )) 3.61 1.81 1.81
assign ((resid 82 and name HN )) ( (resid 84 and name HN )) 3.36 1.56 1.56
assign ((resid 80 and name HA )) ( (resid 84 and name HN )) 3.34 1.54 1.54
assign ((resid 82 and name HA )) ( (resid 85 and name HN )) 3.11 1.31 1.31
assign ((resid 83 and name HA )) ( (resid 86 and name HN )) 2.85 1.06 1.06
assign ((resid 86 and name HN )) ( (resid 88 and name HN )) 3.90 2.10 2.10
assign ((resid 84 and name HA )) ( (resid 87 and name HN )) 3.00 1.20 1.20
assign ((resid 83 and name HA )) ( (resid 87 and name HN )) 3.09 1.29 1.29
assign ((resid 85 and name HA )) ( (resid 88 and name HN )) 3.38 1.58 1.58
assign ((resid 84 and name HA )) ( (resid 88 and name HN )) 3.38 1.58 1.58
assign ((resid 86 and name HA )) ( (resid 89 and name HN )) 3.09 1.29 1.29
assign ((resid 89 and name HN )) ( (resid 91 and name HN )) 3.90 2.10 2.10
assign ((resid 101 and name HA )) ( (resid 103 and name HN )) 3.88 2.08 2.08
assign ((resid 103 and name HN )) ( (resid 105 and name HN )) 3.20 1.40 1.40
assign ((resid 103 and name HA )) ( (resid 105 and name HN )) 3.43 1.63 1.63
assign ((resid 102 and name HA )) ( (resid 105 and name HN )) 2.96 1.17 1.17
assign ((resid 101 and name HA )) ( (resid 105 and name HN )) 3.20 1.40 1.40
assign ((resid 106 and name HN )) ( (resid 108 and name HN )) 3.70 1.90 1.90
assign ((resid 104 and name HA )) ( (resid 107 and name HN )) 3.11 1.31 1.31
assign ((resid 104 and name HA )) ( (resid 108 and name HN )) 3.86 2.07 2.07
assign ((resid 107 and name HN )) ( (resid 109 and name HN )) 3.36 1.56 1.56
assign ((resid 105 and name HA )) ( (resid 109 and name HN )) 3.66 1.87 1.87
assign ((resid 108 and name HA )) ( (resid 110 and name HN )) 3.43 1.63 1.63
assign ((resid 107 and name HA )) ( (resid 110 and name HN )) 2.94 1.15 1.15
assign ((resid 110 and name HN )) ( (resid 112 and name HN )) 3.32 1.53 1.53
assign ((resid 108 and name HA )) ( (resid 111 and name HN )) 2.87 1.08 1.08
assign ((resid 108 and name HA )) ( (resid 112 and name HN )) 3.36 1.56 1.56
assign ((resid 112 and name HA )) ( (resid 115 and name HN )) 3.12 1.33 1.33
assign ((resid 115 and name HN )) ( (resid 117 and name HN )) 3.21 1.42 1.42
assign ((resid 114 and name HN )) ( (resid 116 and name HN )) 3.45 1.65 1.65
assign ((resid 113 and name HA )) ( (resid 116 and name HN )) 3.25 1.45 1.45
assign ((resid 112 and name HA )) ( (resid 116 and name HN )) 3.00 1.20 1.20
assign ((resid 115 and name HA )) ( (resid 118 and name HN )) 2.66 0.86 0.86
assign ((resid 116 and name HA )) ( (resid 119 and name HN )) 2.89 1.09 1.09
assign ((resid 116 and name HA )) ( (resid 120 and name HN )) 3.20 1.40 1.40
assign ((resid 118 and name HA )) ( (resid 121 and name HN )) 3.09 1.29 1.29
assign ((resid 119 and name HA )) ( (resid 122 and name HN )) 2.93 1.13 1.13
assign ((resid 3 and name HA )) ( (resid 5 and name HN )) 3.66 1.87 1.87
assign ((resid 5 and name HN )) ( (resid 8 and name HB* )) 3.42 1.62 1.62
assign ((resid 6 and name HN )) ( (resid 9 and name HB )) 3.48 1.69 1.69
assign ((resid 5 and name HA )) ( (resid 7 and name HN )) 3.30 1.51 1.51
assign ((resid 8 and name HN )) ( (resid 9 and name HB )) 3.43 1.63 1.63
assign ((resid 5 and name HB )) ( (resid 8 and name HN )) 3.36 1.56 1.56
assign ((resid 5 and name HN )) ( (resid 8 and name HN )) 3.27 1.47 1.47
assign ((resid 5 and name HN )) ( (resid 9 and name HN )) 3.30 1.51 1.51
assign ((resid 19 and name HA )) ( (resid 21 and name HN )) 3.27 1.47 1.47
assign ((resid 19 and name HB* )) ( (resid 21 and name HN )) 4.01 2.22 2.22
assign ((resid 18 and name HA )) ( (resid 21 and name HN )) 3.00 1.20 1.20
assign ((resid 20 and name HB1 )) ( (resid 22 and name HN )) 3.20 1.40 1.40
assign ((resid 20 and name HB2 )) ( (resid 22 and name HN )) 3.20 1.40 1.40
assign ((resid 18 and name HA )) ( (resid 22 and name HN )) 3.02 1.22 1.22
assign ((resid 20 and name HN )) ( (resid 22 and name HN )) 2.96 1.17 1.17
assign ((resid 22 and name HA )) ( (resid 24 and name HN )) 3.14 1.35 1.35
assign ((resid 43 and name HA )) ( (resid 46 and name HN )) 2.93 1.13 1.13
assign ((resid 42 and name HA )) ( (resid 47 and name HN )) 2.98 1.18 1.18
assign ((resid 49 and name HN )) ( (resid 51 and name HN )) 3.38 1.58 1.58
assign ((resid 47 and name HA )) ( (resid 49 and name HN )) 3.23 1.44 1.44
assign ((resid 49 and name HA )) ( (resid 52 and name HN )) 3.23 1.44 1.44
assign ((resid 51 and name HA )) ( (resid 54 and name HN )) 3.18 1.38 1.38
assign ((resid 53 and name HA )) ( (resid 55 and name HN )) 3.11 1.31 1.31
assign ((resid 61 and name HN )) ( (resid 64 and name HN )) 3.45 1.65 1.65
assign ((resid 69 and name HN )) ( (resid 70 and name HB2 )) 3.90 2.10 2.10
assign ((resid 72 and name HA )) ( (resid 74 and name HN )) 3.09 1.29 1.29
assign ((resid 72 and name HA )) ( (resid 75 and name HN )) 3.09 1.29 1.29
assign ((resid 73 and name HA )) ( (resid 76 and name HN )) 2.80 1.00 1.00
assign ((resid 76 and name HA )) ( (resid 78 and name HN )) 3.14 1.35 1.35
assign ((resid 98 and name HN )) ( (resid 99 and name HN )) 2.55 0.75 0.75
assign ((resid 100 and name HB1 )) ( (resid 102 and name HN )) 3.34 1.54 1.54
assign ((resid 100 and name HB2 )) ( (resid 102 and name HN )) 3.34 1.54 1.54
assign ((resid 105 and name HN )) ( (resid 106 and name HB* )) 3.78 1.98 1.98
assign ((resid 109 and name HN )) ( (resid 111 and name HN )) 3.21 1.42 1.42
assign ((resid 106 and name HA )) ( (resid 109 and name HN )) 3.02 1.22 1.22
assign ((resid 116 and name HA )) ( (resid 117 and name HN )) 2.67 0.88 0.88
assign ((resid 86 and name HN )) ( (resid 87 and name HN )) 2.67 0.88 0.88
assign ((resid 117 and name HA )) ( (resid 120 and name HN )) 3.09 1.29 1.29
assign ((resid 48 and name HA )) ( (resid 52 and name HN )) 2.80 1.00 1.00
assign ((resid 12 and name HG* )) ( (resid 13 and name HN )) 4.34 2.54 2.54
assign ((resid 13 and name HG2 )) ( (resid 14 and name HN )) 3.90 2.10 2.10
assign ((resid 13 and name HG1 )) ( (resid 14 and name HN )) 3.90 2.10 2.10
assign ((resid 20 and name HG* )) ( (resid 21 and name HN )) 4.34 2.54 2.54
assign ((resid 41 and name HG )) ( (resid 42 and name HN )) 3.57 1.78 1.78
assign ((resid 55 and name HG2 )) ( (resid 56 and name HN )) 3.90 2.10 2.10
assign ((resid 55 and name HG1 )) ( (resid 56 and name HN )) 3.90 2.10 2.10
assign ((resid 59 and name HG* )) ( (resid 60 and name HN )) 4.34 2.54 2.54
assign ((resid 82 and name HG )) ( (resid 83 and name HN )) 3.70 1.90 1.90
assign ((resid 96 and name HG* )) ( (resid 97 and name HN )) 4.34 2.54 2.54
assign ((resid 108 and name HG* )) ( (resid 109 and name HN )) 4.27 2.47 2.47
assign ((resid 114 and name HG )) ( (resid 115 and name HN )) 3.72 1.92 1.92
assign ((resid 116 and name HG12 )) ( (resid 117 and name HN )) 3.90 2.10 2.10
assign ((resid 116 and name HG11 )) ( (resid 117 and name HN )) 3.90 2.10 2.10
assign ((resid 8 and name HN )) ( (resid 8 and name HG2 )) 2.89 1.09 1.09
assign ((resid 8 and name HN )) ( (resid 8 and name HG1 )) 2.89 1.09 1.09
assign ((resid 13 and name HN )) ( (resid 13 and name HG1 )) 2.76 0.97 0.97
assign ((resid 15 and name HN )) ( (resid 15 and name HG2 )) 2.75 0.95 0.95
assign ((resid 15 and name HN )) ( (resid 15 and name HG1 )) 2.75 0.95 0.95
assign ((resid 16 and name HN )) ( (resid 16 and name HG2 )) 2.94 1.15 1.15
assign ((resid 18 and name HN )) ( (resid 18 and name HG )) 3.34 1.54 1.54
assign ((resid 20 and name HN )) ( (resid 20 and name HG* )) 3.55 1.75 1.75
assign ((resid 28 and name HN )) ( (resid 28 and name HG2 )) 3.54 1.74 1.74
assign ((resid 28 and name HN )) ( (resid 28 and name HG1 )) 3.54 1.74 1.74
assign ((resid 32 and name HN )) ( (resid 32 and name HG2 )) 3.05 1.26 1.26
assign ((resid 35 and name HN )) ( (resid 35 and name HG2 )) 3.05 1.26 1.26
assign ((resid 35 and name HN )) ( (resid 35 and name HG1 )) 3.05 1.26 1.26
assign ((resid 36 and name HN )) ( (resid 36 and name HG2 )) 3.05 1.26 1.26
assign ((resid 36 and name HN )) ( (resid 36 and name HG1 )) 3.05 1.26 1.26
assign ((resid 38 and name HN )) ( (resid 38 and name HG12 )) 2.89 1.09 1.09
assign ((resid 38 and name HN )) ( (resid 38 and name HG11 )) 2.89 1.09 1.09
assign ((resid 40 and name HN )) ( (resid 40 and name HG )) 2.73 0.93 0.93
assign ((resid 44 and name HN )) ( (resid 44 and name HG2 )) 3.00 1.20 1.20
assign ((resid 47 and name HN )) ( (resid 47 and name HG )) 2.62 0.82 0.82
assign ((resid 48 and name HN )) ( (resid 48 and name HG2 )) 3.30 1.51 1.51
assign ((resid 48 and name HN )) ( (resid 48 and name HG1 )) 3.30 1.51 1.51
assign ((resid 50 and name HN )) ( (resid 50 and name HG11 )) 3.16 1.36 1.36
assign ((resid 55 and name HN )) ( (resid 55 and name HG2 )) 3.27 1.47 1.47
assign ((resid 55 and name HN )) ( (resid 55 and name HG1 )) 3.27 1.47 1.47
assign ((resid 59 and name HN )) ( (resid 59 and name HG* )) 3.46 1.66 1.66
assign ((resid 60 and name HN )) ( (resid 60 and name HD1 )) 3.03 1.24 1.24
assign ((resid 61 and name HN )) ( (resid 61 and name HG2 )) 3.29 1.49 1.49
assign ((resid 61 and name HN )) ( (resid 61 and name HG1 )) 3.29 1.49 1.49
assign ((resid 65 and name HN )) ( (resid 65 and name HG )) 2.67 0.88 0.88
assign ((resid 67 and name HN )) ( (resid 67 and name HG1 )) 3.34 1.54 1.54
assign ((resid 70 and name HN )) ( (resid 70 and name HG1 )) 3.25 1.45 1.45
assign ((resid 71 and name HN )) ( (resid 71 and name HG )) 2.53 0.73 0.73
assign ((resid 72 and name HN )) ( (resid 72 and name HG )) 2.75 0.95 0.95
assign ((resid 73 and name HN )) ( (resid 73 and name HG1 )) 3.29 1.49 1.49
assign ((resid 79 and name HN )) ( (resid 79 and name HG1* )) 3.65 1.86 1.86
assign ((resid 81 and name HN )) ( (resid 81 and name HG12 )) 2.98 1.18 1.18
assign ((resid 82 and name HN )) ( (resid 82 and name HG )) 2.40 0.61 0.61
assign ((resid 84 and name HN )) ( (resid 84 and name HG2 )) 3.50 1.71 1.71
assign ((resid 84 and name HN )) ( (resid 84 and name HG1 )) 3.50 1.71 1.71
assign ((resid 99 and name HN )) ( (resid 99 and name HG )) 2.75 0.95 0.95
assign ((resid 102 and name HN )) ( (resid 102 and name HG1 )) 3.03 1.24 1.24
assign ((resid 103 and name HN )) ( (resid 103 and name HG* )) 3.47 1.68 1.68
assign ((resid 105 and name HN )) ( (resid 105 and name HG1 )) 3.21 1.42 1.42
assign ((resid 106 and name HN )) ( (resid 106 and name HG2 )) 3.30 1.51 1.51
assign ((resid 106 and name HN )) ( (resid 106 and name HG1 )) 3.30 1.51 1.51
assign ((resid 108 and name HN )) ( (resid 108 and name HG* )) 3.49 1.70 1.70
assign ((resid 109 and name HN )) ( (resid 109 and name HG2 )) 3.21 1.42 1.42
assign ((resid 109 and name HN )) ( (resid 109 and name HG1 )) 3.21 1.42 1.42
assign ((resid 112 and name HN )) ( (resid 112 and name HG2 )) 3.79 1.99 1.99
assign ((resid 113 and name HN )) ( (resid 113 and name HG2 )) 2.89 1.09 1.09
assign ((resid 113 and name HN )) ( (resid 113 and name HG1 )) 2.89 1.09 1.09
assign ((resid 114 and name HN )) ( (resid 114 and name HG )) 2.89 1.09 1.09
assign ((resid 125 and name HN )) ( (resid 125 and name HD2 )) 3.90 2.10 2.10
assign ((resid 53 and name HA )) ( (resid 53 and name HD21 )) 3.57 1.78 1.78
assign ((resid 53 and name HA )) ( (resid 53 and name HD22 )) 3.57 1.78 1.78
assign ((resid 64 and name HA )) ( (resid 64 and name HD21 )) 3.27 1.47 1.47
assign ((resid 64 and name HA )) ( (resid 64 and name HD22 )) 3.27 1.47 1.47
assign ((resid 75 and name HA )) ( (resid 75 and name HD22 )) 3.50 1.71 1.71
assign ((resid 25 and name HB1 )) ( (resid 25 and name HE21 )) 3.90 2.10 2.10
assign ((resid 25 and name HB2 )) ( (resid 25 and name HE21 )) 3.90 2.10 2.10
assign ((resid 25 and name HB1 )) ( (resid 25 and name HE22 )) 3.90 2.10 2.10
assign ((resid 25 and name HB2 )) ( (resid 25 and name HE22 )) 3.90 2.10 2.10
assign ((resid 4 and name HB* )) ( (resid 9 and name HN )) 4.34 2.54 2.54
assign ((resid 20 and name HG* )) ( (resid 22 and name HN )) 4.34 2.54 2.54
assign ((resid 17 and name HB2 )) ( (resid 26 and name HN )) 3.79 1.99 1.99
assign ((resid 17 and name HB1 )) ( (resid 26 and name HN )) 3.18 1.38 1.38
assign ((resid 10 and name HB1 )) ( (resid 33 and name HN )) 3.25 1.45 1.45
assign ((resid 10 and name HB2 )) ( (resid 33 and name HN )) 3.52 1.72 1.72
assign ((resid 41 and name HA )) ( (resid 45 and name HD21 )) 3.68 1.89 1.89
assign ((resid 41 and name HB1 )) ( (resid 45 and name HD21 )) 4.78 2.98 2.98
assign ((resid 41 and name HB1 )) ( (resid 45 and name HD22 )) 4.78 2.98 2.98
assign ((resid 48 and name HB1 )) ( (resid 52 and name HD21 )) 4.38 2.59 2.59
assign ((resid 48 and name HD* )) ( (resid 52 and name HD21 )) 4.34 2.54 2.54
assign ((resid 48 and name HB1 )) ( (resid 52 and name HD22 )) 4.38 2.59 2.59
assign ((resid 48 and name HD* )) ( (resid 52 and name HD22 )) 4.34 2.54 2.54
assign ((resid 49 and name HA )) ( (resid 52 and name HD21 )) 3.65 1.85 1.85
assign ((resid 49 and name HA )) ( (resid 52 and name HD22 )) 3.65 1.85 1.85
assign ((resid 50 and name HA )) ( (resid 53 and name HD21 )) 3.50 1.71 1.71
assign ((resid 50 and name HA )) ( (resid 53 and name HD22 )) 3.50 1.71 1.71
assign ((resid 55 and name HD* )) ( (resid 56 and name HN )) 4.34 2.54 2.54
assign ((resid 64 and name HN )) ( (resid 65 and name HG )) 3.68 1.89 1.89
assign ((resid 79 and name HN )) ( (resid 80 and name HD2 )) 2.87 1.08 1.08
assign ((resid 79 and name HN )) ( (resid 80 and name HD1 )) 2.87 1.08 1.08
assign ((resid 112 and name HN )) ( (resid 112 and name HG1 )) 3.79 1.99 1.99
assign ((resid 9 and name HG2* )) ( (resid 10 and name HN )) 3.44 1.64 1.64
assign ((resid 9 and name HG1* )) ( (resid 10 and name HN )) 3.36 1.57 1.57
assign ((resid 18 and name HD1* )) ( (resid 19 and name HN )) 4.41 2.61 2.61
assign ((resid 23 and name HD2* )) ( (resid 24 and name HN )) 4.05 2.25 2.25
assign ((resid 24 and name HG1* )) ( (resid 25 and name HN )) 3.31 1.51 1.51
assign ((resid 24 and name HG2* )) ( (resid 25 and name HN )) 3.63 1.84 1.84
assign ((resid 38 and name HG2* )) ( (resid 39 and name HN )) 3.74 1.95 1.95
assign ((resid 41 and name HD1* )) ( (resid 42 and name HN )) 3.85 2.05 2.05
assign ((resid 42 and name HG1* )) ( (resid 43 and name HN )) 3.56 1.77 1.77
assign ((resid 54 and name HG2* )) ( (resid 55 and name HN )) 3.76 1.96 1.96
assign ((resid 71 and name HD1* )) ( (resid 72 and name HN )) 3.69 1.89 1.89
assign ((resid 71 and name HD2* )) ( (resid 72 and name HN )) 3.74 1.95 1.95
assign ((resid 82 and name HD1* )) ( (resid 83 and name HN )) 3.94 2.14 2.14
assign ((resid 82 and name HD2* )) ( (resid 83 and name HN )) 4.17 2.38 2.38
assign ((resid 88 and name HG2* )) ( (resid 89 and name HN )) 3.63 1.84 1.84
assign ((resid 97 and name HD2* )) ( (resid 98 and name HN )) 4.41 2.61 2.61
assign ((resid 99 and name HD2* )) ( (resid 100 and name HN )) 3.47 1.68 1.68
assign ((resid 107 and name HD1* )) ( (resid 108 and name HN )) 4.28 2.49 2.49
assign ((resid 111 and name HG2* )) ( (resid 112 and name HN )) 3.63 1.84 1.84
assign ((resid 114 and name HD2* )) ( (resid 115 and name HN )) 4.08 2.29 2.29
assign ((resid 114 and name HD1* )) ( (resid 115 and name HN )) 4.24 2.45 2.45
assign ((resid 116 and name HG2* )) ( (resid 117 and name HN )) 3.07 1.28 1.28
assign ((resid 119 and name HG2* )) ( (resid 120 and name HN )) 3.51 1.71 1.71
assign ((resid 122 and name HD1* )) ( (resid 123 and name HN )) 3.63 1.84 1.84
assign ((resid 122 and name HD2* )) ( (resid 123 and name HN )) 3.97 2.18 2.18
assign ((resid 23 and name HN )) ( (resid 23 and name HD1* )) 3.36 1.57 1.57
assign ((resid 23 and name HN )) ( (resid 23 and name HD2* )) 3.42 1.62 1.62
assign ((resid 38 and name HN )) ( (resid 38 and name HD1* )) 3.44 1.64 1.64
assign ((resid 40 and name HN )) ( (resid 40 and name HD1* )) 3.35 1.55 1.55
assign ((resid 47 and name HN )) ( (resid 47 and name HD1* )) 3.53 1.73 1.73
assign ((resid 65 and name HN )) ( (resid 65 and name HD2* )) 3.76 1.96 1.96
assign ((resid 65 and name HN )) ( (resid 65 and name HD1* )) 3.78 1.98 1.98
assign ((resid 79 and name HN )) ( (resid 79 and name HD1* )) 3.36 1.57 1.57
assign ((resid 82 and name HN )) ( (resid 82 and name HD1* )) 3.60 1.80 1.80
assign ((resid 99 and name HN )) ( (resid 99 and name HD2* )) 3.74 1.95 1.95
assign ((resid 104 and name HN )) ( (resid 104 and name HG1* )) 3.29 1.50 1.50
assign ((resid 104 and name HN )) ( (resid 104 and name HG2* )) 3.25 1.46 1.46
assign ((resid 114 and name HN )) ( (resid 114 and name HD2* )) 3.80 2.00 2.00
assign ((resid 114 and name HN )) ( (resid 114 and name HD1* )) 3.67 1.87 1.87
assign ((resid 116 and name HN )) ( (resid 116 and name HD1* )) 3.31 1.51 1.51
assign ((resid 119 and name HN )) ( (resid 119 and name HG1* )) 3.20 1.41 1.41
assign ((resid 122 and name HN )) ( (resid 122 and name HD1* )) 3.56 1.77 1.77
assign ((resid 122 and name HN )) ( (resid 122 and name HD2* )) 3.51 1.71 1.71
assign ((resid 5 and name HG2* )) ( (resid 8 and name HN )) 4.41 2.61 2.61
assign ((resid 8 and name HN )) ( (resid 119 and name HG2* )) 4.26 2.47 2.47
assign ((resid 8 and name HN )) ( (resid 119 and name HG1* )) 3.80 2.00 2.00
assign ((resid 8 and name HN )) ( (resid 115 and name HG1* )) 3.90 2.11 2.11
assign ((resid 12 and name HN )) ( (resid 33 and name HB* )) 4.19 2.40 2.40
assign ((resid 23 and name HN )) ( (resid 26 and name HB* )) 4.41 2.61 2.61
assign ((resid 23 and name HN )) ( (resid 24 and name HG2* )) 4.35 2.56 2.56
assign ((resid 34 and name HN )) ( (resid 86 and name HB* )) 4.28 2.49 2.49
assign ((resid 34 and name HN )) ( (resid 111 and name HG2* )) 4.08 2.29 2.29
assign ((resid 7 and name HB* )) ( (resid 36 and name HN )) 3.96 2.16 2.16
assign ((resid 37 and name HN )) ( (resid 118 and name HB* )) 3.72 1.93 1.93
assign ((resid 42 and name HG1* )) ( (resid 47 and name HN )) 3.35 1.55 1.55
assign ((resid 47 and name HD2* )) ( (resid 48 and name HN )) 3.67 1.87 1.87
assign ((resid 54 and name HG1* )) ( (resid 56 and name HN )) 3.85 2.05 2.05
assign ((resid 43 and name HD1* )) ( (resid 60 and name HE1 )) 4.26 2.47 2.47
assign ((resid 54 and name HG1* )) ( (resid 60 and name HE1 )) 3.81 2.02 2.02
assign ((resid 50 and name HD1* )) ( (resid 67 and name HN )) 4.35 2.56 2.56
assign ((resid 38 and name HG2* )) ( (resid 69 and name HN )) 3.72 1.93 1.93
assign ((resid 50 and name HD1* )) ( (resid 69 and name HN )) 4.32 2.52 2.52
assign ((resid 69 and name HN )) ( (resid 72 and name HD2* )) 4.21 2.41 2.41
assign ((resid 75 and name HN )) ( (resid 79 and name HD1* )) 4.30 2.50 2.50
assign ((resid 76 and name HN )) ( (resid 79 and name HD1* )) 3.38 1.59 1.59
assign ((resid 78 and name HN )) ( (resid 82 and name HD1* )) 4.19 2.40 2.40
assign ((resid 81 and name HG2* )) ( (resid 84 and name HN )) 4.39 2.59 2.59
assign ((resid 34 and name HB* )) ( (resid 86 and name HN )) 4.39 2.59 2.59
assign ((resid 89 and name HN )) ( (resid 107 and name HD1* )) 3.89 2.09 2.09
assign ((resid 90 and name HN )) ( (resid 91 and name HG2* )) 4.41 2.61 2.61
assign ((resid 103 and name HN )) ( (resid 104 and name HG2* )) 4.12 2.32 2.32
assign ((resid 104 and name HN )) ( (resid 107 and name HD2* )) 4.26 2.47 2.47
assign ((resid 104 and name HG1* )) ( (resid 105 and name HN )) 3.20 1.41 1.41
assign ((resid 82 and name HD2* )) ( (resid 110 and name HN )) 3.65 1.86 1.86
assign ((resid 82 and name HD2* )) ( (resid 113 and name HN )) 3.78 1.98 1.98
assign ((resid 37 and name HB* )) ( (resid 114 and name HN )) 4.41 2.61 2.61
assign ((resid 114 and name HN )) ( (resid 115 and name HG2* )) 3.54 1.75 1.75
assign ((resid 111 and name HG1* )) ( (resid 115 and name HN )) 4.32 2.52 2.52
assign ((resid 37 and name HB* )) ( (resid 115 and name HN )) 4.24 2.45 2.45
assign ((resid 41 and name HD1* )) ( (resid 118 and name HN )) 3.29 1.50 1.50
assign ((resid 118 and name HB* )) ( (resid 121 and name HN )) 4.41 2.61 2.61
assign ((resid 121 and name HN )) ( (resid 122 and name HD1* )) 3.62 1.82 1.82
assign ((resid 14 and name HB* )) ( (resid 29 and name HN )) 3.79 2.00 2.00
assign ((resid 39 and name HN )) ( (resid 40 and name HD1* )) 3.80 2.00 2.00
assign ((resid 37 and name HB* )) ( (resid 118 and name HN )) 3.83 2.04 2.04
assign ((resid 6 and name HN )) ( (resid 6 and name HB2 )) 2.84 1.04 1.04
assign ((resid 6 and name HN )) ( (resid 6 and name HB1 )) 2.84 1.04 1.04
assign ((resid 9 and name HN )) ( (resid 9 and name HB )) 2.44 0.64 0.64
assign ((resid 12 and name HN )) ( (resid 12 and name HB2 )) 2.48 0.68 0.68
assign ((resid 12 and name HN )) ( (resid 12 and name HB1 )) 2.48 0.68 0.68
assign ((resid 15 and name HN )) ( (resid 15 and name HB1 )) 2.87 1.08 1.08
assign ((resid 16 and name HA )) ( (resid 16 and name HB2 )) 2.33 0.54 0.54
assign ((resid 16 and name HA )) ( (resid 16 and name HB1 )) 2.33 0.54 0.54
assign ((resid 16 and name HN )) ( (resid 16 and name HB2 )) 2.55 0.75 0.75
assign ((resid 16 and name HN )) ( (resid 16 and name HB1 )) 2.55 0.75 0.75
assign ((resid 20 and name HN )) ( (resid 20 and name HB2 )) 2.64 0.84 0.84
assign ((resid 22 and name HN )) ( (resid 22 and name HB1 )) 2.60 0.81 0.81
assign ((resid 23 and name HN )) ( (resid 23 and name HB1 )) 2.73 0.93 0.93
assign ((resid 24 and name HN )) ( (resid 24 and name HB )) 2.31 0.52 0.52
assign ((resid 27 and name HN )) ( (resid 27 and name HB1 )) 2.80 1.00 1.00
assign ((resid 32 and name HN )) ( (resid 32 and name HB1 )) 2.76 0.97 0.97
assign ((resid 35 and name HN )) ( (resid 35 and name HB1 )) 2.87 1.08 1.08
assign ((resid 38 and name HN )) ( (resid 38 and name HB )) 2.49 0.70 0.70
assign ((resid 40 and name HN )) ( (resid 40 and name HB1 )) 2.62 0.82 0.82
assign ((resid 41 and name HN )) ( (resid 41 and name HB1 )) 2.93 1.13 1.13
assign ((resid 42 and name HN )) ( (resid 42 and name HB )) 2.51 0.72 0.72
assign ((resid 43 and name HN )) ( (resid 43 and name HB )) 2.40 0.61 0.61
assign ((resid 44 and name HN )) ( (resid 44 and name HB2 )) 2.82 1.02 1.02
assign ((resid 45 and name HN )) ( (resid 45 and name HB2 )) 2.67 0.88 0.88
assign ((resid 46 and name HN )) ( (resid 46 and name HA )) 2.21 0.41 0.41
assign ((resid 46 and name HN )) ( (resid 46 and name HB2 )) 2.71 0.91 0.91
assign ((resid 46 and name HA )) ( (resid 46 and name HB2 )) 2.40 0.61 0.61
assign ((resid 46 and name HN )) ( (resid 46 and name HB1 )) 2.71 0.91 0.91
assign ((resid 46 and name HA )) ( (resid 46 and name HB1 )) 2.40 0.61 0.61
assign ((resid 48 and name HN )) ( (resid 48 and name HB2 )) 2.53 0.73 0.73
assign ((resid 48 and name HN )) ( (resid 48 and name HB1 )) 2.53 0.73 0.73
assign ((resid 49 and name HA )) ( (resid 49 and name HB2 )) 2.37 0.57 0.57
assign ((resid 49 and name HA )) ( (resid 49 and name HB1 )) 2.37 0.57 0.57
assign ((resid 49 and name HN )) ( (resid 49 and name HB2 )) 2.64 0.84 0.84
assign ((resid 49 and name HN )) ( (resid 49 and name HB1 )) 2.64 0.84 0.84
assign ((resid 50 and name HN )) ( (resid 50 and name HB )) 2.49 0.70 0.70
assign ((resid 51 and name HN )) ( (resid 51 and name HB )) 2.78 0.99 0.99
assign ((resid 51 and name HA )) ( (resid 51 and name HB )) 2.35 0.55 0.55
assign ((resid 53 and name HN )) ( (resid 53 and name HB2 )) 2.58 0.79 0.79
assign ((resid 53 and name HN )) ( (resid 53 and name HB1 )) 2.58 0.79 0.79
assign ((resid 54 and name HN )) ( (resid 54 and name HB )) 2.49 0.70 0.70
assign ((resid 55 and name HN )) ( (resid 55 and name HB2 )) 2.53 0.73 0.73
assign ((resid 56 and name HN )) ( (resid 56 and name HB2 )) 2.53 0.73 0.73
assign ((resid 56 and name HN )) ( (resid 56 and name HB1 )) 2.53 0.73 0.73
assign ((resid 57 and name HN )) ( (resid 57 and name HA )) 2.35 0.55 0.55
assign ((resid 57 and name HN )) ( (resid 57 and name HB2 )) 2.51 0.72 0.72
assign ((resid 59 and name HN )) ( (resid 59 and name HB1 )) 2.64 0.84 0.84
assign ((resid 61 and name HN )) ( (resid 61 and name HB2 )) 2.94 1.15 1.15
assign ((resid 61 and name HN )) ( (resid 61 and name HB1 )) 2.94 1.15 1.15
assign ((resid 62 and name HA )) ( (resid 62 and name HB1 )) 2.37 0.57 0.57
assign ((resid 62 and name HN )) ( (resid 62 and name HB2 )) 2.85 1.06 1.06
assign ((resid 62 and name HA )) ( (resid 62 and name HB2 )) 2.37 0.57 0.57
assign ((resid 62 and name HN )) ( (resid 62 and name HB1 )) 2.85 1.06 1.06
assign ((resid 63 and name HN )) ( (resid 63 and name HB2 )) 2.66 0.86 0.86
assign ((resid 63 and name HN )) ( (resid 63 and name HB1 )) 2.66 0.86 0.86
assign ((resid 64 and name HN )) ( (resid 64 and name HB2 )) 2.58 0.79 0.79
assign ((resid 64 and name HN )) ( (resid 64 and name HB1 )) 2.58 0.79 0.79
assign ((resid 66 and name HN )) ( (resid 66 and name HB2 )) 2.84 1.04 1.04
assign ((resid 67 and name HN )) ( (resid 67 and name HB1 )) 2.53 0.73 0.73
assign ((resid 70 and name HN )) ( (resid 70 and name HB1 )) 2.60 0.81 0.81
assign ((resid 70 and name HN )) ( (resid 70 and name HB2 )) 2.53 0.73 0.73
assign ((resid 71 and name HN )) ( (resid 71 and name HB2 )) 2.84 1.04 1.04
assign ((resid 71 and name HN )) ( (resid 71 and name HB1 )) 2.84 1.04 1.04
assign ((resid 73 and name HN )) ( (resid 73 and name HB1 )) 2.53 0.73 0.73
assign ((resid 73 and name HN )) ( (resid 73 and name HB2 )) 2.53 0.73 0.73
assign ((resid 75 and name HN )) ( (resid 75 and name HB2 )) 2.57 0.77 0.77
assign ((resid 75 and name HN )) ( (resid 75 and name HB1 )) 2.57 0.77 0.77
assign ((resid 77 and name HN )) ( (resid 77 and name HB )) 2.64 0.84 0.84
assign ((resid 77 and name HA )) ( (resid 77 and name HB )) 2.37 0.57 0.57
assign ((resid 78 and name HA )) ( (resid 78 and name HB2 )) 2.39 0.59 0.59
assign ((resid 78 and name HA )) ( (resid 78 and name HB1 )) 2.35 0.55 0.55
assign ((resid 78 and name HN )) ( (resid 78 and name HB2 )) 2.40 0.61 0.61
assign ((resid 78 and name HN )) ( (resid 78 and name HB1 )) 2.62 0.82 0.82
assign ((resid 81 and name HN )) ( (resid 81 and name HB )) 2.37 0.57 0.57
assign ((resid 84 and name HN )) ( (resid 84 and name HB2 )) 2.44 0.64 0.64
assign ((resid 84 and name HN )) ( (resid 84 and name HB1 )) 2.44 0.64 0.64
assign ((resid 87 and name HA )) ( (resid 90 and name HN )) 2.89 1.09 1.09
assign ((resid 91 and name HN )) ( (resid 91 and name HB )) 2.55 0.75 0.75
assign ((resid 94 and name HN )) ( (resid 94 and name HB2 )) 2.75 0.95 0.95
assign ((resid 94 and name HN )) ( (resid 94 and name HB1 )) 2.75 0.95 0.95
assign ((resid 97 and name HN )) ( (resid 97 and name HB2 )) 2.80 1.00 1.00
assign ((resid 97 and name HN )) ( (resid 97 and name HB1 )) 2.80 1.00 1.00
assign ((resid 99 and name HN )) ( (resid 99 and name HB1 )) 2.94 1.15 1.15
assign ((resid 101 and name HN )) ( (resid 101 and name HB2 )) 2.64 0.84 0.84
assign ((resid 101 and name HN )) ( (resid 101 and name HB1 )) 2.64 0.84 0.84
assign ((resid 102 and name HN )) ( (resid 102 and name HB2 )) 2.40 0.61 0.61
assign ((resid 102 and name HA )) ( (resid 102 and name HB2 )) 2.40 0.61 0.61
assign ((resid 103 and name HN )) ( (resid 103 and name HB2 )) 2.60 0.81 0.81
assign ((resid 103 and name HN )) ( (resid 103 and name HB1 )) 2.60 0.81 0.81
assign ((resid 107 and name HN )) ( (resid 107 and name HB2 )) 2.84 1.04 1.04
assign ((resid 111 and name HN )) ( (resid 111 and name HB )) 2.64 0.84 0.84
assign ((resid 113 and name HN )) ( (resid 113 and name HB2 )) 2.71 0.91 0.91
assign ((resid 114 and name HA )) ( (resid 114 and name HB1 )) 2.39 0.59 0.59
assign ((resid 114 and name HN )) ( (resid 114 and name HB2 )) 2.51 0.72 0.72
assign ((resid 114 and name HN )) ( (resid 114 and name HB1 )) 2.53 0.73 0.73
assign ((resid 115 and name HN )) ( (resid 115 and name HB )) 2.31 0.52 0.52
assign ((resid 116 and name HN )) ( (resid 116 and name HB )) 2.39 0.59 0.59
assign ((resid 117 and name HN )) ( (resid 117 and name HB2 )) 2.48 0.68 0.68
assign ((resid 117 and name HN )) ( (resid 117 and name HB1 )) 2.51 0.72 0.72
assign ((resid 120 and name HN )) ( (resid 120 and name HB2 )) 2.48 0.68 0.68
assign ((resid 120 and name HN )) ( (resid 120 and name HB1 )) 2.48 0.68 0.68
assign ((resid 121 and name HN )) ( (resid 121 and name HB1 )) 2.62 0.82 0.82
assign ((resid 124 and name HA )) ( (resid 124 and name HB2 )) 2.40 0.61 0.61
assign ((resid 124 and name HA )) ( (resid 124 and name HB1 )) 2.40 0.61 0.61
assign ((resid 124 and name HN )) ( (resid 124 and name HB2 )) 2.73 0.93 0.93
assign ((resid 3 and name HA )) ( (resid 4 and name HN )) 2.12 0.32 0.32
assign ((resid 3 and name HB )) ( (resid 4 and name HN )) 2.51 0.72 0.72
assign ((resid 4 and name HA )) ( (resid 5 and name HN )) 2.26 0.46 0.46
assign ((resid 5 and name HA )) ( (resid 6 and name HN )) 2.30 0.50 0.50
assign ((resid 5 and name HB )) ( (resid 6 and name HN )) 2.57 0.77 0.77
assign ((resid 16 and name HB2 )) ( (resid 17 and name HN )) 2.71 0.91 0.91
assign ((resid 16 and name HB1 )) ( (resid 17 and name HN )) 2.71 0.91 0.91
assign ((resid 18 and name HA )) ( (resid 19 and name HN )) 2.66 0.86 0.86
assign ((resid 22 and name HA )) ( (resid 23 and name HN )) 2.30 0.50 0.50
assign ((resid 23 and name HB1 )) ( (resid 24 and name HN )) 2.96 1.17 1.17
assign ((resid 24 and name HB )) ( (resid 25 and name HN )) 2.49 0.70 0.70
assign ((resid 32 and name HB1 )) ( (resid 33 and name HN )) 2.96 1.17 1.17
assign ((resid 35 and name HB2 )) ( (resid 36 and name HN )) 2.85 1.06 1.06
assign ((resid 38 and name HB )) ( (resid 39 and name HN )) 2.85 1.06 1.06
assign ((resid 40 and name HB2 )) ( (resid 41 and name HN )) 2.62 0.82 0.82
assign ((resid 40 and name HB1 )) ( (resid 41 and name HN )) 2.75 0.95 0.95
assign ((resid 42 and name HB )) ( (resid 43 and name HN )) 2.66 0.86 0.86
assign ((resid 43 and name HB )) ( (resid 44 and name HN )) 2.58 0.79 0.79
assign ((resid 46 and name HA )) ( (resid 47 and name HN )) 2.44 0.64 0.64
assign ((resid 48 and name HB2 )) ( (resid 49 and name HN )) 2.91 1.11 1.11
assign ((resid 50 and name HB )) ( (resid 51 and name HN )) 2.62 0.82 0.82
assign ((resid 54 and name HA )) ( (resid 55 and name HN )) 2.62 0.82 0.82
assign ((resid 54 and name HB )) ( (resid 55 and name HN )) 2.80 1.00 1.00
assign ((resid 56 and name HA )) ( (resid 57 and name HN )) 2.64 0.84 0.84
assign ((resid 56 and name HB2 )) ( (resid 57 and name HN )) 2.82 1.02 1.02
assign ((resid 56 and name HB1 )) ( (resid 57 and name HN )) 2.82 1.02 1.02
assign ((resid 57 and name HA )) ( (resid 58 and name HN )) 2.49 0.70 0.70
assign ((resid 58 and name HA2 )) ( (resid 59 and name HN )) 2.60 0.81 0.81
assign ((resid 58 and name HA1 )) ( (resid 59 and name HN )) 2.60 0.81 0.81
assign ((resid 59 and name HA )) ( (resid 60 and name HN )) 2.31 0.52 0.52
assign ((resid 59 and name HB2 )) ( (resid 60 and name HN )) 2.98 1.18 1.18
assign ((resid 59 and name HB1 )) ( (resid 60 and name HN )) 2.98 1.18 1.18
assign ((resid 60 and name HA )) ( (resid 61 and name HN )) 2.40 0.61 0.61
assign ((resid 63 and name HB2 )) ( (resid 64 and name HN )) 3.02 1.22 1.22
assign ((resid 63 and name HB1 )) ( (resid 64 and name HN )) 3.02 1.22 1.22
assign ((resid 67 and name HB1 )) ( (resid 68 and name HN )) 2.42 0.63 0.63
assign ((resid 67 and name HB2 )) ( (resid 68 and name HN )) 2.53 0.73 0.73
assign ((resid 70 and name HA )) ( (resid 71 and name HN )) 2.67 0.88 0.88
assign ((resid 70 and name HB1 )) ( (resid 71 and name HN )) 2.60 0.81 0.81
assign ((resid 70 and name HB2 )) ( (resid 71 and name HN )) 2.58 0.79 0.79
assign ((resid 73 and name HB1 )) ( (resid 74 and name HN )) 2.93 1.13 1.13
assign ((resid 75 and name HB2 )) ( (resid 76 and name HN )) 2.75 0.95 0.95
assign ((resid 75 and name HB1 )) ( (resid 76 and name HN )) 2.75 0.95 0.95
assign ((resid 76 and name HA )) ( (resid 77 and name HN )) 2.30 0.50 0.50
assign ((resid 77 and name HB )) ( (resid 78 and name HN )) 3.09 1.29 1.29
assign ((resid 80 and name HB1 )) ( (resid 81 and name HN )) 2.98 1.18 1.18
assign ((resid 80 and name HB2 )) ( (resid 81 and name HN )) 2.73 0.93 0.93
assign ((resid 81 and name HB )) ( (resid 82 and name HN )) 2.58 0.79 0.79
assign ((resid 84 and name HB2 )) ( (resid 85 and name HN )) 2.76 0.97 0.97
assign ((resid 84 and name HB1 )) ( (resid 85 and name HN )) 2.76 0.97 0.97
assign ((resid 91 and name HB )) ( (resid 92 and name HN )) 2.69 0.90 0.90
assign ((resid 94 and name HB2 )) ( (resid 95 and name HN )) 3.07 1.27 1.27
assign ((resid 94 and name HB1 )) ( (resid 95 and name HN )) 3.07 1.27 1.27
assign ((resid 97 and name HA )) ( (resid 98 and name HN )) 2.46 0.66 0.66
assign ((resid 99 and name HA )) ( (resid 100 and name HN )) 2.30 0.50 0.50
assign ((resid 100 and name HB2 )) ( (resid 101 and name HN )) 2.71 0.91 0.91
assign ((resid 100 and name HB1 )) ( (resid 101 and name HN )) 2.71 0.91 0.91
assign ((resid 101 and name HB2 )) ( (resid 102 and name HN )) 2.76 0.97 0.97
assign ((resid 101 and name HB1 )) ( (resid 102 and name HN )) 2.76 0.97 0.97
assign ((resid 102 and name HB2 )) ( (resid 103 and name HN )) 2.71 0.91 0.91
assign ((resid 102 and name HB1 )) ( (resid 103 and name HN )) 2.66 0.86 0.86
assign ((resid 103 and name HB2 )) ( (resid 104 and name HN )) 2.89 1.09 1.09
assign ((resid 103 and name HB1 )) ( (resid 104 and name HN )) 2.89 1.09 1.09
assign ((resid 114 and name HB2 )) ( (resid 115 and name HN )) 2.78 0.99 0.99
assign ((resid 117 and name HB2 )) ( (resid 118 and name HN )) 2.75 0.95 0.95
assign ((resid 117 and name HB1 )) ( (resid 118 and name HN )) 2.62 0.82 0.82
assign ((resid 122 and name HA )) ( (resid 123 and name HN )) 2.60 0.81 0.81
assign ((resid 6 and name HA )) ( (resid 9 and name HB )) 2.57 0.77 0.77
assign ((resid 7 and name HA )) ( (resid 10 and name HB1 )) 3.36 1.56 1.56
assign ((resid 8 and name HA )) ( (resid 11 and name HN )) 2.75 0.95 0.95
assign ((resid 8 and name HA )) ( (resid 11 and name HB2 )) 2.89 1.09 1.09
assign ((resid 8 and name HA )) ( (resid 11 and name HB1 )) 2.69 0.90 0.90
assign ((resid 9 and name HA )) ( (resid 12 and name HN )) 2.87 1.08 1.08
assign ((resid 9 and name HA )) ( (resid 12 and name HB2 )) 2.80 1.00 1.00
assign ((resid 9 and name HA )) ( (resid 12 and name HB1 )) 2.80 1.00 1.00
assign ((resid 11 and name HA )) ( (resid 14 and name HN )) 3.14 1.35 1.35
assign ((resid 12 and name HA )) ( (resid 15 and name HN )) 2.78 0.99 0.99
assign ((resid 12 and name HA )) ( (resid 15 and name HB2 )) 2.58 0.79 0.79
assign ((resid 12 and name HA )) ( (resid 15 and name HB1 )) 2.93 1.13 1.13
assign ((resid 13 and name HA )) ( (resid 16 and name HB2 )) 2.67 0.88 0.88
assign ((resid 13 and name HA )) ( (resid 16 and name HB1 )) 2.67 0.88 0.88
assign ((resid 14 and name HA )) ( (resid 17 and name HB2 )) 3.21 1.42 1.42
assign ((resid 15 and name HA )) ( (resid 18 and name HN )) 2.91 1.11 1.11
assign ((resid 15 and name HA )) ( (resid 18 and name HB2 )) 2.80 1.00 1.00
assign ((resid 15 and name HA )) ( (resid 18 and name HB1 )) 2.80 1.00 1.00
assign ((resid 17 and name HA )) ( (resid 19 and name HN )) 2.91 1.11 1.11
assign ((resid 27 and name HA )) ( (resid 30 and name HN )) 2.93 1.13 1.13
assign ((resid 27 and name HA )) ( (resid 30 and name HB* )) 3.49 1.70 1.70
assign ((resid 30 and name HA )) ( (resid 33 and name HN )) 2.94 1.15 1.15
assign ((resid 31 and name HA )) ( (resid 34 and name HN )) 3.02 1.22 1.22
assign ((resid 37 and name HA )) ( (resid 40 and name HN )) 2.91 1.11 1.11
assign ((resid 37 and name HA )) ( (resid 40 and name HB2 )) 2.66 0.86 0.86
assign ((resid 37 and name HA )) ( (resid 40 and name HB1 )) 2.85 1.06 1.06
assign ((resid 39 and name HA )) ( (resid 42 and name HB )) 2.82 1.02 1.02
assign ((resid 40 and name HA )) ( (resid 43 and name HB )) 2.67 0.88 0.88
assign ((resid 64 and name HA )) ( (resid 67 and name HB1 )) 2.80 1.00 1.00
assign ((resid 64 and name HA )) ( (resid 67 and name HB2 )) 2.93 1.13 1.13
assign ((resid 67 and name HA )) ( (resid 70 and name HB1 )) 2.91 1.11 1.11
assign ((resid 67 and name HA )) ( (resid 70 and name HB2 )) 2.78 0.99 0.99
assign ((resid 77 and name HA )) ( (resid 79 and name HN )) 3.52 1.72 1.72
assign ((resid 78 and name HA )) ( (resid 81 and name HN )) 2.85 1.06 1.06
assign ((resid 78 and name HA )) ( (resid 81 and name HB )) 2.67 0.88 0.88
assign ((resid 80 and name HA )) ( (resid 83 and name HB* )) 3.22 1.43 1.43
assign ((resid 81 and name HA )) ( (resid 84 and name HB1 )) 2.58 0.79 0.79
assign ((resid 85 and name HA )) ( (resid 88 and name HB )) 2.58 0.79 0.79
assign ((resid 86 and name HA )) ( (resid 89 and name HB2 )) 3.25 1.45 1.45
assign ((resid 88 and name HA )) ( (resid 91 and name HN )) 3.03 1.24 1.24
assign ((resid 88 and name HA )) ( (resid 91 and name HB )) 2.94 1.15 1.15
assign ((resid 88 and name HA )) ( (resid 92 and name HN )) 3.11 1.31 1.31
assign ((resid 101 and name HA )) ( (resid 104 and name HB )) 2.51 0.72 0.72
assign ((resid 102 and name HA )) ( (resid 105 and name HB* )) 3.29 1.50 1.50
assign ((resid 103 and name HA )) ( (resid 106 and name HN )) 2.76 0.97 0.97
assign ((resid 103 and name HA )) ( (resid 106 and name HB* )) 3.11 1.32 1.32
assign ((resid 105 and name HA )) ( (resid 108 and name HN )) 2.89 1.09 1.09
assign ((resid 106 and name HA )) ( (resid 109 and name HB* )) 3.04 1.25 1.25
assign ((resid 107 and name HA )) ( (resid 110 and name HB2 )) 2.73 0.93 0.93
assign ((resid 107 and name HA )) ( (resid 110 and name HB1 )) 2.87 1.08 1.08
assign ((resid 108 and name HA )) ( (resid 111 and name HB )) 2.96 1.17 1.17
assign ((resid 109 and name HA )) ( (resid 111 and name HN )) 3.39 1.60 1.60
assign ((resid 109 and name HA )) ( (resid 112 and name HN )) 2.69 0.90 0.90
assign ((resid 109 and name HA )) ( (resid 112 and name HB2 )) 2.62 0.82 0.82
assign ((resid 109 and name HA )) ( (resid 112 and name HB1 )) 2.62 0.82 0.82
assign ((resid 110 and name HA )) ( (resid 113 and name HN )) 2.71 0.91 0.91
assign ((resid 110 and name HA )) ( (resid 113 and name HB2 )) 2.67 0.88 0.88
assign ((resid 110 and name HA )) ( (resid 113 and name HB1 )) 3.18 1.38 1.38
assign ((resid 111 and name HA )) ( (resid 114 and name HN )) 2.80 1.00 1.00
assign ((resid 111 and name HA )) ( (resid 114 and name HB2 )) 2.66 0.86 0.86
assign ((resid 111 and name HA )) ( (resid 114 and name HB1 )) 2.91 1.11 1.11
assign ((resid 112 and name HA )) ( (resid 115 and name HB )) 2.98 1.18 1.18
assign ((resid 114 and name HA )) ( (resid 117 and name HN )) 2.71 0.91 0.91
assign ((resid 114 and name HA )) ( (resid 117 and name HB2 )) 2.87 1.08 1.08
assign ((resid 114 and name HA )) ( (resid 117 and name HB1 )) 2.85 1.06 1.06
assign ((resid 115 and name HA )) ( (resid 117 and name HN )) 2.87 1.08 1.08
assign ((resid 116 and name HA )) ( (resid 119 and name HB )) 2.46 0.66 0.66
assign ((resid 117 and name HA )) ( (resid 120 and name HB2 )) 2.98 1.18 1.18
assign ((resid 117 and name HA )) ( (resid 120 and name HB1 )) 2.98 1.18 1.18
assign ((resid 118 and name HA )) ( (resid 121 and name HB2 )) 2.80 1.00 1.00
assign ((resid 118 and name HA )) ( (resid 121 and name HB1 )) 2.80 1.00 1.00
assign ((resid 119 and name HA )) ( (resid 122 and name HB* )) 3.15 1.36 1.36
assign ((resid 5 and name HB )) ( (resid 7 and name HN )) 2.58 0.79 0.79
assign ((resid 17 and name HA )) ( (resid 20 and name HN )) 2.96 1.17 1.17
assign ((resid 17 and name HA )) ( (resid 20 and name HB1 )) 2.93 1.13 1.13
assign ((resid 17 and name HA )) ( (resid 20 and name HB2 )) 2.93 1.13 1.13
assign ((resid 14 and name HA )) ( (resid 26 and name HA )) 2.49 0.70 0.70
assign ((resid 49 and name HA )) ( (resid 52 and name HB* )) 3.51 1.72 1.72
assign ((resid 50 and name HA )) ( (resid 53 and name HB2 )) 3.14 1.35 1.35
assign ((resid 50 and name HA )) ( (resid 53 and name HB1 )) 3.14 1.35 1.35
assign ((resid 51 and name HA )) ( (resid 54 and name HB )) 2.85 1.06 1.06
assign ((resid 48 and name HB1 )) ( (resid 49 and name HN )) 2.91 1.11 1.11
assign ((resid 101 and name HA )) ( (resid 104 and name HN )) 2.62 0.82 0.82
assign ((resid 109 and name HB* )) ( (resid 110 and name HA )) 3.53 1.73 1.73
assign ((resid 73 and name HB2 )) ( (resid 77 and name HA )) 2.89 1.09 1.09
assign ((resid 48 and name HA )) ( (resid 51 and name HN )) 2.71 0.91 0.91
assign ((resid 120 and name HA )) ( (resid 123 and name HB* )) 3.33 1.53 1.53
assign ((resid 54 and name HA )) ( (resid 56 and name HN )) 2.62 0.82 0.82
assign ((resid 81 and name HA )) ( (resid 84 and name HB2 )) 2.58 0.79 0.79
assign ((resid 71 and name HB1 )) ( (resid 72 and name HN )) 2.96 1.17 1.17
assign ((resid 3 and name HA )) ( (resid 3 and name HG11 )) 2.89 1.09 1.09
assign ((resid 3 and name HA )) ( (resid 3 and name HG12 )) 2.89 1.09 1.09
assign ((resid 8 and name HA )) ( (resid 8 and name HG2 )) 2.91 1.11 1.11
assign ((resid 8 and name HA )) ( (resid 8 and name HG1 )) 2.91 1.11 1.11
assign ((resid 12 and name HN )) ( (resid 12 and name HG* )) 3.40 1.61 1.61
assign ((resid 13 and name HA )) ( (resid 13 and name HG2 )) 2.84 1.04 1.04
assign ((resid 13 and name HN )) ( (resid 13 and name HG2 )) 2.76 0.97 0.97
assign ((resid 13 and name HA )) ( (resid 13 and name HG1 )) 2.84 1.04 1.04
assign ((resid 15 and name HA )) ( (resid 15 and name HG2 )) 3.02 1.22 1.22
assign ((resid 15 and name HA )) ( (resid 15 and name HG1 )) 3.02 1.22 1.22
assign ((resid 16 and name HN )) ( (resid 16 and name HG1 )) 2.94 1.15 1.15
assign ((resid 20 and name HA )) ( (resid 20 and name HD* )) 4.18 2.38 2.38
assign ((resid 23 and name HN )) ( (resid 23 and name HG )) 2.39 0.59 0.59
assign ((resid 25 and name HN )) ( (resid 25 and name HG2 )) 2.64 0.84 0.84
assign ((resid 25 and name HA )) ( (resid 25 and name HG2 )) 2.58 0.79 0.79
assign ((resid 25 and name HN )) ( (resid 25 and name HG1 )) 2.64 0.84 0.84
assign ((resid 25 and name HA )) ( (resid 25 and name HG1 )) 2.58 0.79 0.79
assign ((resid 32 and name HN )) ( (resid 32 and name HG1 )) 3.05 1.26 1.26
assign ((resid 32 and name HA )) ( (resid 32 and name HD2 )) 3.25 1.45 1.45
assign ((resid 32 and name HA )) ( (resid 32 and name HD1 )) 3.25 1.45 1.45
assign ((resid 35 and name HA )) ( (resid 35 and name HG2 )) 2.85 1.06 1.06
assign ((resid 35 and name HA )) ( (resid 35 and name HG1 )) 2.85 1.06 1.06
assign ((resid 38 and name HA )) ( (resid 38 and name HG12 )) 2.98 1.18 1.18
assign ((resid 38 and name HA )) ( (resid 38 and name HG11 )) 2.98 1.18 1.18
assign ((resid 40 and name HB1 )) ( (resid 40 and name HG )) 2.35 0.55 0.55
assign ((resid 40 and name HA )) ( (resid 40 and name HG )) 3.02 1.22 1.22
assign ((resid 40 and name HB2 )) ( (resid 40 and name HG )) 2.35 0.55 0.55
assign ((resid 41 and name HN )) ( (resid 41 and name HG )) 2.96 1.17 1.17
assign ((resid 43 and name HA )) ( (resid 43 and name HG12 )) 2.53 0.73 0.73
assign ((resid 43 and name HA )) ( (resid 43 and name HG11 )) 2.89 1.09 1.09
assign ((resid 43 and name HN )) ( (resid 43 and name HG12 )) 2.82 1.02 1.02
assign ((resid 43 and name HN )) ( (resid 43 and name HG11 )) 2.58 0.79 0.79
assign ((resid 44 and name HA )) ( (resid 44 and name HG1 )) 2.82 1.02 1.02
assign ((resid 44 and name HN )) ( (resid 44 and name HG1 )) 3.00 1.20 1.20
assign ((resid 47 and name HB1 )) ( (resid 47 and name HG )) 2.40 0.61 0.61
assign ((resid 48 and name HA )) ( (resid 48 and name HG2 )) 2.78 0.99 0.99
assign ((resid 48 and name HA )) ( (resid 48 and name HG1 )) 2.78 0.99 0.99
assign ((resid 48 and name HA )) ( (resid 48 and name HD* )) 3.56 1.77 1.77
assign ((resid 49 and name HN )) ( (resid 49 and name HG2 )) 3.18 1.38 1.38
assign ((resid 49 and name HA )) ( (resid 49 and name HG2 )) 2.80 1.00 1.00
assign ((resid 49 and name HN )) ( (resid 49 and name HG1 )) 3.18 1.38 1.38
assign ((resid 49 and name HA )) ( (resid 49 and name HG1 )) 2.80 1.00 1.00
assign ((resid 50 and name HN )) ( (resid 50 and name HG12 )) 3.16 1.36 1.36
assign ((resid 55 and name HA )) ( (resid 55 and name HG2 )) 2.73 0.93 0.93
assign ((resid 55 and name HA )) ( (resid 55 and name HG1 )) 2.73 0.93 0.93
assign ((resid 57 and name HA )) ( (resid 57 and name HG1 )) 2.80 1.00 1.00
assign ((resid 57 and name HN )) ( (resid 57 and name HG2 )) 2.91 1.11 1.11
assign ((resid 57 and name HA )) ( (resid 57 and name HG2 )) 2.80 1.00 1.00
assign ((resid 57 and name HN )) ( (resid 57 and name HG1 )) 2.91 1.11 1.11
assign ((resid 57 and name HA )) ( (resid 57 and name HD* )) 3.38 1.59 1.59
assign ((resid 59 and name HA )) ( (resid 59 and name HD* )) 3.83 2.04 2.04
assign ((resid 63 and name HN )) ( (resid 63 and name HG* )) 3.40 1.61 1.61
assign ((resid 65 and name HA )) ( (resid 65 and name HG )) 2.93 1.13 1.13
assign ((resid 67 and name HN )) ( (resid 67 and name HG2 )) 3.34 1.54 1.54
assign ((resid 67 and name HA )) ( (resid 67 and name HG2 )) 2.76 0.97 0.97
assign ((resid 67 and name HA )) ( (resid 67 and name HG1 )) 2.76 0.97 0.97
assign ((resid 70 and name HN )) ( (resid 70 and name HG2 )) 3.25 1.45 1.45
assign ((resid 70 and name HA )) ( (resid 70 and name HG2 )) 2.85 1.06 1.06
assign ((resid 70 and name HA )) ( (resid 70 and name HG1 )) 2.85 1.06 1.06
assign ((resid 70 and name HA )) ( (resid 70 and name HD* )) 3.46 1.66 1.66
assign ((resid 71 and name HA )) ( (resid 71 and name HG )) 2.71 0.91 0.91
assign ((resid 72 and name HA )) ( (resid 72 and name HG )) 2.75 0.95 0.95
assign ((resid 73 and name HA )) ( (resid 73 and name HG2 )) 2.75 0.95 0.95
assign ((resid 73 and name HA )) ( (resid 73 and name HG1 )) 2.75 0.95 0.95
assign ((resid 73 and name HN )) ( (resid 73 and name HG2 )) 3.29 1.49 1.49
assign ((resid 73 and name HN )) ( (resid 73 and name HD* )) 3.62 1.82 1.82
assign ((resid 75 and name HA )) ( (resid 75 and name HD21 )) 3.50 1.71 1.71
assign ((resid 78 and name HN )) ( (resid 78 and name HG2 )) 2.69 0.90 0.90
assign ((resid 78 and name HA )) ( (resid 78 and name HG2 )) 2.53 0.73 0.73
assign ((resid 78 and name HN )) ( (resid 78 and name HG1 )) 2.69 0.90 0.90
assign ((resid 78 and name HA )) ( (resid 78 and name HG1 )) 2.53 0.73 0.73
assign ((resid 81 and name HA )) ( (resid 81 and name HG12 )) 2.87 1.08 1.08
assign ((resid 81 and name HN )) ( (resid 81 and name HG11 )) 2.98 1.18 1.18
assign ((resid 81 and name HA )) ( (resid 81 and name HG11 )) 2.87 1.08 1.08
assign ((resid 82 and name HA )) ( (resid 82 and name HG )) 2.85 1.06 1.06
assign ((resid 84 and name HA )) ( (resid 84 and name HG2 )) 2.93 1.13 1.13
assign ((resid 84 and name HA )) ( (resid 84 and name HG1 )) 2.93 1.13 1.13
assign ((resid 89 and name HN )) ( (resid 89 and name HG )) 3.07 1.27 1.27
assign ((resid 97 and name HA )) ( (resid 97 and name HG )) 2.75 0.95 0.95
assign ((resid 97 and name HN )) ( (resid 97 and name HG )) 3.00 1.20 1.20
assign ((resid 97 and name HB2 )) ( (resid 97 and name HG )) 2.37 0.57 0.57
assign ((resid 97 and name HB1 )) ( (resid 97 and name HG )) 2.37 0.57 0.57
assign ((resid 101 and name HN )) ( (resid 101 and name HG* )) 3.64 1.84 1.84
assign ((resid 102 and name HN )) ( (resid 102 and name HG2 )) 3.03 1.24 1.24
assign ((resid 102 and name HA )) ( (resid 102 and name HG2 )) 2.78 0.99 0.99
assign ((resid 102 and name HA )) ( (resid 102 and name HG1 )) 2.78 0.99 0.99
assign ((resid 105 and name HA )) ( (resid 105 and name HG2 )) 2.96 1.17 1.17
assign ((resid 105 and name HA )) ( (resid 105 and name HG1 )) 2.96 1.17 1.17
assign ((resid 105 and name HN )) ( (resid 105 and name HG2 )) 3.21 1.42 1.42
assign ((resid 105 and name HA )) ( (resid 105 and name HD* )) 4.19 2.40 2.40
assign ((resid 106 and name HA )) ( (resid 106 and name HG1 )) 2.57 0.77 0.77
assign ((resid 106 and name HA )) ( (resid 106 and name HG2 )) 2.57 0.77 0.77
assign ((resid 113 and name HA )) ( (resid 113 and name HG2 )) 2.91 1.11 1.11
assign ((resid 113 and name HA )) ( (resid 113 and name HG1 )) 2.91 1.11 1.11
assign ((resid 114 and name HB1 )) ( (resid 114 and name HG )) 2.19 0.39 0.39
assign ((resid 116 and name HA )) ( (resid 116 and name HG11 )) 2.80 1.00 1.00
assign ((resid 116 and name HN )) ( (resid 116 and name HG12 )) 2.75 0.95 0.95
assign ((resid 116 and name HA )) ( (resid 116 and name HG12 )) 2.80 1.00 1.00
assign ((resid 116 and name HN )) ( (resid 116 and name HG11 )) 2.75 0.95 0.95
assign ((resid 123 and name HA )) ( (resid 123 and name HG2 )) 2.58 0.79 0.79
assign ((resid 123 and name HN )) ( (resid 123 and name HG2 )) 2.85 1.06 1.06
assign ((resid 123 and name HN )) ( (resid 123 and name HG1 )) 2.85 1.06 1.06
assign ((resid 123 and name HA )) ( (resid 123 and name HG1 )) 2.58 0.79 0.79
assign ((resid 125 and name HA )) ( (resid 125 and name HD2 )) 3.41 1.62 1.62
assign ((resid 3 and name HG12 )) ( (resid 4 and name HN )) 3.07 1.27 1.27
assign ((resid 3 and name HG11 )) ( (resid 4 and name HN )) 3.07 1.27 1.27
assign ((resid 63 and name HG* )) ( (resid 64 and name HN )) 3.51 1.72 1.72
assign ((resid 70 and name HD* )) ( (resid 71 and name HN )) 3.80 2.00 2.00
assign ((resid 73 and name HD* )) ( (resid 74 and name HN )) 3.49 1.70 1.70
assign ((resid 99 and name HG )) ( (resid 100 and name HN )) 3.57 1.78 1.78
assign ((resid 101 and name HG* )) ( (resid 102 and name HN )) 3.71 1.91 1.91
assign ((resid 103 and name HG* )) ( (resid 104 and name HN )) 4.34 2.54 2.54
assign ((resid 17 and name HA )) ( (resid 20 and name HG* )) 3.60 1.81 1.81
assign ((resid 17 and name HB2 )) ( (resid 26 and name HA )) 2.96 1.17 1.17
assign ((resid 27 and name HA )) ( (resid 30 and name HD* )) 4.03 2.23 2.23
assign ((resid 10 and name HB1 )) ( (resid 33 and name HA )) 3.36 1.56 1.56
assign ((resid 10 and name HB2 )) ( (resid 33 and name HA )) 2.73 0.93 0.93
assign ((resid 41 and name HA )) ( (resid 45 and name HD22 )) 3.68 1.89 1.89
assign ((resid 42 and name HA )) ( (resid 47 and name HB2 )) 2.85 1.06 1.06
assign ((resid 42 and name HA )) ( (resid 47 and name HG )) 3.23 1.44 1.44
assign ((resid 42 and name HA )) ( (resid 47 and name HB1 )) 3.90 2.10 2.10
assign ((resid 73 and name HG1 )) ( (resid 77 and name HA )) 2.69 0.90 0.90
assign ((resid 73 and name HG2 )) ( (resid 77 and name HA )) 2.69 0.90 0.90
assign ((resid 79 and name HB )) ( (resid 80 and name HD2 )) 2.82 1.02 1.02
assign ((resid 79 and name HB )) ( (resid 80 and name HD1 )) 2.82 1.02 1.02
assign ((resid 86 and name HA )) ( (resid 89 and name HG )) 2.62 0.82 0.82
assign ((resid 89 and name HG )) ( (resid 107 and name HG )) 2.37 0.57 0.57
assign ((resid 44 and name HA )) ( (resid 44 and name HG2 )) 2.82 1.02 1.02
assign ((resid 101 and name HG* )) ( (resid 105 and name HD* )) 4.28 2.48 2.48
assign ((resid 105 and name HN )) ( (resid 105 and name HD* )) 3.73 1.93 1.93
assign ((resid 69 and name HB1 )) ( (resid 80 and name HA )) 3.11 1.31 1.31
assign ((resid 69 and name HB2 )) ( (resid 80 and name HA )) 3.11 1.31 1.31
assign ((resid 82 and name HA )) ( (resid 110 and name HB1 )) 2.98 1.18 1.18
assign ((resid 41 and name HG )) ( (resid 118 and name HA )) 2.93 1.13 1.13
assign ((resid 3 and name HG2* )) ( (resid 3 and name HD1* )) 3.17 1.38 1.38
assign ((resid 5 and name HN )) ( (resid 5 and name HG2* )) 3.13 1.33 1.33
assign ((resid 9 and name HN )) ( (resid 9 and name HG2* )) 2.81 1.01 1.01
assign ((resid 18 and name HA )) ( (resid 18 and name HD2* )) 3.38 1.59 1.59
assign ((resid 18 and name HN )) ( (resid 18 and name HD1* )) 3.70 1.91 1.91
assign ((resid 18 and name HA )) ( (resid 18 and name HD1* )) 2.75 0.96 0.96
assign ((resid 23 and name HA )) ( (resid 23 and name HD1* )) 3.40 1.60 1.60
assign ((resid 23 and name HA )) ( (resid 23 and name HD2* )) 2.79 0.99 0.99
assign ((resid 24 and name HN )) ( (resid 24 and name HG1* )) 3.33 1.53 1.53
assign ((resid 24 and name HN )) ( (resid 24 and name HG2* )) 2.77 0.97 0.97
assign ((resid 24 and name HA )) ( (resid 24 and name HG2* )) 2.77 0.97 0.97
assign ((resid 38 and name HA )) ( (resid 38 and name HD1* )) 2.81 1.01 1.01
assign ((resid 40 and name HA )) ( (resid 40 and name HD1* )) 3.00 1.21 1.21
assign ((resid 40 and name HA )) ( (resid 40 and name HD2* )) 3.07 1.28 1.28
assign ((resid 40 and name HN )) ( (resid 40 and name HD2* )) 3.45 1.66 1.66
assign ((resid 41 and name HA )) ( (resid 41 and name HD1* )) 2.84 1.05 1.05
assign ((resid 41 and name HA )) ( (resid 41 and name HD2* )) 3.16 1.37 1.37
assign ((resid 41 and name HN )) ( (resid 41 and name HD1* )) 3.53 1.73 1.73
assign ((resid 41 and name HN )) ( (resid 41 and name HD2* )) 3.63 1.84 1.84
assign ((resid 42 and name HN )) ( (resid 42 and name HG2* )) 2.95 1.15 1.15
assign ((resid 43 and name HN )) ( (resid 43 and name HG2* )) 3.29 1.50 1.50
assign ((resid 43 and name HN )) ( (resid 43 and name HD1* )) 3.36 1.57 1.57
assign ((resid 43 and name HA )) ( (resid 43 and name HD1* )) 2.93 1.14 1.14
assign ((resid 47 and name HA )) ( (resid 47 and name HD1* )) 3.17 1.37 1.37
assign ((resid 47 and name HA )) ( (resid 47 and name HD2* )) 3.06 1.26 1.26
assign ((resid 50 and name HG2* )) ( (resid 50 and name HD1* )) 3.12 1.32 1.32
assign ((resid 50 and name HN )) ( (resid 50 and name HD1* )) 3.49 1.69 1.69
assign ((resid 50 and name HA )) ( (resid 50 and name HD1* )) 2.81 1.01 1.01
assign ((resid 51 and name HN )) ( (resid 51 and name HG2* )) 2.86 1.06 1.06
assign ((resid 54 and name HN )) ( (resid 54 and name HG2* )) 3.04 1.24 1.24
assign ((resid 65 and name HA )) ( (resid 65 and name HD2* )) 2.93 1.14 1.14
assign ((resid 71 and name HN )) ( (resid 71 and name HD1* )) 3.76 1.96 1.96
assign ((resid 71 and name HN )) ( (resid 71 and name HD2* )) 3.47 1.68 1.68
assign ((resid 71 and name HA )) ( (resid 71 and name HD2* )) 2.64 0.85 0.85
assign ((resid 72 and name HA )) ( (resid 72 and name HD2* )) 2.90 1.10 1.10
assign ((resid 72 and name HN )) ( (resid 72 and name HD1* )) 3.29 1.50 1.50
assign ((resid 72 and name HA )) ( (resid 72 and name HD1* )) 3.81 2.02 2.02
assign ((resid 77 and name HN )) ( (resid 77 and name HG2* )) 2.88 1.08 1.08
assign ((resid 79 and name HA )) ( (resid 79 and name HD1* )) 3.26 1.46 1.46
assign ((resid 81 and name HN )) ( (resid 81 and name HG2* )) 3.34 1.55 1.55
assign ((resid 81 and name HA )) ( (resid 81 and name HD1* )) 3.38 1.59 1.59
assign ((resid 82 and name HN )) ( (resid 82 and name HD2* )) 3.40 1.60 1.60
assign ((resid 82 and name HA )) ( (resid 82 and name HD2* )) 2.86 1.06 1.06
assign ((resid 91 and name HN )) ( (resid 91 and name HG2* )) 2.99 1.19 1.19
assign ((resid 97 and name HA )) ( (resid 97 and name HD1* )) 3.18 1.39 1.39
assign ((resid 97 and name HN )) ( (resid 97 and name HD1* )) 3.58 1.78 1.78
assign ((resid 97 and name HN )) ( (resid 97 and name HD2* )) 3.67 1.87 1.87
assign ((resid 97 and name HA )) ( (resid 97 and name HD2* )) 2.79 0.99 0.99
assign ((resid 99 and name HN )) ( (resid 99 and name HD1* )) 3.89 2.09 2.09
assign ((resid 99 and name HA )) ( (resid 99 and name HD1* )) 3.69 1.89 1.89
assign ((resid 99 and name HA )) ( (resid 99 and name HD2* )) 2.86 1.06 1.06
assign ((resid 107 and name HA )) ( (resid 107 and name HD1* )) 3.78 1.98 1.98
assign ((resid 107 and name HA )) ( (resid 107 and name HD2* )) 2.79 0.99 0.99
assign ((resid 107 and name HN )) ( (resid 107 and name HD2* )) 3.54 1.75 1.75
assign ((resid 111 and name HN )) ( (resid 111 and name HG2* )) 2.97 1.17 1.17
assign ((resid 114 and name HA )) ( (resid 114 and name HD2* )) 3.04 1.24 1.24
assign ((resid 114 and name HA )) ( (resid 114 and name HD1* )) 2.88 1.08 1.08
assign ((resid 115 and name HN )) ( (resid 115 and name HG2* )) 2.84 1.05 1.05
assign ((resid 115 and name HN )) ( (resid 115 and name HG1* )) 3.15 1.35 1.35
assign ((resid 116 and name HA )) ( (resid 116 and name HD1* )) 3.17 1.37 1.37
assign ((resid 116 and name HN )) ( (resid 116 and name HG2* )) 3.18 1.39 1.39
assign ((resid 119 and name HN )) ( (resid 119 and name HG2* )) 2.79 0.99 0.99
assign ((resid 119 and name HA )) ( (resid 119 and name HG2* )) 2.77 0.97 0.97
assign ((resid 122 and name HA )) ( (resid 122 and name HD2* )) 3.00 1.21 1.21
assign ((resid 122 and name HA )) ( (resid 122 and name HD1* )) 3.13 1.33 1.33
assign ((resid 3 and name HG2* )) ( (resid 4 and name HN )) 3.22 1.42 1.42
assign ((resid 3 and name HD1* )) ( (resid 4 and name HN )) 4.39 2.59 2.59
assign ((resid 5 and name HG2* )) ( (resid 6 and name HN )) 3.38 1.59 1.59
assign ((resid 18 and name HD2* )) ( (resid 19 and name HN )) 3.83 2.04 2.04
assign ((resid 23 and name HD1* )) ( (resid 24 and name HN )) 3.42 1.62 1.62
assign ((resid 26 and name HB* )) ( (resid 27 and name HN )) 3.06 1.26 1.26
assign ((resid 33 and name HB* )) ( (resid 34 and name HN )) 3.04 1.24 1.24
assign ((resid 40 and name HD2* )) ( (resid 41 and name HN )) 4.35 2.56 2.56
assign ((resid 42 and name HG2* )) ( (resid 43 and name HN )) 3.56 1.77 1.77
assign ((resid 43 and name HG2* )) ( (resid 44 and name HN )) 3.40 1.60 1.60
assign ((resid 50 and name HG2* )) ( (resid 51 and name HN )) 3.56 1.77 1.77
assign ((resid 54 and name HG1* )) ( (resid 55 and name HN )) 3.42 1.62 1.62
assign ((resid 77 and name HG2* )) ( (resid 78 and name HN )) 3.76 1.96 1.96
assign ((resid 81 and name HG2* )) ( (resid 82 and name HN )) 3.49 1.69 1.69
assign ((resid 91 and name HG2* )) ( (resid 92 and name HN )) 3.58 1.78 1.78
assign ((resid 91 and name HG1* )) ( (resid 92 and name HN )) 3.69 1.89 1.89
assign ((resid 97 and name HD1* )) ( (resid 98 and name HN )) 3.35 1.55 1.55
assign ((resid 99 and name HD1* )) ( (resid 100 and name HN )) 3.53 1.73 1.73
assign ((resid 104 and name HG2* )) ( (resid 105 and name HN )) 3.33 1.53 1.53
assign ((resid 111 and name HG1* )) ( (resid 112 and name HN )) 3.90 2.11 2.11
assign ((resid 119 and name HG1* )) ( (resid 120 and name HN )) 3.49 1.69 1.69
assign ((resid 11 and name HA )) ( (resid 14 and name HB* )) 3.16 1.37 1.37
assign ((resid 23 and name HA )) ( (resid 26 and name HB* )) 2.88 1.08 1.08
assign ((resid 30 and name HA )) ( (resid 33 and name HB* )) 3.25 1.46 1.46
assign ((resid 31 and name HA )) ( (resid 34 and name HB* )) 3.29 1.50 1.50
assign ((resid 34 and name HA )) ( (resid 37 and name HB* )) 2.95 1.15 1.15
assign ((resid 37 and name HA )) ( (resid 118 and name HB* )) 2.89 1.10 1.10
assign ((resid 65 and name HA )) ( (resid 68 and name HB* )) 3.33 1.53 1.53
assign ((resid 83 and name HA )) ( (resid 86 and name HB* )) 3.20 1.41 1.41
assign ((resid 115 and name HA )) ( (resid 118 and name HB* )) 3.11 1.32 1.32
assign ((resid 4 and name HB* )) ( (resid 9 and name HG2* )) 3.68 1.88 1.88
assign ((resid 4 and name HA )) ( (resid 5 and name HG2* )) 4.41 2.61 2.61
assign ((resid 5 and name HG2* )) ( (resid 7 and name HN )) 3.90 2.11 2.11
assign ((resid 4 and name HB* )) ( (resid 5 and name HG2* )) 4.85 3.05 3.05
assign ((resid 5 and name HB )) ( (resid 40 and name HD2* )) 3.31 1.51 1.51
assign ((resid 7 and name HB* )) ( (resid 37 and name HN )) 3.20 1.41 1.41
assign ((resid 7 and name HB* )) ( (resid 37 and name HA )) 3.38 1.59 1.59
assign ((resid 7 and name HB* )) ( (resid 36 and name HB2 )) 3.25 1.46 1.46
assign ((resid 7 and name HB* )) ( (resid 36 and name HB1 )) 3.25 1.46 1.46
assign ((resid 7 and name HB* )) ( (resid 115 and name HG2* )) 3.76 1.97 1.97
assign ((resid 7 and name HB* )) ( (resid 115 and name HG1* )) 3.98 2.19 2.19
assign ((resid 8 and name HG1 )) ( (resid 115 and name HG1* )) 4.03 2.23 2.23
assign ((resid 6 and name HA )) ( (resid 9 and name HG2* )) 3.15 1.35 1.35
assign ((resid 14 and name HB* )) ( (resid 29 and name HB1 )) 3.09 1.30 1.30
assign ((resid 14 and name HB* )) ( (resid 29 and name HB2 )) 3.09 1.30 1.30
assign ((resid 14 and name HB* )) ( (resid 26 and name HA )) 3.31 1.51 1.51
assign ((resid 14 and name HB* )) ( (resid 29 and name HA )) 4.39 2.59 2.59
assign ((resid 14 and name HB* )) ( (resid 30 and name HN )) 3.22 1.42 1.42
assign ((resid 15 and name HA )) ( (resid 18 and name HD2* )) 4.16 2.36 2.36
assign ((resid 23 and name HD1* )) ( (resid 101 and name HN )) 3.26 1.46 1.46
assign ((resid 23 and name HD2* )) ( (resid 101 and name HN )) 3.99 2.20 2.20
assign ((resid 23 and name HD1* )) ( (resid 101 and name HA )) 3.08 1.28 1.28
assign ((resid 97 and name HD1* )) ( (resid 98 and name HA )) 4.34 2.54 2.54
assign ((resid 23 and name HD1* )) ( (resid 100 and name HA )) 3.44 1.64 1.64
assign ((resid 18 and name HA )) ( (resid 23 and name HD2* )) 3.60 1.80 1.80
assign ((resid 23 and name HD2* )) ( (resid 101 and name HA )) 3.04 1.24 1.24
assign ((resid 23 and name HD1* )) ( (resid 101 and name HB1 )) 3.45 1.66 1.66
assign ((resid 23 and name HD1* )) ( (resid 101 and name HB2 )) 3.45 1.66 1.66
assign ((resid 23 and name HD2* )) ( (resid 101 and name HB2 )) 3.61 1.82 1.82
assign ((resid 23 and name HD2* )) ( (resid 101 and name HB1 )) 3.61 1.82 1.82
assign ((resid 24 and name HG1* )) ( (resid 25 and name HA )) 3.69 1.89 1.89
assign ((resid 24 and name HG1* )) ( (resid 98 and name HA )) 3.33 1.53 1.53
assign ((resid 24 and name HG2* )) ( (resid 98 and name HA )) 4.33 2.54 2.54
assign ((resid 24 and name HG1* )) ( (resid 25 and name HG2 )) 3.98 2.18 2.18
assign ((resid 24 and name HG1* )) ( (resid 25 and name HG1 )) 3.98 2.18 2.18
assign ((resid 18 and name HD1* )) ( (resid 26 and name HB* )) 4.20 2.40 2.40
assign ((resid 23 and name HD2* )) ( (resid 26 and name HB* )) 4.54 2.74 2.74
assign ((resid 14 and name HB* )) ( (resid 26 and name HB* )) 4.75 2.96 2.96
assign ((resid 17 and name HB2 )) ( (resid 26 and name HB* )) 3.45 1.66 1.66
assign ((resid 17 and name HB1 )) ( (resid 26 and name HB* )) 3.34 1.55 1.55
assign ((resid 14 and name HA )) ( (resid 26 and name HB* )) 3.74 1.95 1.95
assign ((resid 18 and name HA )) ( (resid 26 and name HB* )) 3.31 1.51 1.51
assign ((resid 18 and name HN )) ( (resid 26 and name HB* )) 3.15 1.35 1.35
assign ((resid 26 and name HB* )) ( (resid 27 and name HA )) 3.69 1.89 1.89
assign ((resid 30 and name HA )) ( (resid 111 and name HG1* )) 3.81 2.02 2.02
assign ((resid 14 and name HB* )) ( (resid 30 and name HA )) 3.61 1.82 1.82
assign ((resid 31 and name HA )) ( (resid 86 and name HB* )) 3.51 1.71 1.71
assign ((resid 11 and name HN )) ( (resid 33 and name HB* )) 3.15 1.35 1.35
assign ((resid 11 and name HA )) ( (resid 33 and name HB* )) 3.33 1.53 1.53
assign ((resid 10 and name HB1 )) ( (resid 33 and name HB* )) 3.45 1.66 1.66
assign ((resid 11 and name HB1 )) ( (resid 33 and name HB* )) 3.65 1.86 1.86
assign ((resid 11 and name HB2 )) ( (resid 33 and name HB* )) 3.47 1.68 1.68
assign ((resid 10 and name HB2 )) ( (resid 33 and name HB* )) 3.33 1.53 1.53
assign ((resid 34 and name HA )) ( (resid 114 and name HD2* )) 3.60 1.80 1.80
assign ((resid 34 and name HB* )) ( (resid 111 and name HG1* )) 4.02 2.22 2.22
assign ((resid 34 and name HB* )) ( (resid 86 and name HB* )) 3.58 1.79 1.79
assign ((resid 34 and name HB* )) ( (resid 111 and name HG2* )) 4.05 2.26 2.26
assign ((resid 31 and name HE* )) ( (resid 34 and name HB* )) 4.61 2.81 2.81
assign ((resid 35 and name HG2 )) ( (resid 65 and name HD1* )) 3.69 1.89 1.89
assign ((resid 35 and name HG1 )) ( (resid 65 and name HD1* )) 3.69 1.89 1.89
assign ((resid 35 and name HG2 )) ( (resid 65 and name HD2* )) 3.92 2.13 2.13
assign ((resid 35 and name HG1 )) ( (resid 65 and name HD2* )) 3.92 2.13 2.13
assign ((resid 37 and name HA )) ( (resid 40 and name HD1* )) 3.13 1.33 1.33
assign ((resid 7 and name HB* )) ( (resid 37 and name HB* )) 3.40 1.61 1.61
assign ((resid 37 and name HB* )) ( (resid 114 and name HD1* )) 4.70 2.91 2.91
assign ((resid 37 and name HB* )) ( (resid 115 and name HG2* )) 3.98 2.19 2.19
assign ((resid 38 and name HA )) ( (resid 41 and name HD1* )) 3.08 1.28 1.28
assign ((resid 38 and name HD1* )) ( (resid 72 and name HD1* )) 3.96 2.17 2.17
assign ((resid 38 and name HD1* )) ( (resid 114 and name HD1* )) 3.91 2.11 2.11
assign ((resid 38 and name HD1* )) ( (resid 114 and name HD2* )) 4.13 2.33 2.33
assign ((resid 38 and name HD1* )) ( (resid 68 and name HB* )) 4.92 3.12 3.12
assign ((resid 38 and name HD1* )) ( (resid 83 and name HB* )) 4.49 2.69 2.69
assign ((resid 38 and name HD1* )) ( (resid 69 and name HA )) 3.42 1.62 1.62
assign ((resid 5 and name HB )) ( (resid 40 and name HD1* )) 3.58 1.78 1.78
assign ((resid 41 and name HD2* )) ( (resid 45 and name HD22 )) 3.61 1.82 1.82
assign ((resid 41 and name HD2* )) ( (resid 45 and name HD21 )) 3.61 1.82 1.82
assign ((resid 38 and name HD1* )) ( (resid 41 and name HD1* )) 4.29 2.49 2.49
assign ((resid 42 and name HG2* )) ( (resid 50 and name HG2* )) 3.80 2.01 2.01
assign ((resid 38 and name HG2* )) ( (resid 42 and name HG2* )) 3.94 2.15 2.15
assign ((resid 42 and name HA )) ( (resid 47 and name HD1* )) 3.13 1.33 1.33
assign ((resid 42 and name HG1* )) ( (resid 50 and name HB )) 3.11 1.32 1.32
assign ((resid 42 and name HG1* )) ( (resid 47 and name HB2 )) 2.95 1.15 1.15
assign ((resid 42 and name HG2* )) ( (resid 50 and name HB )) 3.94 2.14 2.14
assign ((resid 42 and name HG2* )) ( (resid 47 and name HB2 )) 3.54 1.75 1.75
assign ((resid 39 and name HA )) ( (resid 42 and name HG2* )) 3.54 1.75 1.75
assign ((resid 42 and name HG2* )) ( (resid 60 and name HZ3 )) 3.96 2.16 2.16
assign ((resid 81 and name HN )) ( (resid 81 and name HD1* )) 3.49 1.69 1.69
assign ((resid 40 and name HA )) ( (resid 43 and name HD1* )) 3.38 1.59 1.59
assign ((resid 43 and name HG2* )) ( (resid 44 and name HA )) 3.27 1.48 1.48
assign ((resid 42 and name HG1* )) ( (resid 43 and name HA )) 3.38 1.59 1.59
assign ((resid 42 and name HG1* )) ( (resid 47 and name HB1 )) 3.54 1.75 1.75
assign ((resid 42 and name HG1* )) ( (resid 47 and name HG )) 3.83 2.04 2.04
assign ((resid 42 and name HB )) ( (resid 47 and name HD1* )) 3.99 2.20 2.20
assign ((resid 42 and name HG1* )) ( (resid 50 and name HG2* )) 3.87 2.08 2.08
assign ((resid 50 and name HD1* )) ( (resid 67 and name HB1 )) 2.99 1.19 1.19
assign ((resid 50 and name HD1* )) ( (resid 67 and name HG2 )) 3.35 1.55 1.55
assign ((resid 50 and name HD1* )) ( (resid 68 and name HB* )) 3.80 2.01 2.01
assign ((resid 50 and name HD1* )) ( (resid 68 and name HA )) 3.08 1.28 1.28
assign ((resid 50 and name HD1* )) ( (resid 68 and name HN )) 3.31 1.51 1.51
assign ((resid 50 and name HG2* )) ( (resid 51 and name HA )) 4.26 2.47 2.47
assign ((resid 50 and name HG2* )) ( (resid 54 and name HG2* )) 4.92 3.12 3.12
assign ((resid 51 and name HA )) ( (resid 54 and name HG2* )) 3.38 1.59 1.59
assign ((resid 54 and name HG1* )) ( (resid 60 and name HA )) 3.79 2.00 2.00
assign ((resid 54 and name HG2* )) ( (resid 60 and name HA )) 4.05 2.25 2.25
assign ((resid 65 and name HD2* )) ( (resid 83 and name HZ2 )) 3.17 1.37 1.37
assign ((resid 68 and name HA )) ( (resid 71 and name HD1* )) 3.20 1.41 1.41
assign ((resid 38 and name HG2* )) ( (resid 68 and name HB* )) 3.84 2.04 2.04
assign ((resid 69 and name HA )) ( (resid 72 and name HD1* )) 3.58 1.78 1.78
assign ((resid 69 and name HA )) ( (resid 79 and name HG2* )) 3.27 1.48 1.48
assign ((resid 38 and name HG2* )) ( (resid 72 and name HD1* )) 4.02 2.22 2.22
assign ((resid 47 and name HD1* )) ( (resid 72 and name HA )) 3.58 1.78 1.78
assign ((resid 38 and name HA )) ( (resid 72 and name HD1* )) 3.61 1.82 1.82
assign ((resid 75 and name HB2 )) ( (resid 79 and name HD1* )) 3.36 1.57 1.57
assign ((resid 75 and name HB1 )) ( (resid 79 and name HD1* )) 3.36 1.57 1.57
assign ((resid 76 and name HB1 )) ( (resid 79 and name HD1* )) 3.44 1.64 1.64
assign ((resid 76 and name HA )) ( (resid 77 and name HG2* )) 3.67 1.87 1.87
assign ((resid 78 and name HB2 )) ( (resid 82 and name HD1* )) 3.56 1.77 1.77
assign ((resid 78 and name HB1 )) ( (resid 82 and name HD1* )) 3.47 1.68 1.68
assign ((resid 78 and name HB1 )) ( (resid 82 and name HD2* )) 3.81 2.02 2.02
assign ((resid 78 and name HG2 )) ( (resid 82 and name HD1* )) 4.01 2.22 2.22
assign ((resid 78 and name HG1 )) ( (resid 82 and name HD1* )) 4.01 2.22 2.22
assign ((resid 79 and name HD1* )) ( (resid 117 and name HE* )) 5.47 3.68 3.68
assign ((resid 79 and name HD1* )) ( (resid 117 and name HD* )) 5.17 3.37 3.37
assign ((resid 76 and name HB2 )) ( (resid 79 and name HD1* )) 3.44 1.64 1.64
assign ((resid 34 and name HB* )) ( (resid 114 and name HD1* )) 4.09 2.29 2.29
assign ((resid 38 and name HD1* )) ( (resid 79 and name HG2* )) 3.86 2.06 2.06
assign ((resid 81 and name HG2* )) ( (resid 82 and name HA )) 3.85 2.05 2.05
assign ((resid 78 and name HA )) ( (resid 81 and name HG2* )) 3.83 2.04 2.04
assign ((resid 78 and name HA )) ( (resid 81 and name HD1* )) 3.06 1.26 1.26
assign ((resid 81 and name HG2* )) ( (resid 82 and name HG )) 3.45 1.66 1.66
assign ((resid 79 and name HA )) ( (resid 82 and name HD1* )) 3.29 1.50 1.50
assign ((resid 82 and name HD2* )) ( (resid 110 and name HA )) 2.98 1.19 1.19
assign ((resid 82 and name HD1* )) ( (resid 114 and name HN )) 3.63 1.84 1.84
assign ((resid 86 and name HA )) ( (resid 111 and name HG2* )) 3.65 1.86 1.86
assign ((resid 34 and name HB* )) ( (resid 86 and name HA )) 4.30 2.50 2.50
assign ((resid 88 and name HA )) ( (resid 91 and name HG2* )) 3.38 1.59 1.59
assign ((resid 97 and name HD1* )) ( (resid 99 and name HN )) 3.87 2.07 2.07
assign ((resid 89 and name HA )) ( (resid 99 and name HD1* )) 3.33 1.53 1.53
assign ((resid 101 and name HA )) ( (resid 104 and name HG2* )) 3.25 1.46 1.46
assign ((resid 101 and name HG* )) ( (resid 104 and name HG2* )) 4.06 2.26 2.26
assign ((resid 23 and name HD1* )) ( (resid 101 and name HG* )) 4.34 2.55 2.55
assign ((resid 23 and name HD2* )) ( (resid 101 and name HG* )) 4.15 2.35 2.35
assign ((resid 104 and name HA )) ( (resid 107 and name HD2* )) 3.04 1.24 1.24
assign ((resid 106 and name HE* )) ( (resid 107 and name HD2* )) 4.11 2.31 2.31
assign ((resid 82 and name HD2* )) ( (resid 113 and name HE* )) 3.73 1.94 1.94
assign ((resid 82 and name HD1* )) ( (resid 113 and name HE* )) 4.09 2.30 2.30
assign ((resid 82 and name HD2* )) ( (resid 110 and name HB2 )) 3.61 1.82 1.82
assign ((resid 33 and name HB* )) ( (resid 111 and name HG1* )) 3.96 2.17 2.17
assign ((resid 30 and name HB* )) ( (resid 111 and name HG2* )) 4.00 2.21 2.21
assign ((resid 108 and name HA )) ( (resid 111 and name HG2* )) 3.31 1.51 1.51
assign ((resid 30 and name HB* )) ( (resid 111 and name HG1* )) 4.34 2.55 2.55
assign ((resid 34 and name HA )) ( (resid 111 and name HG1* )) 3.65 1.86 1.86
assign ((resid 34 and name HN )) ( (resid 111 and name HG1* )) 3.40 1.60 1.60
assign ((resid 31 and name HN )) ( (resid 111 and name HG2* )) 4.41 2.61 2.61
assign ((resid 86 and name HN )) ( (resid 111 and name HG2* )) 4.32 2.52 2.52
assign ((resid 11 and name HE* )) ( (resid 111 and name HG2* )) 5.47 3.68 3.68
assign ((resid 112 and name HA )) ( (resid 115 and name HG2* )) 3.63 1.84 1.84
assign ((resid 82 and name HD2* )) ( (resid 113 and name HB2 )) 3.96 2.16 2.16
assign ((resid 82 and name HD2* )) ( (resid 113 and name HB1 )) 3.83 2.04 2.04
assign ((resid 82 and name HD1* )) ( (resid 113 and name HB1 )) 3.69 1.89 1.89
assign ((resid 111 and name HG2* )) ( (resid 114 and name HB2 )) 4.21 2.41 2.41
assign ((resid 82 and name HD1* )) ( (resid 114 and name HB2 )) 3.85 2.05 2.05
assign ((resid 23 and name HD2* )) ( (resid 104 and name HB )) 3.85 2.05 2.05
assign ((resid 37 and name HB* )) ( (resid 115 and name HA )) 2.89 1.10 1.10
assign ((resid 7 and name HB* )) ( (resid 115 and name HA )) 3.72 1.93 1.93
assign ((resid 33 and name HB* )) ( (resid 115 and name HG2* )) 4.23 2.44 2.44
assign ((resid 37 and name HB* )) ( (resid 115 and name HG1* )) 4.70 2.91 2.91
assign ((resid 8 and name HA )) ( (resid 115 and name HG1* )) 3.22 1.42 1.42
assign ((resid 116 and name HG2* )) ( (resid 120 and name HD22 )) 3.18 1.39 1.39
assign ((resid 116 and name HG2* )) ( (resid 120 and name HD21 )) 3.18 1.39 1.39
assign ((resid 113 and name HA )) ( (resid 116 and name HD1* )) 3.24 1.44 1.44
assign ((resid 116 and name HG2* )) ( (resid 117 and name HA )) 3.63 1.84 1.84
assign ((resid 41 and name HN )) ( (resid 118 and name HB* )) 3.63 1.84 1.84
assign ((resid 41 and name HD1* )) ( (resid 118 and name HA )) 2.82 1.03 1.03
assign ((resid 115 and name HG2* )) ( (resid 118 and name HB* )) 4.11 2.31 2.31
assign ((resid 115 and name HG1* )) ( (resid 119 and name HG2* )) 3.85 2.06 2.06
assign ((resid 116 and name HA )) ( (resid 119 and name HG2* )) 3.18 1.39 1.39
assign ((resid 119 and name HA )) ( (resid 122 and name HD1* )) 3.27 1.48 1.48
assign ((resid 5 and name HG2* )) ( (resid 122 and name HD1* )) 4.72 2.92 2.92
assign ((resid 5 and name HG2* )) ( (resid 122 and name HD2* )) 4.83 3.03 3.03
assign ((resid 8 and name HG2 )) ( (resid 115 and name HG1* )) 4.03 2.23 2.23
assign ((resid 37 and name HN )) ( (resid 115 and name HG2* )) 3.74 1.95 1.95
assign ((resid 50 and name HD1* )) ( (resid 67 and name HB2 )) 3.58 1.78 1.78
assign ((resid 85 and name HA )) ( (resid 107 and name HD2* )) 3.15 1.35 1.35
assign ((resid 86 and name HB* )) ( (resid 111 and name HG2* )) 4.03 2.24 2.24
assign ((resid 82 and name HD1* )) ( (resid 113 and name HB2 )) 3.44 1.64 1.64
assign ((resid 34 and name HB* )) ( (resid 114 and name HD2* )) 3.94 2.15 2.15
assign ((resid 41 and name HD1* )) ( (resid 114 and name HA )) 3.29 1.50 1.50
assign ((resid 82 and name HD1* )) ( (resid 114 and name HA )) 3.56 1.77 1.77
assign ((resid 37 and name HB* )) ( (resid 114 and name HD2* )) 4.41 2.62 2.62
assign ((resid 83 and name HA )) ( (resid 114 and name HD2* )) 3.90 2.11 2.11
assign ((resid 83 and name HA )) ( (resid 114 and name HD1* )) 4.10 2.31 2.31
assign ((resid 49 and name HG1 )) ( (resid 71 and name HD1* )) 3.71 1.91 1.91
assign ((resid 49 and name HG2 )) ( (resid 71 and name HD1* )) 3.71 1.91 1.91
assign ((resid 50 and name HD1* )) ( (resid 67 and name HG1 )) 3.35 1.55 1.55
assign ((resid 82 and name HD2* )) ( (resid 110 and name HB1 )) 3.25 1.46 1.46
assign ((resid 87 and name HE1 )) ( (resid 91 and name HG2* )) 3.33 1.53 1.53
assign ((resid 37 and name HA )) ( (resid 40 and name HD2* )) 3.47 1.68 1.68
assign ((resid 88 and name HB )) ( (resid 107 and name HD2* )) 3.20 1.41 1.41
assign ((resid 89 and name HD1* )) ( (resid 111 and name HG2* )) 4.32 2.53 2.53
assign ((resid 41 and name HD1* )) ( (resid 118 and name HB* )) 3.80 2.01 2.01
assign ((resid 115 and name HG1* )) ( (resid 118 and name HB* )) 4.92 3.12 3.12
assign ((resid 40 and name HB2 )) ( (resid 118 and name HB* )) 3.07 1.28 1.28
assign ((resid 119 and name HG1* )) ( (resid 120 and name HA )) 3.71 1.91 1.91
assign ((resid 104 and name HA )) ( (resid 107 and name HD1* )) 3.92 2.13 2.13
assign ((resid 82 and name HD2* )) ( (resid 114 and name HB2 )) 4.24 2.45 2.45
assign ((resid 89 and name HD2* )) ( (resid 111 and name HG2* )) 4.32 2.53 2.53
assign ((resid 111 and name HG2* )) ( (resid 114 and name HD1* )) 3.87 2.08 2.08
assign ((resid 7 and name HB* )) ( (resid 40 and name HD1* )) 4.75 2.96 2.96
assign ((resid 30 and name HB* )) ( (resid 33 and name HB* )) 4.36 2.57 2.57
assign ((resid 7 and name HB* )) ( (resid 33 and name HB* )) 4.14 2.35 2.35
assign ((resid 33 and name HB* )) ( (resid 111 and name HG2* )) 4.92 3.12 3.12
assign ((resid 45 and name HD22 )) ( (resid 72 and name HD2* )) 3.65 1.86 1.86
assign ((resid 42 and name HA )) ( (resid 72 and name HD2* )) 3.44 1.64 1.64
assign ((resid 50 and name HG2* )) ( (resid 68 and name HB* )) 4.00 2.20 2.20
assign ((resid 18 and name HD1* )) ( (resid 101 and name HG* )) 3.97 2.17 2.17
assign ((resid 40 and name HD2* )) ( (resid 118 and name HB* )) 4.05 2.26 2.26
assign ((resid 18 and name HD1* )) ( (resid 23 and name HA )) 3.18 1.39 1.39
assign ((resid 82 and name HD2* )) ( (resid 114 and name HD1* )) 3.78 1.99 1.99
assign ((resid 45 and name HD21 )) ( (resid 72 and name HD2* )) 3.65 1.86 1.86
assign ((resid 50 and name HN )) ( (resid 51 and name HG2* )) 3.58 1.78 1.78
assign ((resid 115 and name HG2* )) ( (resid 116 and name HN )) 3.56 1.77 1.77
assign ((resid 60 and name HB2 )) ( (resid 60 and name HD1 )) 2.69 0.90 0.90
assign ((resid 83 and name HA )) ( (resid 83 and name HE3 )) 2.33 0.54 0.54
assign ((resid 83 and name HA )) ( (resid 83 and name HZ3 )) 3.14 1.35 1.35
assign ((resid 35 and name HA )) ( (resid 83 and name HZ3 )) 2.37 0.57 0.57
assign ((resid 87 and name HB2 )) ( (resid 87 and name HD1 )) 2.66 0.86 0.86
assign ((resid 87 and name HA )) ( (resid 87 and name HE3 )) 2.62 0.82 0.82
assign ((resid 117 and name HD* )) ( (resid 118 and name HN )) 4.64 2.84 2.84
assign ((resid 7 and name HA )) ( (resid 10 and name HD* )) 3.86 2.07 2.07
assign ((resid 10 and name HD* )) ( (resid 33 and name HN )) 4.12 2.32 2.32
assign ((resid 10 and name HD* )) ( (resid 36 and name HG1 )) 3.72 1.92 1.92
assign ((resid 10 and name HD* )) ( (resid 36 and name HB2 )) 4.33 2.54 2.54
assign ((resid 10 and name HD* )) ( (resid 36 and name HB1 )) 4.33 2.54 2.54
assign ((resid 10 and name HD* )) ( (resid 32 and name HB2 )) 3.92 2.12 2.12
assign ((resid 10 and name HD* )) ( (resid 32 and name HB1 )) 3.92 2.12 2.12
assign ((resid 11 and name HD* )) ( (resid 12 and name HN )) 3.99 2.19 2.19
assign ((resid 11 and name HD* )) ( (resid 12 and name HA )) 4.21 2.41 2.41
assign ((resid 11 and name HD* )) ( (resid 12 and name HG* )) 5.40 3.61 3.61
assign ((resid 10 and name HE* )) ( (resid 36 and name HG1 )) 3.85 2.05 2.05
assign ((resid 11 and name HE* )) ( (resid 112 and name HN )) 3.76 1.96 1.96
assign ((resid 11 and name HE* )) ( (resid 112 and name HA )) 4.14 2.34 2.34
assign ((resid 31 and name HD* )) ( (resid 86 and name HA )) 4.96 3.17 3.17
assign ((resid 31 and name HE* )) ( (resid 83 and name HH2 )) 3.51 1.71 1.71
assign ((resid 31 and name HD* )) ( (resid 32 and name HN )) 4.21 2.41 2.41
assign ((resid 31 and name HE* )) ( (resid 87 and name HE3 )) 3.51 1.71 1.71
assign ((resid 31 and name HE* )) ( (resid 87 and name HA )) 3.79 2.00 2.00
assign ((resid 31 and name HE* )) ( (resid 35 and name HG2 )) 4.62 2.83 2.83
assign ((resid 31 and name HE* )) ( (resid 35 and name HG1 )) 4.62 2.83 2.83
assign ((resid 63 and name HA )) ( (resid 66 and name HD* )) 4.46 2.66 2.66
assign ((resid 66 and name HN )) ( (resid 66 and name HE* )) 4.96 3.17 3.17
assign ((resid 17 and name HD* )) ( (resid 18 and name HN )) 4.87 3.08 3.08
assign ((resid 10 and name HD* )) ( (resid 36 and name HG2 )) 3.72 1.92 1.92
assign ((resid 10 and name HE* )) ( (resid 32 and name HE* )) 4.60 2.80 2.80
assign ((resid 10 and name HE* )) ( (resid 36 and name HG2 )) 3.85 2.05 2.05
assign ((resid 17 and name HE* )) ( (resid 25 and name HB2 )) 3.79 2.00 2.00
assign ((resid 17 and name HE* )) ( (resid 25 and name HB1 )) 4.06 2.27 2.27
assign ((resid 14 and name HA )) ( (resid 17 and name HD* )) 3.94 2.14 2.14
assign ((resid 17 and name HD* )) ( (resid 22 and name HB2 )) 4.37 2.57 2.57
assign ((resid 17 and name HD* )) ( (resid 22 and name HB1 )) 4.37 2.57 2.57
assign ((resid 39 and name HA )) ( (resid 60 and name HH2 )) 2.76 0.97 0.97
assign ((resid 43 and name HG12 )) ( (resid 60 and name HZ2 )) 2.87 1.08 1.08
assign ((resid 42 and name HB )) ( (resid 60 and name HH2 )) 2.66 0.86 0.86
assign ((resid 60 and name HE3 )) ( (resid 65 and name HG )) 3.16 1.36 1.36
assign ((resid 35 and name HA )) ( (resid 83 and name HE3 )) 2.94 1.15 1.15
assign ((resid 35 and name HA )) ( (resid 83 and name HH2 )) 2.66 0.86 0.86
assign ((resid 65 and name HB1 )) ( (resid 83 and name HZ2 )) 2.51 0.72 0.72
assign ((resid 65 and name HB1 )) ( (resid 83 and name HH2 )) 2.98 1.18 1.18
assign ((resid 10 and name HD* )) ( (resid 33 and name HA )) 3.90 2.10 2.10
assign ((resid 31 and name HD* )) ( (resid 32 and name HA )) 4.53 2.73 2.73
assign ((resid 17 and name HD* )) ( (resid 26 and name HA )) 4.75 2.95 2.95
assign ((resid 10 and name HD* )) ( (resid 33 and name HB* )) 4.17 2.38 2.38
assign ((resid 9 and name HG2* )) ( (resid 10 and name HD* )) 4.90 3.10 3.10
assign ((resid 9 and name HG1* )) ( (resid 10 and name HD* )) 4.55 2.76 2.76
assign ((resid 11 and name HD* )) ( (resid 33 and name HB* )) 4.35 2.56 2.56
assign ((resid 11 and name HD* )) ( (resid 111 and name HG1* )) 4.05 2.25 2.25
assign ((resid 11 and name HD* )) ( (resid 115 and name HG2* )) 4.21 2.42 2.42
assign ((resid 11 and name HE* )) ( (resid 33 and name HB* )) 5.33 3.53 3.53
assign ((resid 11 and name HE* )) ( (resid 111 and name HG1* )) 4.14 2.35 2.35
assign ((resid 11 and name HE* )) ( (resid 115 and name HG2* )) 4.36 2.56 2.56
assign ((resid 31 and name HD* )) ( (resid 34 and name HB* )) 5.09 3.30 3.30
assign ((resid 31 and name HD* )) ( (resid 86 and name HB* )) 4.17 2.38 2.38
assign ((resid 31 and name HE* )) ( (resid 86 and name HB* )) 4.52 2.72 2.72
assign ((resid 41 and name HD1* )) ( (resid 117 and name HE* )) 4.18 2.38 2.38
assign ((resid 87 and name HD1 )) ( (resid 91 and name HG2* )) 3.20 1.41 1.41
assign ((resid 11 and name HD* )) ( (resid 115 and name HG1* )) 4.98 3.19 3.19
assign ((resid 41 and name HD1* )) ( (resid 117 and name HD* )) 3.98 2.18 2.18
assign ((resid 41 and name HD2* )) ( (resid 117 and name HD* )) 4.10 2.31 2.31
assign ((resid 41 and name HD2* )) ( (resid 117 and name HE* )) 4.23 2.44 2.44
assign ((resid 60 and name HZ3 )) ( (resid 68 and name HB* )) 3.99 2.20 2.20
assign ((resid 42 and name HG1* )) ( (resid 60 and name HH2 )) 2.79 0.99 0.99
assign ((resid 43 and name HD1* )) ( (resid 60 and name HH2 )) 3.67 1.87 1.87
assign ((resid 50 and name HG2* )) ( (resid 60 and name HH2 )) 3.09 1.30 1.30
assign ((resid 42 and name HG1* )) ( (resid 60 and name HZ2 )) 3.20 1.41 1.41
assign ((resid 43 and name HD1* )) ( (resid 60 and name HZ2 )) 3.13 1.33 1.33
assign ((resid 54 and name HG1* )) ( (resid 60 and name HZ2 )) 3.69 1.89 1.89
assign ((resid 54 and name HG2* )) ( (resid 60 and name HZ2 )) 3.36 1.57 1.57
assign ((resid 42 and name HG1* )) ( (resid 60 and name HZ3 )) 3.15 1.35 1.35
assign ((resid 50 and name HG2* )) ( (resid 60 and name HZ3 )) 3.13 1.33 1.33
assign ((resid 60 and name HZ3 )) ( (resid 65 and name HD2* )) 3.40 1.60 1.60
assign ((resid 60 and name HE3 )) ( (resid 65 and name HD1* )) 3.54 1.75 1.75
assign ((resid 50 and name HG2* )) ( (resid 60 and name HE3 )) 3.92 2.13 2.13
assign ((resid 60 and name HE3 )) ( (resid 65 and name HD2* )) 3.09 1.30 1.30
assign ((resid 54 and name HG1* )) ( (resid 60 and name HD1 )) 3.27 1.48 1.48
assign ((resid 11 and name HE* )) ( (resid 115 and name HG1* )) 4.74 2.94 2.94
assign ((resid 87 and name HZ2 )) ( (resid 91 and name HG1* )) 3.65 1.86 1.86
assign ((resid 87 and name HZ2 )) ( (resid 91 and name HG2* )) 3.18 1.39 1.39
assign ((resid 34 and name HB* )) ( (resid 83 and name HE3 )) 2.95 1.15 1.15
assign ((resid 38 and name HD1* )) ( (resid 83 and name HE3 )) 3.40 1.60 1.60
assign ((resid 34 and name HB* )) ( (resid 83 and name HZ3 )) 2.89 1.10 1.10
assign ((resid 38 and name HD1* )) ( (resid 83 and name HZ3 )) 3.53 1.73 1.73
assign ((resid 65 and name HD1* )) ( (resid 83 and name HZ2 )) 3.09 1.30 1.30
assign ((resid 65 and name HD1* )) ( (resid 83 and name HH2 )) 2.86 1.06 1.06
assign ((resid 65 and name HD2* )) ( (resid 83 and name HH2 )) 3.31 1.51 1.51
assign ((resid 42 and name HG2* )) ( (resid 60 and name HH2 )) 2.80 1.01 1.01
assign ((resid 51 and name HG2* )) ( (resid 60 and name HZ2 )) 3.11 1.32 1.32
assign ((resid 3 and name HA )) ( (resid 3 and name HG1* )) 2.74 0.95 0.95
assign ((resid 3 and name HG1* )) ( (resid 4 and name HN )) 2.99 1.19 1.19
assign ((resid 6 and name HN )) ( (resid 6 and name HB* )) 2.69 0.89 0.89
assign ((resid 6 and name HB* )) ( (resid 10 and name HD* )) 4.93 3.14 3.14
assign ((resid 6 and name HB* )) ( (resid 10 and name HE* )) 4.38 2.58 2.58
assign ((resid 7 and name HA )) ( (resid 36 and name HB* )) 3.47 1.68 1.68
assign ((resid 7 and name HB* )) ( (resid 36 and name HB* )) 3.16 1.36 1.36
assign ((resid 8 and name HN )) ( (resid 8 and name HG* )) 2.77 0.97 0.97
assign ((resid 8 and name HA )) ( (resid 8 and name HG* )) 2.80 1.01 1.01
assign ((resid 8 and name HG* )) ( (resid 9 and name HN )) 4.34 2.54 2.54
assign ((resid 8 and name HG* )) ( (resid 115 and name HG1* )) 3.75 1.96 1.96
assign ((resid 8 and name HG* )) ( (resid 119 and name HG2* )) 3.66 1.86 1.86
assign ((resid 9 and name HA )) ( (resid 12 and name HB* )) 2.68 0.89 0.89
assign ((resid 10 and name HD* )) ( (resid 32 and name HB* )) 3.83 2.04 2.04
assign ((resid 10 and name HD* )) ( (resid 32 and name HD* )) 4.39 2.60 2.60
assign ((resid 10 and name HD* )) ( (resid 36 and name HB* )) 4.17 2.38 2.38
assign ((resid 10 and name HD* )) ( (resid 36 and name HG* )) 3.61 1.82 1.82
assign ((resid 10 and name HE* )) ( (resid 32 and name HD* )) 4.31 2.51 2.51
assign ((resid 10 and name HE* )) ( (resid 36 and name HG* )) 3.66 1.87 1.87
assign ((resid 11 and name HE* )) ( (resid 15 and name HG* )) 4.32 2.53 2.53
assign ((resid 11 and name HE* )) ( (resid 112 and name HB* )) 4.13 2.33 2.33
assign ((resid 11 and name HE* )) ( (resid 112 and name HG* )) 4.40 2.60 2.60
assign ((resid 13 and name HN )) ( (resid 13 and name HB* )) 2.66 0.86 0.86
assign ((resid 13 and name HN )) ( (resid 13 and name HG* )) 2.64 0.84 0.84
assign ((resid 13 and name HA )) ( (resid 13 and name HG* )) 2.67 0.87 0.87
assign ((resid 13 and name HA )) ( (resid 16 and name HB* )) 2.58 0.79 0.79
assign ((resid 13 and name HB* )) ( (resid 14 and name HN )) 2.67 0.87 0.87
assign ((resid 13 and name HG* )) ( (resid 14 and name HN )) 3.66 1.87 1.87
assign ((resid 14 and name HN )) ( (resid 29 and name HB* )) 3.53 1.73 1.73
assign ((resid 14 and name HA )) ( (resid 29 and name HB* )) 3.53 1.73 1.73
assign ((resid 14 and name HB* )) ( (resid 29 and name HB* )) 2.98 1.18 1.18
assign ((resid 15 and name HA )) ( (resid 15 and name HG* )) 2.75 0.95 0.95
assign ((resid 15 and name HA )) ( (resid 18 and name HB* )) 2.64 0.84 0.84
assign ((resid 15 and name HG* )) ( (resid 16 and name HN )) 3.73 1.93 1.93
assign ((resid 16 and name HN )) ( (resid 16 and name HG* )) 2.73 0.93 0.93
assign ((resid 16 and name HA )) ( (resid 16 and name HG* )) 2.77 0.98 0.98
assign ((resid 16 and name HB* )) ( (resid 17 and name HN )) 2.62 0.82 0.82
assign ((resid 16 and name HG* )) ( (resid 17 and name HN )) 4.10 2.31 2.31
assign ((resid 17 and name HA )) ( (resid 20 and name HB* )) 2.78 0.98 0.98
assign ((resid 17 and name HA )) ( (resid 22 and name HB* )) 4.34 2.54 2.54
assign ((resid 17 and name HD* )) ( (resid 22 and name HB* )) 4.23 2.44 2.44
assign ((resid 17 and name HD* )) ( (resid 29 and name HG* )) 4.38 2.58 2.58
assign ((resid 17 and name HE* )) ( (resid 29 and name HG* )) 4.41 2.62 2.62
assign ((resid 18 and name HN )) ( (resid 18 and name HB* )) 2.60 0.81 0.81
assign ((resid 18 and name HB* )) ( (resid 19 and name HN )) 2.72 0.93 0.93
assign ((resid 18 and name HB* )) ( (resid 26 and name HB* )) 4.22 2.42 2.42
assign ((resid 20 and name HN )) ( (resid 20 and name HB* )) 2.53 0.73 0.73
assign ((resid 20 and name HB* )) ( (resid 22 and name HN )) 3.12 1.32 1.32
assign ((resid 22 and name HN )) ( (resid 22 and name HB* )) 2.52 0.72 0.72
assign ((resid 23 and name HD1* )) ( (resid 101 and name HB* )) 3.35 1.56 1.56
assign ((resid 23 and name HD2* )) ( (resid 101 and name HB* )) 3.46 1.67 1.67
assign ((resid 24 and name HA )) ( (resid 27 and name HB* )) 3.17 1.37 1.37
assign ((resid 24 and name HG1* )) ( (resid 25 and name HG* )) 3.90 2.11 2.11
assign ((resid 25 and name HN )) ( (resid 25 and name HG* )) 2.52 0.73 0.73
assign ((resid 25 and name HA )) ( (resid 25 and name HG* )) 2.50 0.70 0.70
assign ((resid 25 and name HA )) ( (resid 25 and name HE2* )) 4.33 2.53 2.53
assign ((resid 25 and name HG* )) ( (resid 26 and name HN )) 4.34 2.54 2.54
assign ((resid 26 and name HA )) ( (resid 29 and name HB* )) 3.17 1.37 1.37
assign ((resid 26 and name HB* )) ( (resid 27 and name HB* )) 4.63 2.84 2.84
assign ((resid 27 and name HN )) ( (resid 27 and name HB* )) 2.56 0.77 0.77
assign ((resid 27 and name HA )) ( (resid 89 and name HD* )) 4.57 2.77 2.77
assign ((resid 27 and name HB* )) ( (resid 28 and name HN )) 2.88 1.09 1.09
assign ((resid 28 and name HN )) ( (resid 28 and name HG* )) 3.42 1.63 1.63
assign ((resid 29 and name HB* )) ( (resid 30 and name HN )) 3.02 1.23 1.23
assign ((resid 31 and name HN )) ( (resid 31 and name HB* )) 2.55 0.76 0.76
assign ((resid 31 and name HD* )) ( (resid 35 and name HG* )) 4.32 2.53 2.53
assign ((resid 31 and name HE* )) ( (resid 35 and name HB* )) 4.99 3.20 3.20
assign ((resid 31 and name HE* )) ( (resid 35 and name HG* )) 4.43 2.63 2.63
assign ((resid 32 and name HN )) ( (resid 32 and name HB* )) 2.63 0.83 0.83
assign ((resid 32 and name HN )) ( (resid 32 and name HG* )) 2.98 1.18 1.18
assign ((resid 32 and name HA )) ( (resid 32 and name HG* )) 2.89 1.10 1.10
assign ((resid 32 and name HA )) ( (resid 32 and name HD* )) 3.17 1.38 1.38
assign ((resid 32 and name HA )) ( (resid 35 and name HB* )) 3.31 1.52 1.52
assign ((resid 32 and name HB* )) ( (resid 33 and name HN )) 2.83 1.03 1.03
assign ((resid 33 and name HA )) ( (resid 36 and name HB* )) 3.35 1.55 1.55
assign ((resid 35 and name HN )) ( (resid 35 and name HB* )) 2.75 0.95 0.95
assign ((resid 35 and name HN )) ( (resid 35 and name HG* )) 2.95 1.16 1.16
assign ((resid 35 and name HA )) ( (resid 35 and name HG* )) 2.76 0.96 0.96
assign ((resid 35 and name HB* )) ( (resid 83 and name HZ2 )) 3.11 1.32 1.32
assign ((resid 35 and name HB* )) ( (resid 83 and name HH2 )) 3.10 1.30 1.30
assign ((resid 35 and name HG* )) ( (resid 36 and name HN )) 3.67 1.88 1.88
assign ((resid 35 and name HG* )) ( (resid 65 and name HD1* )) 3.58 1.79 1.79
assign ((resid 35 and name HG* )) ( (resid 65 and name HD2* )) 3.74 1.95 1.95
assign ((resid 35 and name HG* )) ( (resid 83 and name HH2 )) 3.01 1.21 1.21
assign ((resid 36 and name HN )) ( (resid 36 and name HB* )) 2.70 0.91 0.91
assign ((resid 36 and name HN )) ( (resid 36 and name HG* )) 2.86 1.07 1.07
assign ((resid 36 and name HB* )) ( (resid 37 and name HN )) 3.09 1.30 1.30
assign ((resid 36 and name HG* )) ( (resid 37 and name HN )) 4.34 2.54 2.54
assign ((resid 38 and name HN )) ( (resid 38 and name HG1* )) 2.66 0.87 0.87
assign ((resid 38 and name HA )) ( (resid 38 and name HG1* )) 2.80 1.00 1.00
assign ((resid 38 and name HA )) ( (resid 41 and name HB* )) 3.13 1.34 1.34
assign ((resid 38 and name HG1* )) ( (resid 83 and name HE3 )) 2.93 1.14 1.14
assign ((resid 38 and name HG1* )) ( (resid 83 and name HZ3 )) 3.13 1.34 1.34
assign ((resid 38 and name HD1* )) ( (resid 69 and name HB* )) 4.40 2.60 2.60
assign ((resid 41 and name HN )) ( (resid 41 and name HB* )) 2.71 0.92 0.92
assign ((resid 41 and name HA )) ( (resid 45 and name HD2* )) 3.59 1.79 1.79
assign ((resid 41 and name HB* )) ( (resid 45 and name HD2* )) 4.31 2.52 2.52
assign ((resid 41 and name HB2 )) ( (resid 45 and name HD21 )) 4.78 2.98 2.98
assign ((resid 41 and name HB2 )) ( (resid 45 and name HD22 )) 4.78 2.98 2.98
assign ((resid 41 and name HD1* )) ( (resid 45 and name HD2* )) 3.92 2.12 2.12
assign ((resid 41 and name HD2* )) ( (resid 45 and name HD2* )) 3.53 1.74 1.74
assign ((resid 44 and name HN )) ( (resid 44 and name HB* )) 2.69 0.90 0.90
assign ((resid 44 and name HN )) ( (resid 44 and name HG* )) 2.87 1.08 1.08
assign ((resid 44 and name HA )) ( (resid 44 and name HG* )) 2.70 0.91 0.91
assign ((resid 44 and name HB* )) ( (resid 45 and name HN )) 2.96 1.17 1.17
assign ((resid 44 and name HB* )) ( (resid 45 and name HD2* )) 4.77 2.97 2.97
assign ((resid 45 and name HN )) ( (resid 45 and name HB* )) 2.57 0.78 0.78
assign ((resid 45 and name HB* )) ( (resid 47 and name HD1* )) 4.76 2.96 2.96
assign ((resid 45 and name HD2* )) ( (resid 72 and name HD2* )) 3.56 1.77 1.77
assign ((resid 46 and name HN )) ( (resid 46 and name HB* )) 2.59 0.79 0.79
assign ((resid 48 and name HN )) ( (resid 48 and name HG* )) 3.04 1.24 1.24
assign ((resid 48 and name HA )) ( (resid 48 and name HG* )) 2.59 0.79 0.79
assign ((resid 48 and name HB* )) ( (resid 49 and name HN )) 2.73 0.93 0.93
assign ((resid 48 and name HB* )) ( (resid 52 and name HN )) 4.07 2.27 2.27
assign ((resid 48 and name HB* )) ( (resid 52 and name HD2* )) 3.79 2.00 2.00
assign ((resid 48 and name HB2 )) ( (resid 52 and name HD21 )) 4.38 2.59 2.59
assign ((resid 48 and name HB2 )) ( (resid 52 and name HD22 )) 4.38 2.59 2.59
assign ((resid 49 and name HN )) ( (resid 49 and name HG* )) 2.98 1.19 1.19
assign ((resid 49 and name HA )) ( (resid 49 and name HG* )) 2.64 0.84 0.84
assign ((resid 49 and name HA )) ( (resid 52 and name HD2* )) 3.57 1.77 1.77
assign ((resid 49 and name HB* )) ( (resid 50 and name HN )) 3.09 1.30 1.30
assign ((resid 49 and name HG* )) ( (resid 50 and name HN )) 3.33 1.53 1.53
assign ((resid 49 and name HG* )) ( (resid 71 and name HD1* )) 3.58 1.79 1.79
assign ((resid 50 and name HN )) ( (resid 50 and name HG1* )) 2.96 1.17 1.17
assign ((resid 50 and name HA )) ( (resid 53 and name HD2* )) 3.43 1.63 1.63
assign ((resid 50 and name HG2* )) ( (resid 64 and name HD2* )) 4.66 2.86 2.86
assign ((resid 50 and name HG1* )) ( (resid 51 and name HN )) 4.34 2.54 2.54
assign ((resid 50 and name HD1* )) ( (resid 67 and name HD* )) 3.61 1.81 1.81
assign ((resid 53 and name HB* )) ( (resid 64 and name HD2* )) 4.75 2.95 2.95
assign ((resid 55 and name HN )) ( (resid 55 and name HG* )) 3.13 1.33 1.33
assign ((resid 55 and name HB* )) ( (resid 56 and name HN )) 2.70 0.90 0.90
assign ((resid 55 and name HG* )) ( (resid 56 and name HN )) 3.71 1.91 1.91
assign ((resid 56 and name HN )) ( (resid 56 and name HB* )) 2.43 0.64 0.64
assign ((resid 56 and name HB* )) ( (resid 57 and name HN )) 2.67 0.87 0.87
assign ((resid 57 and name HN )) ( (resid 57 and name HG* )) 2.79 0.99 0.99
assign ((resid 57 and name HA )) ( (resid 57 and name HG* )) 2.68 0.89 0.89
assign ((resid 57 and name HB* )) ( (resid 58 and name HN )) 3.01 1.21 1.21
assign ((resid 57 and name HG* )) ( (resid 58 and name HN )) 4.14 2.34 2.34
assign ((resid 59 and name HB* )) ( (resid 60 and name HN )) 2.82 1.03 1.03
assign ((resid 61 and name HN )) ( (resid 61 and name HB* )) 2.77 0.98 0.98
assign ((resid 61 and name HN )) ( (resid 61 and name HG* )) 3.20 1.41 1.41
assign ((resid 61 and name HB* )) ( (resid 62 and name HN )) 3.13 1.34 1.34
assign ((resid 61 and name HB* )) ( (resid 64 and name HN )) 3.49 1.70 1.70
assign ((resid 61 and name HB* )) ( (resid 64 and name HD2* )) 4.77 2.97 2.97
assign ((resid 62 and name HN )) ( (resid 62 and name HB* )) 2.77 0.98 0.98
assign ((resid 62 and name HB* )) ( (resid 65 and name HD1* )) 3.89 2.10 2.10
assign ((resid 63 and name HB* )) ( (resid 64 and name HN )) 2.93 1.13 1.13
assign ((resid 63 and name HG* )) ( (resid 64 and name HD2* )) 4.77 2.97 2.97
assign ((resid 64 and name HN )) ( (resid 64 and name HB* )) 2.45 0.66 0.66
assign ((resid 64 and name HA )) ( (resid 64 and name HD2* )) 3.07 1.27 1.27
assign ((resid 64 and name HB* )) ( (resid 65 and name HN )) 3.00 1.20 1.20
assign ((resid 66 and name HN )) ( (resid 66 and name HB* )) 2.66 0.87 0.87
assign ((resid 66 and name HB* )) ( (resid 67 and name HN )) 2.85 1.05 1.05
assign ((resid 67 and name HA )) ( (resid 67 and name HD* )) 3.19 1.39 1.39
assign ((resid 67 and name HB2 )) ( (resid 67 and name HD* )) 2.78 0.99 0.99
assign ((resid 67 and name HB1 )) ( (resid 67 and name HD* )) 2.64 0.85 0.85
assign ((resid 68 and name HA )) ( (resid 71 and name HB* )) 3.26 1.46 1.46
assign ((resid 69 and name HN )) ( (resid 69 and name HB* )) 2.69 0.89 0.89
assign ((resid 69 and name HN )) ( (resid 70 and name HG* )) 4.34 2.54 2.54
assign ((resid 69 and name HB* )) ( (resid 79 and name HG2* )) 4.07 2.28 2.28
assign ((resid 69 and name HB* )) ( (resid 80 and name HA )) 2.95 1.15 1.15
assign ((resid 70 and name HN )) ( (resid 70 and name HG* )) 3.07 1.28 1.28
assign ((resid 70 and name HA )) ( (resid 70 and name HG* )) 2.71 0.92 0.92
assign ((resid 70 and name HG* )) ( (resid 71 and name HN )) 3.47 1.68 1.68
assign ((resid 71 and name HN )) ( (resid 71 and name HB* )) 2.63 0.84 0.84
assign ((resid 71 and name HB* )) ( (resid 72 and name HN )) 2.74 0.94 0.94
assign ((resid 72 and name HN )) ( (resid 72 and name HB* )) 2.79 0.99 0.99
assign ((resid 72 and name HB* )) ( (resid 79 and name HG2* )) 3.60 1.81 1.81
assign ((resid 72 and name HB* )) ( (resid 79 and name HD1* )) 3.64 1.85 1.85
assign ((resid 73 and name HN )) ( (resid 73 and name HG* )) 3.12 1.33 1.33
assign ((resid 73 and name HA )) ( (resid 73 and name HG* )) 2.66 0.86 0.86
assign ((resid 73 and name HB1 )) ( (resid 73 and name HG* )) 2.23 0.43 0.43
assign ((resid 73 and name HG* )) ( (resid 74 and name HN )) 4.34 2.54 2.54
assign ((resid 73 and name HG* )) ( (resid 77 and name HA )) 2.60 0.80 0.80
assign ((resid 75 and name HN )) ( (resid 75 and name HB* )) 2.48 0.69 0.69
assign ((resid 75 and name HA )) ( (resid 75 and name HD2* )) 3.19 1.40 1.40
assign ((resid 76 and name HB* )) ( (resid 79 and name HD1* )) 3.24 1.44 1.44
assign ((resid 78 and name HN )) ( (resid 78 and name HG* )) 2.57 0.78 0.78
assign ((resid 78 and name HG* )) ( (resid 82 and name HD1* )) 3.94 2.15 2.15
assign ((resid 78 and name HG* )) ( (resid 82 and name HD2* )) 4.11 2.31 2.31
assign ((resid 79 and name HN )) ( (resid 80 and name HD* )) 2.76 0.96 0.96
assign ((resid 79 and name HB )) ( (resid 80 and name HD* )) 2.74 0.94 0.94
assign ((resid 80 and name HD* )) ( (resid 81 and name HN )) 3.49 1.70 1.70
assign ((resid 81 and name HN )) ( (resid 81 and name HG1* )) 2.75 0.95 0.95
assign ((resid 81 and name HA )) ( (resid 81 and name HG1* )) 2.77 0.97 0.97
assign ((resid 81 and name HA )) ( (resid 84 and name HB* )) 2.47 0.68 0.68
assign ((resid 82 and name HN )) ( (resid 82 and name HB* )) 2.82 1.03 1.03
assign ((resid 82 and name HA )) ( (resid 85 and name HB* )) 3.82 2.02 2.02
assign ((resid 82 and name HB* )) ( (resid 83 and name HN )) 2.89 1.10 1.10
assign ((resid 82 and name HB* )) ( (resid 114 and name HD2* )) 3.79 1.99 1.99
assign ((resid 82 and name HD1* )) ( (resid 113 and name HD* )) 4.06 2.26 2.26
assign ((resid 82 and name HD2* )) ( (resid 113 and name HD* )) 3.55 1.76 1.76
assign ((resid 84 and name HN )) ( (resid 84 and name HG* )) 3.20 1.40 1.40
assign ((resid 84 and name HA )) ( (resid 84 and name HG* )) 2.84 1.05 1.05
assign ((resid 85 and name HN )) ( (resid 85 and name HB* )) 2.74 0.94 0.94
assign ((resid 85 and name HB* )) ( (resid 86 and name HN )) 2.89 1.10 1.10
assign ((resid 85 and name HB* )) ( (resid 107 and name HD2* )) 3.91 2.12 2.12
assign ((resid 87 and name HA )) ( (resid 90 and name HB* )) 3.19 1.39 1.39
assign ((resid 88 and name HG2* )) ( (resid 92 and name HG* )) 4.31 2.51 2.51
assign ((resid 89 and name HN )) ( (resid 89 and name HD* )) 4.35 2.56 2.56
assign ((resid 89 and name HD* )) ( (resid 104 and name HA )) 4.07 2.27 2.27
assign ((resid 89 and name HD* )) ( (resid 111 and name HG2* )) 3.98 2.18 2.18
assign ((resid 90 and name HN )) ( (resid 90 and name HB* )) 2.61 0.81 0.81
assign ((resid 90 and name HB* )) ( (resid 90 and name HD2 )) 2.68 0.88 0.88
assign ((resid 90 and name HB* )) ( (resid 91 and name HN )) 3.04 1.24 1.24
assign ((resid 92 and name HN )) ( (resid 92 and name HG* )) 3.87 2.07 2.07
assign ((resid 92 and name HA )) ( (resid 92 and name HG* )) 2.78 0.99 0.99
assign ((resid 94 and name HN )) ( (resid 95 and name HB* )) 3.53 1.73 1.73
assign ((resid 94 and name HB* )) ( (resid 95 and name HN )) 2.96 1.16 1.16
assign ((resid 95 and name HN )) ( (resid 95 and name HB* )) 2.80 1.00 1.00
assign ((resid 97 and name HB* )) ( (resid 97 and name HG )) 2.23 0.43 0.43
assign ((resid 100 and name HN )) ( (resid 103 and name HB* )) 3.35 1.55 1.55
assign ((resid 100 and name HB* )) ( (resid 102 and name HN )) 3.18 1.39 1.39
assign ((resid 101 and name HN )) ( (resid 101 and name HB* )) 2.55 0.75 0.75
assign ((resid 101 and name HB* )) ( (resid 102 and name HN )) 2.66 0.87 0.87
assign ((resid 101 and name HG* )) ( (resid 105 and name HG* )) 3.95 2.16 2.16
assign ((resid 101 and name HG* )) ( (resid 105 and name HE* )) 4.15 2.35 2.35
assign ((resid 102 and name HN )) ( (resid 102 and name HG* )) 2.90 1.10 1.10
assign ((resid 102 and name HN )) ( (resid 102 and name HD* )) 3.55 1.75 1.75
assign ((resid 102 and name HA )) ( (resid 102 and name HG* )) 2.68 0.89 0.89
assign ((resid 102 and name HA )) ( (resid 102 and name HD* )) 3.49 1.70 1.70
assign ((resid 102 and name HG* )) ( (resid 103 and name HN )) 4.34 2.54 2.54
assign ((resid 103 and name HB* )) ( (resid 104 and name HN )) 2.80 1.01 1.01
assign ((resid 103 and name HB* )) ( (resid 104 and name HG2* )) 3.98 2.19 2.19
assign ((resid 104 and name HA )) ( (resid 107 and name HB* )) 3.20 1.41 1.41
assign ((resid 105 and name HN )) ( (resid 105 and name HG* )) 2.86 1.06 1.06
assign ((resid 105 and name HA )) ( (resid 105 and name HG* )) 2.88 1.08 1.08
assign ((resid 105 and name HA )) ( (resid 105 and name HE* )) 3.65 1.86 1.86
assign ((resid 105 and name HG* )) ( (resid 105 and name HE* )) 2.67 0.88 0.88
assign ((resid 106 and name HN )) ( (resid 106 and name HG* )) 3.05 1.26 1.26
assign ((resid 107 and name HN )) ( (resid 107 and name HB* )) 2.68 0.88 0.88
assign ((resid 107 and name HB* )) ( (resid 108 and name HN )) 3.02 1.23 1.23
assign ((resid 108 and name HN )) ( (resid 108 and name HB* )) 2.80 1.01 1.01
assign ((resid 108 and name HA )) ( (resid 108 and name HD* )) 4.34 2.54 2.54
assign ((resid 108 and name HG* )) ( (resid 112 and name HD* )) 4.78 2.98 2.98
assign ((resid 108 and name HD* )) ( (resid 109 and name HN )) 3.62 1.82 1.82
assign ((resid 109 and name HN )) ( (resid 109 and name HG* )) 3.06 1.27 1.27
assign ((resid 109 and name HA )) ( (resid 112 and name HB* )) 2.52 0.73 0.73
assign ((resid 109 and name HG* )) ( (resid 110 and name HN )) 4.28 2.49 2.49
assign ((resid 112 and name HG* )) ( (resid 113 and name HN )) 4.19 2.40 2.40
assign ((resid 112 and name HG* )) ( (resid 116 and name HD1* )) 3.84 2.04 2.04
assign ((resid 112 and name HD* )) ( (resid 116 and name HD1* )) 4.42 2.62 2.62
assign ((resid 113 and name HN )) ( (resid 113 and name HG* )) 2.66 0.87 0.87
assign ((resid 113 and name HN )) ( (resid 113 and name HD* )) 3.71 1.91 1.91
assign ((resid 113 and name HA )) ( (resid 113 and name HG* )) 2.69 0.90 0.90
assign ((resid 116 and name HN )) ( (resid 116 and name HG1* )) 2.61 0.82 0.82
assign ((resid 116 and name HA )) ( (resid 116 and name HG1* )) 2.69 0.90 0.90
assign ((resid 116 and name HG2* )) ( (resid 120 and name HD2* )) 3.10 1.30 1.30
assign ((resid 116 and name HG1* )) ( (resid 117 and name HN )) 3.72 1.93 1.93
assign ((resid 117 and name HA )) ( (resid 120 and name HB* )) 2.76 0.97 0.97
assign ((resid 118 and name HA )) ( (resid 121 and name HB* )) 2.54 0.74 0.74
assign ((resid 120 and name HN )) ( (resid 120 and name HB* )) 2.39 0.59 0.59
assign ((resid 120 and name HB* )) ( (resid 121 and name HN )) 2.82 1.03 1.03
assign ((resid 121 and name HB* )) ( (resid 122 and name HN )) 3.02 1.22 1.22
assign ((resid 121 and name HB* )) ( (resid 122 and name HD1* )) 3.98 2.19 2.19
assign ((resid 123 and name HN )) ( (resid 123 and name HG* )) 2.69 0.89 0.89
assign ((resid 124 and name HN )) ( (resid 124 and name HB* )) 2.58 0.79 0.79
assign ((resid 125 and name HN )) ( (resid 125 and name HB* )) 2.61 0.82 0.82
assign ((resid 126 and name HN )) ( (resid 126 and name HB* )) 2.77 0.97 0.97
list of removed NOE constraints
409-> SER 121 HN - LEU 122 HB* 1.79 6.13 # NoRestrctn S [2.00 6.01] -- sequential
449-> LYS 48 HN - LYS 48 HD* 1.79 6.77 # NoRestrctn I [2.29 6.01] -- intra
453-> LYS 55 HN - LYS 55 HD* 1.80 6.88 # NoRestrctn I [2.29 6.01] -- intra
454-> LYS 57 HN - LYS 57 HD* 1.80 6.88 # NoRestrctn I [2.29 6.01] -- intra
462-> LYS 70 HN - LYS 70 HD* 1.80 6.88 # NoRestrctn I [2.29 6.01] -- intra
471-> GLU 96 HN - GLU 96 HG* 1.79 6.23 # NoRestrctn I [2.29 6.01] -- intra
478-> LYS 106 HN - LYS 106 HD* 1.80 6.74 # NoRestrctn I [2.29 6.01] -- intra
561-> SER 6 HN - ALA 7 HB* 1.79 6.95 # NoRestrctn S [2.00 6.01] -- sequential
572-> GLU 36 HN - ALA 37 HB* 1.80 7.02 # NoRestrctn S [2.00 6.01] -- sequential
878-> LYS 20 HN - LYS 20 HD* 1.80 6.20 # NoRestrctn I [2.29 6.01] -- intra
919-> ARG 59 HN - ARG 59 HD* 1.79 6.77 # NoRestrctn I [2.29 6.01] -- intra
1006-> LEU 18 HN - LEU 18 HD2* 1.80 6.12 # NoRestrctn I [2.29 6.01] -- intra
1028-> LEU 47 HN - LEU 47 HD2* 1.79 6.33 # NoRestrctn I [2.29 6.01] -- intra
1037-> LEU 71 HA - LEU 71 HD1* 1.80 6.30 # NoRestrctn I [2.11 5.99] -- intra
1059-> LEU 107 HN - LEU 107 HD1* 1.80 6.26 # NoRestrctn I [2.29 6.01] -- intra
1144-> GLN 25 HN - ALA 26 HB* 1.79 6.59 # NoRestrctn S [2.00 6.01] -- sequential
1476-> ALA 26 HN - CYS 27 HB* 1.79 6.23 # NoRestrctn S [2.00 6.01] -- sequential
1527-> ASN 45 HN - ASN 45 HD2* 1.79 6.25 # NoRestrctn I [2.29 6.01] -- intra
1566-> SER 62 HA - SER 62 HB* 1.80 2.60 # FixedDistn I [0.00 0.00] -- intra
1575-> LYS 67 HN - LYS 67 HD* 1.79 6.77 # NoRestrctn I [2.29 6.01] -- intra
====== TOTAL ======: 20
table of distance constraints violations
Residual Violations greater than 0.10
1-> SER 4 HN - THR 5 HN [ 1.80 3.52] 0.00 0.00 0.00 0.00 0.00 0.00 0.12 0.02 0.03 0.01 0.00 0.04 0.00 0.00 0.00 0.00 0.01 0.00 0.03 0.00 - 7 [ 0.01 .. 0.12]
18-> GLU 16 HA - PHE 17 HN [ 1.80 3.44] 0.10 0.08 0.10 0.08 0.11 0.09 0.11 0.11 0.08 0.10 0.11 0.10 0.11 0.09 0.10 0.11 0.09 0.10 0.10 0.11 - 20 [ 0.08 .. 0.11]
44-> GLU 35 HB3 - GLU 36 HN [ 1.79 3.91] 0.00 0.00 0.00 0.03 0.00 0.05 0.00 0.04 0.02 0.02 0.03 0.10 0.06 0.00 0.11 0.00 0.00 0.00 0.00 0.04 - 10 [ 0.02 .. 0.11]
57-> ASN 46 HN - LEU 47 HN [ 1.80 3.62] 0.00 0.00 0.00 0.00 0.00 0.00 0.00 0.00 0.00 0.00 0.00 0.00 0.00 0.00 0.00 0.00 0.14 0.00 0.00 0.00 - 1 [ 0.14 .. 0.14]
68-> LYS 55 HB2 - ASN 56 HN [ 1.80 3.88] 0.10 0.00 0.10 0.00 0.05 0.10 0.00 0.00 0.11 0.07 0.00 0.06 0.00 0.00 0.17 0.08 0.00 0.00 0.00 0.11 - 10 [ 0.05 .. 0.17]
69-> LYS 55 HB3 - ASN 56 HN [ 1.80 3.88] 0.00 0.00 0.00 0.00 0.00 0.00 0.00 0.06 0.00 0.00 0.00 0.00 0.00 0.00 0.00 0.00 0.00 0.00 0.18 0.00 - 2 [ 0.06 .. 0.18]
74-> TRP 60 HB2 - LYS 61 HN [ 1.80 4.20] 0.00 0.15 0.00 0.04 0.00 0.00 0.00 0.00 0.05 0.00 0.04 0.00 0.00 0.00 0.06 0.00 0.00 0.00 0.04 0.00 - 7 [ 0.00 .. 0.15]
76-> LYS 61 HA - SER 62 HN [ 1.79 3.41] 0.00 0.00 0.00 0.00 0.00 0.11 0.12 0.00 0.11 0.15 0.00 0.00 0.00 0.15 0.00 0.10 0.00 0.00 0.00 0.00 - 6 [ 0.10 .. 0.15]
145-> GLU 123 HN - HIS 124 HN [ 1.79 3.37] 0.00 0.01 0.00 0.00 0.00 0.00 0.00 0.00 0.03 0.00 0.00 0.04 0.00 0.00 0.00 0.05 0.00 0.00 0.13 0.00 - 5 [ 0.01 .. 0.13]
146-> GLU 123 HA - HIS 124 HN [ 1.80 3.30] 0.00 0.00 0.00 0.00 0.08 0.02 0.00 0.15 0.00 0.00 0.00 0.00 0.01 0.00 0.02 0.00 0.00 0.00 0.00 0.00 - 5 [ 0.01 .. 0.15]
160-> ASP 22 HN - ASP 22 HB2 [ 1.79 3.41] 0.00 0.00 0.00 0.00 0.00 0.00 0.00 0.00 0.00 0.00 0.12 0.00 0.00 0.00 0.00 0.00 0.00 0.00 0.00 0.00 - 1 [ 0.12 .. 0.12]
175-> LEU 47 HN - LEU 47 HB3 [ 1.79 3.55] 0.08 0.08 0.07 0.05 0.04 0.06 0.09 0.12 0.04 0.06 0.05 0.06 0.09 0.07 0.09 0.10 0.06 0.04 0.07 0.06 - 20 [ 0.04 .. 0.12]
176-> LYS 55 HN - LYS 55 HB3 [ 1.80 3.26] 0.00 0.00 0.00 0.16 0.00 0.00 0.00 0.25 0.00 0.00 0.00 0.00 0.00 0.07 0.00 0.00 0.00 0.18 0.23 0.00 - 5 [ 0.07 .. 0.25]
177-> LYS 57 HN - LYS 57 HB3 [ 1.79 3.23] 0.31 0.29 0.26 0.17 0.14 0.00 0.31 0.35 0.28 0.17 0.28 0.28 0.20 0.36 0.17 0.00 0.28 0.26 0.14 0.29 - 18 [ 0.14 .. 0.36]
219-> HIS 125 HN - HIS 125 HB2 [ 1.80 3.66] 0.00 0.00 0.00 0.00 0.00 0.00 0.00 0.00 0.11 0.00 0.00 0.00 0.00 0.00 0.00 0.00 0.01 0.00 0.00 0.00 - 2 [ 0.01 .. 0.11]
220-> HIS 125 HN - HIS 125 HB3 [ 1.80 3.66] 0.00 0.00 0.00 0.00 0.02 0.00 0.00 0.00 0.00 0.00 0.00 0.00 0.00 0.00 0.00 0.00 0.00 0.00 0.00 0.13 - 2 [ 0.02 .. 0.13]
222-> HIS 126 HN - HIS 126 HB3 [ 1.80 3.98] 0.00 0.00 0.00 0.00 0.00 0.00 0.00 0.00 0.00 0.00 0.00 0.00 0.07 0.00 0.00 0.00 0.10 0.00 0.00 0.00 - 2 [ 0.07 .. 0.10]
249-> ARG 59 HN - TRP 60 HN [ 1.79 4.27] 0.10 0.03 0.05 0.19 0.09 0.06 0.22 0.15 0.11 0.05 0.08 0.08 0.06 0.06 0.00 0.08 0.14 0.14 0.13 0.05 - 19 [ 0.03 .. 0.22]
264-> GLY 93 HN - PHE 94 HN [ 1.79 3.91] 0.00 0.00 0.02 0.00 0.25 0.00 0.00 0.06 0.04 0.00 0.00 0.00 0.00 0.00 0.00 0.00 0.13 0.00 0.00 0.00 - 5 [ 0.02 .. 0.25]
265-> HIS 95 HN - GLU 96 HN [ 1.80 3.80] 0.00 0.00 0.00 0.00 0.13 0.00 0.00 0.00 0.00 0.00 0.00 0.00 0.00 0.00 0.00 0.00 0.00 0.00 0.00 0.00 - 1 [ 0.13 .. 0.13]
266-> LEU 97 HN - SER 98 HN [ 1.80 4.02] 0.11 0.01 0.09 0.12 0.10 0.02 0.03 0.00 0.22 0.00 0.00 0.00 0.00 0.03 0.00 0.02 0.17 0.00 0.00 0.00 - 11 [ 0.01 .. 0.22]
292-> GLU 13 HA - PHE 17 HN [ 1.80 3.98] 0.10 0.04 0.00 0.02 0.01 0.06 0.09 0.00 0.04 0.06 0.00 0.00 0.00 0.02 0.01 0.03 0.06 0.02 0.07 0.04 - 15 [ 0.01 .. 0.10]
322-> GLU 63 HA - PHE 66 HN [ 1.79 3.95] 0.00 0.12 0.00 0.00 0.00 0.00 0.05 0.00 0.00 0.00 0.00 0.00 0.00 0.00 0.00 0.00 0.00 0.00 0.00 0.04 - 3 [ 0.04 .. 0.12]
379-> THR 5 HN - GLU 8 HN [ 1.80 4.74] 0.00 0.15 0.00 0.00 0.00 0.00 0.00 0.00 0.00 0.00 0.00 0.00 0.00 0.00 0.00 0.00 0.00 0.00 0.00 0.08 - 2 [ 0.08 .. 0.15]
380-> THR 5 HN - VAL 9 HN [ 1.79 4.81] 0.00 0.14 0.00 0.03 0.00 0.00 0.00 0.00 0.00 0.00 0.00 0.00 0.00 0.00 0.00 0.00 0.00 0.05 0.01 0.10 - 6 [ 0.00 .. 0.14]
389-> ILE 43 HA - ASN 46 HN [ 1.80 4.06] 0.00 0.00 0.00 0.00 0.00 0.17 0.00 0.00 0.00 0.00 0.00 0.00 0.00 0.00 0.00 0.00 0.10 0.00 0.00 0.00 - 2 [ 0.10 .. 0.17]
396-> LYS 61 HN - ASN 64 HN [ 1.80 5.10] 0.00 0.00 0.00 0.00 0.00 0.00 0.00 0.18 0.00 0.00 0.00 0.00 0.00 0.00 0.00 0.00 0.00 0.00 0.00 0.00 - 1 [ 0.18 .. 0.18]
402-> SER 98 HN - LEU 99 HN [ 1.80 3.30] 0.00 0.00 0.03 0.00 0.00 0.00 0.23 0.00 0.00 0.00 0.00 0.00 0.00 0.00 0.00 0.00 0.00 0.00 0.09 0.04 - 4 [ 0.03 .. 0.23]
412-> LYS 48 HA - ASN 52 HN [ 1.80 3.80] 0.02 0.00 0.00 0.11 0.00 0.02 0.00 0.02 0.00 0.00 0.02 0.00 0.00 0.00 0.00 0.00 0.00 0.00 0.00 0.05 - 6 [ 0.02 .. 0.11]
428-> GLU 8 HN - GLU 8 HG3 [ 1.80 3.98] 0.00 0.00 0.00 0.00 0.00 0.00 0.00 0.00 0.00 0.00 0.00 0.00 0.00 0.00 0.00 0.00 0.00 0.26 0.00 0.00 - 1 [ 0.26 .. 0.26]
432-> GLU 16 HN - GLU 16 HG2 [ 1.79 4.09] 0.00 0.00 0.00 0.00 0.00 0.27 0.00 0.00 0.00 0.00 0.00 0.00 0.00 0.00 0.00 0.00 0.00 0.00 0.00 0.00 - 1 [ 0.27 .. 0.27]
437-> LYS 32 HN - LYS 32 HG2 [ 1.79 4.31] 0.00 0.00 0.00 0.00 0.00 0.00 0.00 0.00 0.00 0.00 0.00 0.00 0.11 0.00 0.00 0.00 0.00 0.00 0.00 0.00 - 1 [ 0.11 .. 0.11]
438-> GLU 35 HN - GLU 35 HG2 [ 1.79 4.31] 0.09 0.11 0.00 0.00 0.00 0.00 0.00 0.00 0.00 0.00 0.00 0.00 0.00 0.08 0.00 0.04 0.08 0.07 0.00 0.00 - 6 [ 0.04 .. 0.11]
439-> GLU 35 HN - GLU 35 HG3 [ 1.79 4.31] 0.10 0.10 0.00 0.00 0.00 0.00 0.00 0.00 0.00 0.00 0.00 0.00 0.00 0.10 0.00 0.14 0.10 0.12 0.00 0.00 - 6 [ 0.10 .. 0.14]
440-> GLU 36 HN - GLU 36 HG2 [ 1.79 4.31] 0.00 0.00 0.00 0.00 0.00 0.00 0.00 0.18 0.00 0.00 0.00 0.00 0.00 0.00 0.00 0.00 0.00 0.00 0.00 0.00 - 1 [ 0.18 .. 0.18]
444-> LEU 40 HN - LEU 40 HG [ 1.80 3.66] 0.16 0.21 0.16 0.19 0.00 0.00 0.00 0.15 0.10 0.17 0.19 0.13 0.18 0.00 0.13 0.00 0.16 0.20 0.00 0.13 - 14 [ 0.10 .. 0.21]
445-> GLU 44 HN - GLU 44 HG2 [ 1.80 4.20] 0.00 0.00 0.00 0.00 0.00 0.00 0.00 0.10 0.00 0.00 0.00 0.00 0.00 0.08 0.01 0.00 0.00 0.17 0.00 0.00 - 4 [ 0.01 .. 0.17]
472-> LEU 99 HN - LEU 99 HG [ 1.80 3.70] 0.00 0.00 0.00 0.00 0.00 0.00 0.00 0.42 0.00 0.00 0.00 0.00 0.16 0.00 0.00 0.00 0.00 0.00 0.15 0.00 - 3 [ 0.15 .. 0.42]
473-> LYS 102 HN - LYS 102 HG3 [ 1.79 4.27] 0.15 0.01 0.03 0.14 0.10 0.09 0.06 0.05 0.06 0.11 0.08 0.11 0.04 0.14 0.02 0.16 0.13 0.08 0.12 0.08 - 20 [ 0.01 .. 0.16]
483-> LYS 113 HN - LYS 113 HG2 [ 1.80 3.98] 0.00 0.00 0.00 0.00 0.00 0.38 0.00 0.01 0.00 0.01 0.00 0.00 0.00 0.00 0.00 0.00 0.00 0.00 0.00 0.00 - 3 [ 0.01 .. 0.38]
485-> LEU 114 HN - LEU 114 HG [ 1.80 3.98] 0.03 0.30 0.28 0.00 0.19 0.01 0.02 0.04 0.03 0.00 0.00 0.03 0.26 0.04 0.03 0.33 0.03 0.33 0.00 0.32 - 16 [ 0.01 .. 0.33]
512-> ILE 50 HA - ASN 53 HD22 [ 1.79 5.21] 0.00 0.10 0.00 0.00 0.12 0.00 0.00 0.00 0.00 0.09 0.00 0.12 0.14 0.00 0.00 0.06 0.06 0.00 0.00 0.00 - 8 [ 0.00 .. 0.14]
564-> GLU 8 HN - VAL 119 HG1* [ 1.80 5.80] 0.06 0.00 0.05 0.01 0.00 0.02 0.12 0.01 0.06 0.00 0.09 0.04 0.10 0.03 0.05 0.00 0.02 0.00 0.04 0.09 - 15 [ 0.01 .. 0.12]
601-> SER 121 HN - LEU 122 HD1* [ 1.80 5.44] 0.15 0.00 0.00 0.00 0.00 0.03 0.00 0.12 0.00 0.00 0.00 0.00 0.00 0.00 0.00 0.00 0.00 0.00 0.00 0.00 - 3 [ 0.03 .. 0.15]
609-> GLU 12 HN - GLU 12 HB3 [ 1.80 3.16] 0.00 0.00 0.00 0.00 0.00 0.00 0.00 0.00 0.00 0.00 0.00 0.00 0.00 0.00 0.00 0.00 0.00 0.00 0.00 0.27 - 1 [ 0.27 .. 0.27]
611-> GLU 16 HA - GLU 16 HB2 [ 1.79 2.87] 0.00 0.10 0.13 0.11 0.11 0.00 0.00 0.00 0.11 0.08 0.12 0.13 0.11 0.04 0.10 0.11 0.09 0.08 0.13 0.11 - 16 [ 0.04 .. 0.13]
612-> GLU 16 HA - GLU 16 HB3 [ 1.79 2.87] 0.00 0.00 0.00 0.00 0.00 0.13 0.07 0.09 0.00 0.00 0.00 0.00 0.00 0.00 0.00 0.00 0.00 0.00 0.00 0.00 - 3 [ 0.07 .. 0.13]
614-> GLU 16 HN - GLU 16 HB3 [ 1.80 3.30] 0.00 0.18 0.24 0.18 0.18 0.00 0.00 0.00 0.19 0.15 0.20 0.22 0.20 0.07 0.18 0.20 0.18 0.15 0.22 0.20 - 16 [ 0.07 .. 0.24]
616-> ASP 22 HN - ASP 22 HB3 [ 1.79 3.41] 0.00 0.00 0.00 0.00 0.00 0.00 0.00 0.00 0.00 0.00 0.00 0.00 0.00 0.00 0.14 0.00 0.00 0.00 0.00 0.00 - 1 [ 0.14 .. 0.14]
623-> LEU 40 HN - LEU 40 HB3 [ 1.80 3.44] 0.02 0.01 0.03 0.00 0.09 0.08 0.12 0.02 0.05 0.03 0.01 0.02 0.03 0.07 0.05 0.12 0.02 0.01 0.07 0.04 - 19 [ 0.01 .. 0.12]
629-> ASN 46 HN - ASN 46 HA [ 1.80 2.62] 0.00 0.00 0.00 0.00 0.00 0.10 0.00 0.00 0.00 0.00 0.00 0.00 0.00 0.00 0.00 0.00 0.12 0.00 0.00 0.00 - 2 [ 0.10 .. 0.12]
635-> LYS 48 HN - LYS 48 HB3 [ 1.80 3.26] 0.00 0.00 0.00 0.00 0.00 0.00 0.12 0.00 0.00 0.25 0.00 0.00 0.00 0.00 0.00 0.00 0.26 0.25 0.00 0.00 - 4 [ 0.12 .. 0.26]
639-> GLU 49 HN - GLU 49 HB3 [ 1.80 3.48] 0.08 0.00 0.00 0.09 0.00 0.11 0.07 0.00 0.09 0.00 0.09 0.08 0.08 0.09 0.07 0.00 0.06 0.06 0.00 0.00 - 12 [ 0.06 .. 0.11]
642-> THR 51 HA - THR 51 HB [ 1.80 2.90] 0.00 0.00 0.00 0.00 0.00 0.09 0.00 0.00 0.00 0.10 0.00 0.00 0.00 0.00 0.00 0.10 0.10 0.00 0.00 0.00 - 4 [ 0.09 .. 0.10]
644-> ASN 53 HN - ASN 53 HB3 [ 1.79 3.37] 0.08 0.00 0.13 0.16 0.00 0.15 0.09 0.14 0.19 0.00 0.14 0.00 0.00 0.14 0.07 0.00 0.00 0.17 0.16 0.17 - 13 [ 0.07 .. 0.19]
647-> ASN 56 HN - ASN 56 HB2 [ 1.80 3.26] 0.00 0.00 0.00 0.00 0.00 0.00 0.13 0.00 0.00 0.00 0.00 0.00 0.00 0.00 0.00 0.00 0.00 0.00 0.00 0.00 - 1 [ 0.13 .. 0.13]
648-> ASN 56 HN - ASN 56 HB3 [ 1.80 3.26] 0.00 0.00 0.00 0.00 0.00 0.00 0.00 0.00 0.00 0.00 0.00 0.19 0.00 0.00 0.00 0.00 0.00 0.00 0.00 0.00 - 1 [ 0.19 .. 0.19]
661-> ASN 64 HN - ASN 64 HB3 [ 1.79 3.37] 0.00 0.16 0.00 0.00 0.00 0.15 0.00 0.14 0.18 0.16 0.00 0.00 0.16 0.16 0.00 0.16 0.00 0.14 0.18 0.17 - 11 [ 0.14 .. 0.18]
664-> LYS 70 HN - LYS 70 HB3 [ 1.79 3.41] 0.00 0.00 0.00 0.00 0.00 0.00 0.00 0.00 0.00 0.00 0.00 0.00 0.00 0.10 0.00 0.00 0.00 0.00 0.11 0.00 - 2 [ 0.10 .. 0.11]
671-> ASN 75 HN - ASN 75 HB3 [ 1.80 3.34] 0.00 0.00 0.00 0.00 0.00 0.00 0.19 0.00 0.19 0.20 0.00 0.00 0.20 0.00 0.00 0.00 0.00 0.00 0.00 0.00 - 4 [ 0.19 .. 0.20]
672-> THR 77 HN - THR 77 HB [ 1.80 3.48] 0.00 0.00 0.00 0.00 0.00 0.00 0.00 0.00 0.00 0.11 0.05 0.00 0.00 0.00 0.00 0.00 0.00 0.00 0.00 0.00 - 2 [ 0.05 .. 0.11]
677-> GLU 78 HN - GLU 78 HB3 [ 1.80 3.44] 0.14 0.15 0.13 0.13 0.12 0.14 0.14 0.14 0.14 0.12 0.12 0.13 0.13 0.10 0.11 0.12 0.15 0.15 0.13 0.13 - 20 [ 0.10 .. 0.15]
684-> PHE 94 HN - PHE 94 HB3 [ 1.80 3.70] 0.01 0.00 0.00 0.00 0.30 0.00 0.00 0.00 0.00 0.00 0.00 0.05 0.00 0.00 0.00 0.00 0.00 0.00 0.00 0.00 - 4 [ 0.00 .. 0.30]
693-> GLU 103 HN - GLU 103 HB3 [ 1.79 3.41] 0.16 0.16 0.16 0.14 0.16 0.12 0.00 0.13 0.14 0.15 0.00 0.00 0.17 0.00 0.00 0.00 0.11 0.10 0.00 0.15 - 13 [ 0.10 .. 0.17]
699-> LEU 114 HN - LEU 114 HB3 [ 1.80 3.26] 0.16 0.00 0.00 0.06 0.07 0.17 0.17 0.12 0.15 0.20 0.18 0.00 0.00 0.15 0.15 0.00 0.16 0.00 0.19 0.00 - 13 [ 0.06 .. 0.20]
703-> PHE 117 HN - PHE 117 HB3 [ 1.79 3.23] 0.00 0.00 0.00 0.22 0.00 0.00 0.00 0.00 0.08 0.00 0.00 0.00 0.00 0.08 0.00 0.00 0.13 0.00 0.00 0.00 - 4 [ 0.08 .. 0.22]
706-> SER 121 HN - SER 121 HB3 [ 1.80 3.44] 0.10 0.00 0.00 0.00 0.00 0.00 0.09 0.00 0.10 0.09 0.09 0.00 0.00 0.10 0.10 0.00 0.00 0.00 0.00 0.00 - 7 [ 0.09 .. 0.10]
710-> ILE 3 HA - SER 4 HN [ 1.80 2.44] 0.06 0.00 0.00 0.12 0.04 0.02 0.08 0.00 0.00 0.07 0.03 0.06 0.00 0.12 0.07 0.00 0.04 0.00 0.00 0.00 - 11 [ 0.02 .. 0.12]
711-> ILE 3 HB - SER 4 HN [ 1.79 3.23] 0.00 0.00 0.00 0.00 0.00 0.00 0.00 0.00 0.09 0.00 0.00 0.00 0.00 0.00 0.00 0.17 0.00 0.00 0.00 0.00 - 2 [ 0.09 .. 0.17]
732-> VAL 54 HB - LYS 55 HN [ 1.80 3.80] 0.13 0.00 0.00 0.00 0.14 0.00 0.00 0.00 0.00 0.00 0.00 0.00 0.00 0.00 0.14 0.00 0.05 0.00 0.00 0.00 - 4 [ 0.05 .. 0.14]
734-> ASN 56 HB2 - LYS 57 HN [ 1.80 3.84] 0.06 0.12 0.02 0.00 0.03 0.00 0.00 0.00 0.00 0.00 0.01 0.00 0.00 0.00 0.02 0.00 0.08 0.06 0.00 0.00 - 8 [ 0.01 .. 0.12]
736-> LYS 57 HA - GLY 58 HN [ 1.79 3.19] 0.00 0.00 0.00 0.11 0.17 0.17 0.00 0.00 0.00 0.00 0.00 0.00 0.00 0.00 0.00 0.17 0.00 0.00 0.04 0.00 - 5 [ 0.04 .. 0.17]
737-> GLY 58 HA2 - ARG 59 HN [ 1.79 3.41] 0.00 0.00 0.00 0.10 0.08 0.00 0.00 0.00 0.15 0.00 0.00 0.04 0.00 0.00 0.00 0.06 0.06 0.00 0.11 0.00 - 8 [ 0.00 .. 0.15]
738-> GLY 58 HA3 - ARG 59 HN [ 1.79 3.41] 0.00 0.00 0.00 0.00 0.00 0.00 0.12 0.00 0.00 0.00 0.00 0.00 0.00 0.00 0.02 0.00 0.00 0.00 0.00 0.00 - 2 [ 0.02 .. 0.12]
742-> TRP 60 HA - LYS 61 HN [ 1.79 3.01] 0.00 0.00 0.00 0.00 0.00 0.00 0.00 0.18 0.00 0.00 0.00 0.00 0.00 0.00 0.00 0.00 0.00 0.00 0.00 0.00 - 1 [ 0.18 .. 0.18]
746-> LYS 67 HB2 - ALA 68 HN [ 1.80 3.26] 0.04 0.00 0.03 0.11 0.07 0.22 0.05 0.23 0.20 0.19 0.20 0.04 0.22 0.01 0.16 0.06 0.21 0.21 0.11 0.11 - 19 [ 0.01 .. 0.23]
749-> LYS 70 HB2 - LEU 71 HN [ 1.79 3.37] 0.00 0.18 0.17 0.00 0.11 0.00 0.17 0.19 0.10 0.09 0.14 0.09 0.14 0.00 0.21 0.13 0.09 0.12 0.00 0.10 - 15 [ 0.09 .. 0.21]
761-> PHE 94 HB2 - HIS 95 HN [ 1.80 4.34] 0.10 0.00 0.00 0.00 0.00 0.00 0.05 0.00 0.00 0.00 0.00 0.13 0.00 0.00 0.00 0.00 0.04 0.00 0.00 0.00 - 4 [ 0.04 .. 0.13]
765-> ASN 100 HB2 - GLU 101 HN [ 1.80 3.62] 0.09 0.15 0.00 0.00 0.00 0.00 0.05 0.10 0.11 0.12 0.05 0.00 0.14 0.10 0.00 0.14 0.00 0.05 0.01 0.17 - 13 [ 0.01 .. 0.17]
766-> ASN 100 HB3 - GLU 101 HN [ 1.80 3.62] 0.00 0.00 0.00 0.00 0.03 0.00 0.00 0.00 0.00 0.00 0.00 0.18 0.00 0.00 0.05 0.00 0.00 0.00 0.00 0.00 - 3 [ 0.03 .. 0.18]
767-> GLU 101 HB2 - LYS 102 HN [ 1.79 3.73] 0.14 0.07 0.00 0.07 0.03 0.09 0.06 0.07 0.09 0.04 0.03 0.05 0.07 0.06 0.07 0.05 0.00 0.06 0.07 0.03 - 18 [ 0.03 .. 0.14]
769-> LYS 102 HB2 - GLU 103 HN [ 1.80 3.62] 0.09 0.00 0.11 0.05 0.13 0.14 0.13 0.10 0.11 0.13 0.10 0.00 0.00 0.09 0.09 0.07 0.02 0.10 0.05 0.01 - 18 [ 0.00 .. 0.14]
776-> LEU 122 HA - GLU 123 HN [ 1.79 3.41] 0.06 0.11 0.14 0.10 0.00 0.07 0.00 0.11 0.10 0.00 0.12 0.07 0.01 0.03 0.01 0.12 0.15 0.04 0.00 0.06 - 16 [ 0.01 .. 0.15]
788-> GLU 12 HA - GLU 15 HB3 [ 1.80 4.06] 0.09 0.06 0.12 0.17 0.09 0.02 0.07 0.03 0.08 0.09 0.15 0.10 0.10 0.15 0.05 0.03 0.08 0.07 0.00 0.13 - 20 [ 0.00 .. 0.17]
790-> GLU 13 HA - GLU 16 HB3 [ 1.79 3.55] 0.00 0.07 0.08 0.08 0.11 0.00 0.00 0.00 0.12 0.06 0.08 0.04 0.06 0.08 0.06 0.07 0.05 0.07 0.04 0.02 - 16 [ 0.02 .. 0.12]
802-> ALA 37 HA - LEU 40 HB3 [ 1.79 3.91] 0.09 0.08 0.06 0.18 0.07 0.10 0.01 0.11 0.06 0.04 0.09 0.10 0.03 0.11 0.09 0.04 0.11 0.07 0.16 0.06 - 20 [ 0.01 .. 0.18]
807-> LYS 67 HA - LYS 70 HB3 [ 1.80 4.02] 0.00 0.00 0.00 0.00 0.00 0.00 0.00 0.00 0.00 0.00 0.00 0.00 0.00 0.11 0.00 0.00 0.00 0.00 0.10 0.00 - 2 [ 0.10 .. 0.11]
815-> ALA 86 HA - LEU 89 HB2 [ 1.80 4.70] 0.00 0.00 0.00 0.00 0.00 0.00 0.00 0.00 0.00 0.12 0.00 0.13 0.00 0.00 0.00 0.00 0.00 0.00 0.00 0.00 - 2 [ 0.12 .. 0.13]
830-> GLU 109 HA - ARG 112 HB2 [ 1.80 3.44] 0.00 0.15 0.00 0.00 0.00 0.00 0.00 0.00 0.00 0.00 0.00 0.00 0.00 0.01 0.00 0.00 0.00 0.00 0.00 0.00 - 2 [ 0.01 .. 0.15]
837-> VAL 111 HA - LEU 114 HB3 [ 1.80 4.02] 0.02 0.00 0.00 0.06 0.04 0.04 0.00 0.10 0.05 0.00 0.07 0.00 0.00 0.04 0.04 0.00 0.05 0.00 0.05 0.00 - 12 [ 0.00 .. 0.10]
842-> VAL 115 HA - PHE 117 HN [ 1.79 3.95] 0.12 0.10 0.14 0.11 0.10 0.12 0.12 0.12 0.14 0.11 0.15 0.11 0.07 0.11 0.12 0.10 0.14 0.10 0.11 0.14 - 20 [ 0.07 .. 0.15]
847-> ALA 118 HA - SER 121 HB3 [ 1.80 3.80] 0.10 0.00 0.00 0.00 0.00 0.00 0.12 0.00 0.11 0.13 0.15 0.00 0.00 0.12 0.11 0.00 0.00 0.00 0.00 0.00 - 8 [ 0.00 .. 0.15]
856-> ILE 50 HA - ASN 53 HB3 [ 1.79 4.49] 0.00 0.00 0.02 0.00 0.00 0.00 0.16 0.00 0.01 0.00 0.02 0.00 0.00 0.02 0.00 0.00 0.00 0.03 0.00 0.00 - 6 [ 0.01 .. 0.16]
861-> ARG 73 HB2 - THR 77 HA [ 1.80 3.98] 0.06 0.02 0.00 0.00 0.00 0.02 0.00 0.00 0.00 0.04 0.02 0.06 0.04 0.07 0.00 0.02 0.11 0.05 0.00 0.00 - 12 [ 0.00 .. 0.11]
863-> ASN 120 HA - GLU 123 HB* [ 1.80 4.86] 0.12 0.00 0.00 0.00 0.00 0.00 0.00 0.00 0.00 0.00 0.00 0.00 0.00 0.00 0.00 0.00 0.00 0.00 0.00 0.00 - 1 [ 0.12 .. 0.12]
864-> VAL 54 HA - ASN 56 HN [ 1.80 3.44] 0.08 0.04 0.08 0.03 0.03 0.12 0.04 0.04 0.13 0.00 0.06 0.01 0.00 0.01 0.07 0.09 0.04 0.03 0.04 0.02 - 19 [ 0.00 .. 0.13]
865-> ILE 81 HA - LYS 84 HB2 [ 1.79 3.37] 0.00 0.00 0.00 0.00 0.00 0.00 0.00 0.00 0.00 0.00 0.00 0.00 0.10 0.00 0.00 0.03 0.00 0.00 0.00 0.00 - 2 [ 0.03 .. 0.10]
867-> ILE 3 HA - ILE 3 HG13 [ 1.80 3.98] 0.00 0.00 0.00 0.00 0.00 0.00 0.00 0.00 0.00 0.00 0.00 0.24 0.00 0.00 0.00 0.00 0.00 0.00 0.00 0.00 - 1 [ 0.24 .. 0.24]
877-> GLU 16 HN - GLU 16 HG3 [ 1.79 4.09] 0.06 0.00 0.00 0.00 0.00 0.13 0.22 0.19 0.00 0.00 0.00 0.00 0.00 0.00 0.00 0.00 0.00 0.00 0.00 0.00 - 4 [ 0.06 .. 0.22]
881-> GLN 25 HN - GLN 25 HG2 [ 1.80 3.48] 0.00 0.00 0.02 0.00 0.02 0.00 0.00 0.02 0.00 0.00 0.26 0.00 0.00 0.00 0.00 0.00 0.00 0.02 0.00 0.00 - 7 [ 0.00 .. 0.26]
882-> GLN 25 HA - GLN 25 HG2 [ 1.79 3.37] 0.00 0.00 0.00 0.00 0.00 0.00 0.00 0.00 0.00 0.00 0.16 0.00 0.00 0.00 0.00 0.00 0.00 0.00 0.00 0.00 - 1 [ 0.16 .. 0.16]
883-> GLN 25 HN - GLN 25 HG3 [ 1.80 3.48] 0.00 0.00 0.00 0.00 0.00 0.12 0.00 0.00 0.00 0.00 0.00 0.00 0.00 0.00 0.00 0.00 0.00 0.00 0.00 0.00 - 1 [ 0.12 .. 0.12]
884-> GLN 25 HA - GLN 25 HG3 [ 1.79 3.37] 0.00 0.00 0.00 0.00 0.00 0.25 0.00 0.00 0.00 0.00 0.00 0.00 0.00 0.00 0.00 0.00 0.00 0.00 0.00 0.00 - 1 [ 0.25 .. 0.25]
892-> LEU 40 HB3 - LEU 40 HG [ 1.80 2.90] 0.00 0.00 0.00 0.00 0.10 0.10 0.10 0.00 0.00 0.00 0.00 0.00 0.00 0.10 0.00 0.10 0.00 0.00 0.10 0.00 - 6 [ 0.10 .. 0.10]
899-> ILE 43 HN - ILE 43 HG13 [ 1.79 3.37] 0.11 0.13 0.00 0.00 0.00 0.00 0.00 0.12 0.00 0.00 0.08 0.00 0.00 0.10 0.12 0.11 0.00 0.09 0.10 0.11 - 10 [ 0.08 .. 0.13]
901-> GLU 44 HN - GLU 44 HG3 [ 1.80 4.20] 0.21 0.00 0.00 0.00 0.00 0.00 0.00 0.12 0.00 0.15 0.00 0.00 0.00 0.19 0.18 0.00 0.00 0.17 0.00 0.00 - 6 [ 0.12 .. 0.21]
914-> LYS 57 HN - LYS 57 HG2 [ 1.80 4.02] 0.00 0.00 0.00 0.00 0.00 0.12 0.00 0.00 0.00 0.00 0.00 0.00 0.08 0.00 0.00 0.01 0.00 0.00 0.09 0.04 - 5 [ 0.01 .. 0.12]
916-> LYS 57 HN - LYS 57 HG3 [ 1.80 4.02] 0.00 0.00 0.00 0.15 0.00 0.29 0.00 0.00 0.00 0.13 0.00 0.00 0.00 0.00 0.05 0.25 0.00 0.03 0.00 0.00 - 7 [ 0.00 .. 0.29]
948-> LEU 97 HN - LEU 97 HG [ 1.80 4.20] 0.00 0.18 0.00 0.00 0.00 0.01 0.00 0.01 0.00 0.02 0.00 0.07 0.00 0.00 0.00 0.00 0.00 0.00 0.00 0.00 - 5 [ 0.01 .. 0.18]
952-> LYS 102 HN - LYS 102 HG2 [ 1.79 4.27] 0.19 0.00 0.19 0.00 0.19 0.11 0.18 0.17 0.18 0.14 0.11 0.00 0.00 0.02 0.17 0.00 0.00 0.15 0.00 0.00 - 13 [ 0.00 .. 0.19]
959-> LYS 106 HA - LYS 106 HG3 [ 1.80 3.34] 0.00 0.04 0.00 0.00 0.00 0.00 0.00 0.00 0.00 0.00 0.00 0.00 0.00 0.00 0.18 0.13 0.00 0.00 0.00 0.19 - 4 [ 0.04 .. 0.19]
963-> LEU 114 HB3 - LEU 114 HG [ 1.80 2.58] 0.00 0.00 0.00 0.39 0.00 0.00 0.00 0.00 0.00 0.00 0.00 0.38 0.00 0.00 0.00 0.00 0.00 0.00 0.00 0.00 - 2 [ 0.38 .. 0.39]
970-> GLU 123 HN - GLU 123 HG3 [ 1.79 3.91] 0.00 0.00 0.27 0.00 0.00 0.00 0.02 0.00 0.00 0.00 0.00 0.00 0.00 0.00 0.00 0.00 0.23 0.00 0.00 0.00 - 3 [ 0.02 .. 0.27]
971-> GLU 123 HA - GLU 123 HG3 [ 1.79 3.37] 0.00 0.00 0.13 0.00 0.00 0.00 0.23 0.00 0.00 0.00 0.00 0.00 0.00 0.00 0.24 0.00 0.04 0.00 0.00 0.00 - 4 [ 0.04 .. 0.24]
973-> ILE 3 HG12 - SER 4 HN [ 1.80 4.34] 0.00 0.09 0.24 0.00 0.06 0.13 0.04 0.09 0.18 0.05 0.10 0.00 0.12 0.02 0.02 0.00 0.08 0.11 0.14 0.15 - 17 [ 0.00 .. 0.24]
974-> ILE 3 HG13 - SER 4 HN [ 1.80 4.34] 0.00 0.00 0.00 0.00 0.00 0.00 0.00 0.00 0.04 0.00 0.00 0.00 0.00 0.00 0.00 0.31 0.00 0.00 0.00 0.00 - 2 [ 0.04 .. 0.31]
977-> ARG 73 HD* - SER 74 HN [ 1.79 5.19] 0.00 0.00 0.00 0.00 0.00 0.00 0.00 0.00 0.00 0.00 0.00 0.00 0.11 0.00 0.00 0.00 0.00 0.00 0.00 0.07 - 2 [ 0.07 .. 0.11]
986-> LEU 41 HA - ASN 45 HD22 [ 1.79 5.57] 0.00 0.00 0.00 0.24 0.00 0.00 0.00 0.00 0.00 0.00 0.00 0.00 0.00 0.00 0.00 0.00 0.00 0.00 0.00 0.00 - 1 [ 0.24 .. 0.24]
995-> LEU 89 HG - LEU 107 HG [ 1.80 2.94] 0.00 0.08 0.06 0.05 0.05 0.03 0.00 0.02 0.02 0.11 0.00 0.17 0.00 0.01 0.06 0.00 0.01 0.06 0.00 0.10 - 14 [ 0.01 .. 0.17]
996-> GLU 44 HA - GLU 44 HG2 [ 1.80 3.84] 0.00 0.00 0.00 0.16 0.00 0.00 0.03 0.00 0.00 0.00 0.00 0.06 0.00 0.00 0.00 0.02 0.31 0.00 0.00 0.00 - 6 [ 0.00 .. 0.31]
1002-> LEU 41 HG - ALA 118 HA [ 1.80 4.06] 0.00 0.10 0.01 0.00 0.06 0.00 0.06 0.04 0.00 0.00 0.03 0.06 0.00 0.02 0.02 0.01 0.00 0.07 0.07 0.03 - 13 [ 0.01 .. 0.10]
1271-> ILE 116 HG2* - ASN 120 HD22 [ 1.79 4.57] 0.05 0.10 0.08 0.03 0.08 0.05 0.06 0.11 0.06 0.13 0.15 0.10 0.10 0.06 0.11 0.03 0.08 0.07 0.05 0.01 - 20 [ 0.01 .. 0.15]
1276-> LEU 41 HD1* - ALA 118 HA [ 1.79 3.85] 0.02 0.00 0.00 0.17 0.00 0.00 0.00 0.00 0.04 0.06 0.00 0.00 0.00 0.00 0.00 0.00 0.03 0.00 0.00 0.00 - 5 [ 0.02 .. 0.17]
1301-> THR 88 HB - LEU 107 HD2* [ 1.79 4.61] 0.03 0.03 0.00 0.00 0.00 0.00 0.00 0.04 0.00 0.14 0.06 0.00 0.03 0.00 0.01 0.00 0.00 0.00 0.00 0.01 - 8 [ 0.01 .. 0.14]
1323-> ILE 50 HN - THR 51 HG2* [ 1.80 5.36] 0.00 0.00 0.00 0.00 0.00 0.12 0.00 0.00 0.00 0.08 0.00 0.00 0.00 0.00 0.00 0.07 0.06 0.00 0.00 0.00 - 4 [ 0.06 .. 0.12]
1327-> TRP 83 HA - TRP 83 HZ3 [ 1.79 4.49] 0.07 0.09 0.04 0.05 0.05 0.01 0.07 0.08 0.16 0.11 0.08 0.02 0.04 0.10 0.05 0.05 0.11 0.03 0.12 0.04 - 20 [ 0.01 .. 0.16]
1329-> TRP 87 HB2 - TRP 87 HD1 [ 1.80 3.52] 0.14 0.04 0.11 0.14 0.13 0.07 0.08 0.16 0.11 0.16 0.06 0.07 0.11 0.07 0.12 0.03 0.02 0.12 0.09 0.09 - 20 [ 0.02 .. 0.16]
1367-> GLU 35 HA - TRP 83 HE3 [ 1.79 4.09] 0.05 0.09 0.00 0.00 0.12 0.00 0.05 0.07 0.13 0.09 0.09 0.02 0.01 0.11 0.04 0.10 0.11 0.04 0.09 0.02 - 18 [ 0.00 .. 0.13]
1418-> THR 51 HG2* - TRP 60 HZ2 [ 1.79 4.43] 0.00 0.00 0.00 0.00 0.00 0.03 0.14 0.00 0.00 0.00 0.00 0.00 0.00 0.00 0.00 0.00 0.00 0.04 0.00 0.00 - 3 [ 0.03 .. 0.14]
1453-> GLU 16 HN - GLU 16 HG* [ 1.80 3.66] 0.00 0.00 0.00 0.00 0.00 0.16 0.00 0.01 0.00 0.00 0.00 0.00 0.00 0.00 0.00 0.00 0.00 0.00 0.00 0.00 - 2 [ 0.01 .. 0.16]
1498-> GLU 35 HB* - TRP 83 HZ2 [ 1.79 4.43] 0.01 0.03 0.11 0.00 0.11 0.00 0.07 0.00 0.00 0.00 0.00 0.00 0.00 0.03 0.00 0.04 0.03 0.07 0.00 0.00 - 9 [ 0.01 .. 0.11]
-------------------------------------------
Number of Violations greater than 0.10 20 23 21 27 24 26 24 30 29 27 21 16 22 19 22 20 26 20 22 22
-------------------------------------------
---- Summary Of Residual Distance Constraint Violations ----
Mod 1 Mod 2 Mod 3 Mod 4 Mod 5 Mod 6 Mod 7 Mod 8 Mod 9 Mod 10 Mod 11 Mod 12 Mod 13 Mod 14 Mod 15 Mod 16 Mod 17 Mod 18 Mod 19 Mod 20 Averages
~~~~~~ ~~~~~~ ~~~~~~ ~~~~~~ ~~~~~~ ~~~~~~ ~~~~~~ ~~~~~~ ~~~~~~ ~~~~~~ ~~~~~~ ~~~~~~ ~~~~~~ ~~~~~~ ~~~~~~ ~~~~~~ ~~~~~~ ~~~~~~ ~~~~~~ ~~~~~~ ~~~~~~~~~~~
0.1 - 0.2 ang: 18 20 16 24 22 21 19 26 26 25 17 12 18 18 20 17 21 14 20 19 19.65
0.2 - 0.5 ang: 2 3 5 3 2 5 5 4 3 2 4 4 4 1 2 3 5 6 2 3 3.40
> 0.5 ang: 0 0 0 0 0 0 0 0 0 0 0 0 0 0 0 0 0 0 0 0 0.00
Total : 87 91 84 89 96 98 89 97 101 99 84 92 84 90 94 107 97 95 90 94 92.90
Minimum Violation : 0.000 0.000 0.000 0.000 0.000 0.000 0.000 0.000 0.000 0.000 0.000 0.000 0.000 0.000 0.000 0.000 0.000 0.000 0.000 0.000 0.000
Maximum Violation : 0.309 0.301 0.281 0.385 0.302 0.378 0.306 0.419 0.284 0.253 0.281 0.384 0.260 0.362 0.244 0.332 0.309 0.334 0.226 0.318 0.419
Max Intra Viol : 0.309 0.301 0.281 0.385 0.302 0.378 0.306 0.419 0.284 0.253 0.281 0.384 0.260 0.362 0.244 0.332 0.309 0.334 0.226 0.318 0.419
Max Seque Viol : 0.148 0.175 0.238 0.189 0.247 0.216 0.230 0.234 0.224 0.189 0.201 0.178 0.219 0.146 0.212 0.313 0.209 0.214 0.175 0.171 0.313
Max Medium Viol : 0.123 0.154 0.136 0.241 0.119 0.173 0.155 0.180 0.143 0.134 0.150 0.127 0.141 0.145 0.124 0.096 0.140 0.100 0.157 0.142 0.241
Max Long Viol : 0.070 0.105 0.107 0.168 0.119 0.067 0.143 0.068 0.125 0.135 0.088 0.169 0.102 0.106 0.059 0.096 0.109 0.074 0.088 0.101 0.169
Average Violation : 0.004 0.004 0.003 0.004 0.004 0.004 0.004 0.004 0.004 0.004 0.004 0.004 0.003 0.003 0.003 0.004 0.004 0.004 0.004 0.004 0.00382
Avge Intra Viol : 0.005 0.005 0.005 0.006 0.005 0.007 0.005 0.007 0.006 0.006 0.005 0.005 0.005 0.005 0.005 0.005 0.006 0.007 0.005 0.006 0.00567
Avge Seque Viol : 0.002 0.004 0.002 0.003 0.002 0.002 0.002 0.002 0.003 0.003 0.003 0.003 0.002 0.003 0.002 0.003 0.003 0.002 0.002 0.003 0.00261
Avge Mediu Viol : 0.004 0.003 0.004 0.004 0.005 0.005 0.005 0.006 0.007 0.004 0.004 0.004 0.004 0.003 0.004 0.006 0.005 0.003 0.005 0.004 0.00457
Avge Long Viol : 0.001 0.001 0.001 0.001 0.002 0.001 0.002 0.001 0.001 0.002 0.001 0.001 0.001 0.001 0.001 0.002 0.001 0.002 0.001 0.002 0.00139
RMS Violation : 0.020 0.021 0.021 0.023 0.020 0.024 0.022 0.026 0.023 0.021 0.021 0.021 0.020 0.019 0.020 0.022 0.023 0.023 0.020 0.022 0.02165
RMS Intra : 0.027 0.029 0.030 0.031 0.025 0.035 0.028 0.035 0.028 0.029 0.028 0.030 0.027 0.027 0.026 0.028 0.032 0.034 0.026 0.031 0.02941
RMS Sequential : 0.014 0.018 0.012 0.020 0.013 0.015 0.014 0.015 0.015 0.015 0.017 0.015 0.013 0.016 0.013 0.012 0.016 0.011 0.014 0.015 0.01475
RMS Medium range : 0.020 0.019 0.022 0.020 0.025 0.023 0.026 0.028 0.029 0.020 0.019 0.018 0.021 0.015 0.022 0.028 0.024 0.019 0.021 0.020 0.02232
RMS Long range : 0.008 0.010 0.009 0.011 0.012 0.006 0.013 0.006 0.009 0.013 0.009 0.011 0.006 0.008 0.007 0.011 0.009 0.009 0.010 0.010 0.00954
Final --global-- Summary for 20 models, 1681 NOEs/model, 33620 NOEs total
~~~~~~~~~~~~~~~~~~~~~~~~~~~~~~~~~~~~~~~~~~~~~~~~~~~~~~~~~~~~~~~~~~~~~~~~~~~
Summ of viol : 128.307
Summ sq. viol : 15.766
Maximum viol : 0.419
Average viol : 0.00382
RMSD viol : 0.02165
Std. Dev. viol : 0.02132
RMS Intra : 0.02941
RMS Seque : 0.01475
RMS Medi : 0.02232
RMS Long : 0.00954
table of dihedral angle constraints violations
2-> [ALA A 7] PSI -53.0 -25.0 0.0 1.4 0.0 0.0 0.0 0.2 0.0 0.0 0.5 0.0 0.0 0.2 0.7 0.0 0.0 0.0 0.2 0.0 0.0 2.4 - 7 [ 0.0 .. 2.4]
3-> [GLU A 8] PHI -75.0 -53.0 0.0 2.8 0.0 0.0 0.0 0.0 0.0 0.0 0.0 0.0 0.0 0.0 0.0 0.0 0.0 0.0 0.0 0.0 0.0 1.4 - 2 [ 0.0 .. 2.8]
7-> [TYR A 10] PHI -73.0 -51.0 0.0 0.0 0.0 0.0 0.0 0.6 0.0 0.0 1.0 0.0 0.0 0.0 0.0 0.0 0.0 0.0 0.0 0.0 0.0 0.0 - 2 [ 0.0 .. 1.0]
20-> [GLU A 16] PSI -53.0 -27.0 0.4 0.0 0.0 0.0 0.0 0.0 0.2 1.5 0.0 0.0 0.1 0.4 0.2 0.0 0.2 0.0 0.0 0.0 0.0 0.0 - 8 [ 0.0 .. 1.5]
22-> [PHE A 17] PSI -51.0 -29.0 0.0 0.2 0.0 0.0 0.0 0.2 0.0 0.0 0.0 0.0 0.0 1.6 0.1 0.0 0.0 0.0 0.5 0.0 0.0 0.0 - 5 [ 0.0 .. 1.6]
25-> [VAL A 24] PHI -77.0 -51.0 0.0 0.0 0.0 0.0 0.0 0.0 0.0 1.1 0.0 0.7 0.0 0.0 0.0 0.0 0.0 0.0 0.0 0.0 0.0 0.5 - 3 [ 0.0 .. 1.1]
26-> [VAL A 24] PSI -57.0 -33.0 0.0 0.0 0.0 0.0 0.0 0.0 0.0 1.9 0.0 0.2 1.5 0.0 0.0 0.0 0.0 0.3 0.0 0.0 0.0 0.0 - 4 [ 0.0 .. 1.9]
27-> [GLN A 25] PHI -71.0 -49.0 0.0 0.0 0.5 0.0 1.3 0.0 0.0 1.8 0.0 0.7 0.0 0.0 0.0 0.0 0.0 1.5 0.0 0.0 0.0 0.0 - 5 [ 0.0 .. 1.8]
28-> [GLN A 25] PSI -57.0 -29.0 0.7 0.0 0.0 0.0 0.7 0.0 0.0 0.0 0.0 1.5 0.0 0.0 0.0 0.3 0.3 0.6 1.6 2.2 1.4 0.0 - 9 [ 0.0 .. 2.2]
34-> [GLU A 28] PSI -57.0 -33.0 0.0 0.0 0.1 0.0 1.4 0.1 0.0 0.0 0.0 0.0 0.0 0.8 0.5 0.0 1.0 0.0 0.0 0.0 0.0 0.0 - 6 [ 0.0 .. 1.4]
42-> [LYS A 32] PSI -57.0 -29.0 0.2 0.0 1.0 0.4 0.0 0.0 0.0 0.6 0.0 0.1 1.0 1.1 0.0 0.0 0.0 2.1 0.0 0.0 0.0 1.1 - 9 [ 0.0 .. 2.1]
54-> [ILE A 38] PSI -49.0 -27.0 0.0 1.0 0.0 0.0 1.0 0.0 1.0 0.0 0.0 0.7 0.0 0.0 0.6 0.0 0.0 0.8 0.1 0.8 0.0 0.0 - 8 [ 0.0 .. 1.0]
56-> [LYS A 39] PSI -57.0 -33.0 0.0 0.0 0.0 0.2 0.7 2.0 1.7 0.0 0.0 0.0 0.0 0.0 0.0 0.0 0.0 2.0 0.0 0.0 0.0 0.0 - 6 [ 0.0 .. 2.0]
60-> [LEU A 41] PSI -59.0 -27.0 0.0 0.0 0.0 4.2 0.0 0.0 0.0 0.0 0.0 0.0 0.0 0.0 0.0 0.0 0.0 0.0 0.0 0.0 0.0 0.0 - 1 [ 0.0 .. 4.2]
63-> [ILE A 43] PHI -73.0 -47.0 0.0 0.0 0.0 1.3 0.0 1.6 0.0 0.0 0.0 0.0 0.0 0.5 0.6 0.0 0.0 0.0 1.4 0.0 0.0 0.0 - 5 [ 0.0 .. 1.6]
64-> [ILE A 43] PSI -51.0 -27.0 0.0 0.3 0.4 2.6 0.0 0.0 0.0 0.5 0.0 0.0 0.0 0.0 0.0 0.0 1.3 0.0 0.0 0.0 0.0 0.0 - 5 [ 0.0 .. 2.6]
66-> [GLU A 44] PSI -49.0 -15.0 1.1 0.0 0.0 0.0 0.0 0.0 0.0 0.0 0.0 0.0 0.0 0.0 0.0 2.5 0.0 0.0 0.0 0.0 0.0 0.0 - 3 [ 0.0 .. 2.5]
69-> [ILE A 50] PHI -99.0 -47.0 0.0 0.0 0.0 0.0 0.0 0.0 1.9 0.0 0.4 0.0 0.0 0.7 0.0 0.0 0.0 0.0 0.0 0.0 0.0 0.0 - 3 [ 0.0 .. 1.9]
74-> [ASN A 52] PSI -53.0 -19.0 0.0 0.0 0.0 0.0 0.2 0.0 0.0 0.0 0.0 0.1 0.0 1.0 0.0 0.0 0.0 1.0 0.4 0.0 0.0 0.0 - 5 [ 0.0 .. 1.0]
78-> [ASN A 64] PSI -55.0 -25.0 0.0 0.9 0.0 0.0 0.0 0.0 1.0 0.0 0.0 0.0 0.0 0.0 0.0 0.0 0.0 0.0 0.0 0.2 0.0 0.0 - 3 [ 0.0 .. 1.0]
80-> [LEU A 65] PSI -51.0 -29.0 0.0 1.1 0.0 0.0 0.0 0.0 0.6 0.0 0.0 0.0 0.0 0.0 0.0 0.0 0.0 0.0 0.0 0.0 0.0 0.4 - 3 [ 0.0 .. 1.1]
81-> [PHE A 66] PHI -75.0 -53.0 0.0 2.0 0.0 0.0 0.0 0.0 0.7 0.0 0.0 0.0 0.0 0.0 0.0 0.0 0.0 0.0 0.0 0.0 0.0 1.0 - 3 [ 0.0 .. 2.0]
84-> [LYS A 67] PSI -51.0 -27.0 0.0 0.0 0.0 0.0 0.0 0.0 0.0 1.0 0.0 0.0 0.0 0.0 0.0 0.0 0.0 0.0 0.0 1.8 0.0 0.2 - 3 [ 0.0 .. 1.8]
88-> [SER A 69] PSI -61.0 -27.0 0.3 0.0 1.4 0.0 0.0 0.0 0.0 0.0 1.3 2.0 0.0 0.0 0.0 0.0 0.0 0.0 0.0 0.0 0.4 0.0 - 5 [ 0.0 .. 2.0]
90-> [LYS A 70] PSI -65.0 -45.0 2.7 0.0 1.7 1.5 0.6 1.0 0.0 0.0 2.0 2.3 0.0 0.0 0.0 1.0 0.0 0.0 1.2 0.0 2.6 0.0 - 10 [ 0.0 .. 2.7]
110-> [LEU A 89] PSI -53.0 -25.0 0.0 0.0 0.0 0.0 0.0 0.0 0.0 0.0 0.2 1.0 0.0 0.9 0.0 0.0 0.0 0.0 0.0 0.0 0.0 0.0 - 3 [ 0.0 .. 1.0]
111-> [HIS A 90] PHI -85.0 -47.0 0.0 0.0 0.0 0.7 0.0 0.0 0.0 0.0 0.8 0.9 0.0 0.0 0.0 0.0 0.0 0.1 1.4 0.0 0.0 0.0 - 5 [ 0.0 .. 1.4]
113-> [VAL A 91] PHI -73.0 -51.0 0.0 0.0 0.0 1.5 0.2 0.4 1.0 1.8 0.0 0.0 0.0 1.7 0.0 1.1 0.8 0.0 0.0 0.0 0.0 0.0 - 8 [ 0.0 .. 1.8]
114-> [VAL A 91] PSI -57.0 -29.0 0.0 0.0 0.0 0.0 0.4 0.0 0.0 0.0 0.2 0.0 0.0 1.2 0.0 0.4 0.0 0.0 0.6 0.0 0.0 0.0 - 5 [ 0.0 .. 1.2]
116-> [GLU A 101] PSI -45.0 -23.0 0.0 0.0 0.0 0.0 0.0 0.0 0.0 0.0 0.7 0.0 0.0 0.0 0.0 1.7 0.0 0.0 0.0 0.1 0.2 0.0 - 4 [ 0.0 .. 1.7]
117-> [LYS A 102] PHI -77.0 -51.0 0.4 0.0 0.0 0.0 0.0 1.1 0.0 0.0 0.0 0.0 0.0 0.0 0.0 0.0 0.0 0.0 0.0 0.0 0.0 0.0 - 2 [ 0.0 .. 1.1]
118-> [LYS A 102] PSI -47.0 -25.0 0.3 0.0 0.0 0.0 0.0 0.2 0.0 0.2 1.4 2.2 0.2 0.0 0.0 0.0 0.0 0.0 0.0 0.0 0.0 0.0 - 6 [ 0.0 .. 2.2]
122-> [VAL A 104] PSI -55.0 -33.0 0.0 0.0 0.0 0.0 0.0 0.0 0.0 0.0 0.0 0.0 1.5 0.0 0.0 0.0 0.0 0.0 0.0 0.0 0.0 1.2 - 2 [ 0.0 .. 1.5]
125-> [LYS A 106] PHI -75.0 -53.0 0.0 0.3 0.0 0.8 0.0 1.2 0.0 0.0 0.0 0.0 0.0 0.0 0.0 0.0 1.7 0.0 1.0 0.2 0.0 0.0 - 6 [ 0.0 .. 1.7]
126-> [LYS A 106] PSI -57.0 -29.0 1.2 0.0 0.0 0.0 0.0 0.0 0.2 0.0 0.0 0.0 0.0 0.0 1.0 0.0 0.0 0.0 0.0 0.0 0.0 0.0 - 3 [ 0.0 .. 1.2]
128-> [LEU A 107] PSI -57.0 -35.0 0.0 0.4 0.0 0.0 0.0 0.0 0.2 0.0 0.8 0.1 0.9 0.0 1.2 1.6 1.2 0.0 0.4 0.0 1.2 0.6 - 11 [ 0.0 .. 1.6]
130-> [LYS A 108] PSI -57.0 -23.0 0.0 0.0 0.0 0.0 0.0 0.0 0.0 0.0 0.0 0.0 0.0 1.3 0.0 0.0 0.0 0.0 0.0 0.0 0.0 0.0 - 1 [ 0.0 .. 1.3]
131-> [GLU A 109] PHI -81.0 -47.0 0.0 0.0 0.0 2.3 0.0 2.4 0.0 0.0 0.6 2.1 0.0 0.0 0.3 0.0 0.9 0.0 0.9 0.0 0.0 0.0 - 7 [ 0.0 .. 2.4]
141-> [LEU A 114] PHI -77.0 -55.0 0.0 0.0 0.2 0.0 0.0 0.0 0.0 0.0 0.0 0.0 0.0 0.0 0.0 0.0 0.0 1.3 0.0 0.0 0.0 0.0 - 3 [ 0.0 .. 1.3]
150-> [ALA A 118] PSI -53.0 -27.0 0.0 0.0 0.0 0.0 0.0 1.0 0.0 0.0 0.0 0.0 0.0 0.0 0.0 0.0 0.0 0.0 0.0 0.0 0.0 0.0 - 1 [ 0.0 .. 1.0]
152-> [VAL A 119] PSI -55.0 -33.0 0.3 0.0 0.0 1.9 0.0 0.0 0.0 0.5 0.0 0.0 0.0 1.5 1.7 0.3 0.0 0.0 0.0 0.0 0.4 0.3 - 8 [ 0.0 .. 1.9]
178-> [ASN A 45] PSI -85.0 105.0 0.0 0.0 0.0 0.0 0.0 3.6 0.0 0.0 0.0 0.0 0.0 0.0 0.0 0.0 0.0 0.0 0.0 0.0 0.0 0.0 - 1 [ 0.0 .. 3.6]
180-> [ASN A 46] PHI 175.0 65.0 0.0 0.0 0.0 0.0 0.0 4.3 0.3 0.1 0.0 0.0 0.0 0.0 0.0 0.0 0.0 0.0 1.5 0.0 0.0 0.0 - 4 [ 0.0 .. 4.3]
199-> [LYS A 57] PSI -15.0 -165.0 0.0 0.0 0.0 0.0 0.0 1.4 0.0 0.0 0.0 0.0 0.0 0.0 0.0 0.0 0.0 1.2 0.0 0.0 0.0 0.0 - 2 [ 0.0 .. 1.4]
206-> [TRP A 60] PSI 5.0 -165.0 0.0 0.0 0.0 0.0 0.0 0.0 0.0 2.3 0.0 0.0 0.0 0.0 0.0 0.0 0.0 0.0 0.0 0.0 0.0 0.0 - 1 [ 0.0 .. 2.3]
210-> [SER A 62] PHI 45.0 -45.0 0.0 0.0 0.0 3.0 0.0 0.0 0.0 0.0 0.0 0.0 0.0 0.0 0.0 0.0 0.0 0.0 0.0 0.0 0.0 0.0 - 1 [ 0.0 .. 3.0]
215-> [LEU A 71] PSI 135.0 -35.0 0.0 1.5 0.0 0.5 0.1 0.0 1.2 0.0 0.0 0.0 0.6 1.6 1.3 0.6 0.3 1.7 0.9 1.1 0.0 0.7 - 13 [ 0.0 .. 1.7]
216-> [LEU A 71] PSI -85.0 -155.0 0.0 0.0 0.0 0.0 0.0 0.0 0.0 0.0 1.2 0.0 0.0 0.0 0.0 0.0 0.0 0.0 0.0 0.0 3.9 0.0 - 2 [ 0.0 .. 3.9]
218-> [LEU A 72] PHI -155.0 65.0 1.1 0.0 1.7 0.0 0.0 0.0 0.0 0.0 1.8 2.4 0.0 0.0 0.0 0.0 0.0 0.0 0.0 0.0 3.1 0.0 - 5 [ 0.0 .. 3.1]
239-> [GLY A 93] PSI -145.0 145.0 0.0 0.0 0.0 0.0 1.7 0.0 0.0 0.0 0.0 0.0 0.0 0.0 0.0 0.0 0.0 0.0 0.0 0.0 0.0 0.0 - 1 [ 0.0 .. 1.7]
246-> [HIS A 95] PSI -85.0 135.0 0.0 1.1 0.0 0.0 0.0 0.0 0.0 0.0 0.0 0.0 0.0 0.0 0.0 0.0 0.0 0.0 0.0 0.0 0.0 0.0 - 1 [ 0.0 .. 1.1]
265-> [GLU A 123] PHI 45.0 -45.0 0.0 0.0 1.9 0.0 0.0 0.0 0.0 0.0 0.0 0.0 0.0 0.0 0.0 0.0 0.0 0.0 0.3 0.0 0.0 0.0 - 2 [ 0.0 .. 1.9]
267-> [GLU A 123] PSI -35.0 105.0 0.0 0.0 0.0 0.0 0.0 0.0 0.0 1.1 0.0 0.0 0.0 0.0 0.0 0.0 0.0 0.0 0.0 0.0 0.0 0.0 - 1 [ 0.0 .. 1.1]
---- ACOSummary Of Residual ACO Constraint Violations ----
Mod 1 Mod 2 Mod 3 Mod 4 Mod 5 Mod 6 Mod 7 Mod 8 Mod 9 Mod 10 Mod 11 Mod 12 Mod 13 Mod 14 Mod 15 Mod 16 Mod 17 Mod 18 Mod 19 Mod 20 Averages
~~~~~~ ~~~~~~ ~~~~~~ ~~~~~~ ~~~~~~ ~~~~~~ ~~~~~~ ~~~~~~ ~~~~~~ ~~~~~~ ~~~~~~ ~~~~~~ ~~~~~~ ~~~~~~ ~~~~~~ ~~~~~~ ~~~~~~ ~~~~~~ ~~~~~~ ~~~~~~ ~~~~~~~~~~~
1 - 10. degrees : 4 7 5 8 4 9 6 8 6 7 3 8 4 4 3 6 5 3 5 4 5.45
> 10. degrees : 0 0 0 0 0 0 0 0 0 0 0 0 0 0 0 0 0 0 0 0 0.00
Total : 15 19 11 17 13 22 16 20 23 22 11 17 17 12 12 21 18 7 15 14 16.10
Minimum Violation : 0.0 0.0 0.0 0.0 0.0 0.0 0.0 0.0 0.0 0.0 0.0 0.0 0.0 0.0 0.0 0.0 0.0 0.0 0.0 0.0 0.00
Maximum Violation : 2.7 2.8 1.9 4.2 1.7 4.3 1.9 2.3 2.0 2.4 1.5 1.7 1.7 2.5 1.7 2.1 1.6 2.2 3.9 2.4 4.30
Max PHI Viol : 1.1 2.8 1.9 3.0 1.3 4.3 1.9 1.8 1.8 2.4 0.8 1.7 0.6 1.1 1.7 1.5 1.5 0.2 3.1 1.4 4.30
Max PSI Viol : 2.7 1.5 1.7 4.2 1.7 3.6 1.7 2.3 2.0 2.3 1.5 1.6 1.7 2.5 1.3 2.1 1.6 2.2 3.9 2.4 4.16
Average Violation : 0.0 0.1 0.0 0.1 0.0 0.1 0.0 0.1 0.1 0.1 0.0 0.1 0.0 0.0 0.0 0.1 0.1 0.0 0.1 0.0 0.049
Avge PHI Viol : 0.117 0.193 0.179 0.276 0.105 0.288 0.169 0.202 0.190 0.241 0.098 0.161 0.103 0.102 0.162 0.195 0.214 0.039 0.169 0.162 0.179
Avge PSI Viol : 0.261 0.268 0.194 0.308 0.237 0.304 0.248 0.297 0.285 0.290 0.218 0.303 0.243 0.267 0.192 0.307 0.242 0.219 0.293 0.247 0.264
RMS Violation : 0.224 0.277 0.222 0.433 0.184 0.438 0.222 0.293 0.252 0.337 0.169 0.260 0.180 0.236 0.180 0.283 0.235 0.188 0.365 0.227 0.271
RMS PHI Viol : 0.106 0.285 0.221 0.375 0.109 0.453 0.190 0.236 0.196 0.296 0.084 0.168 0.067 0.096 0.173 0.195 0.234 0.018 0.264 0.167 0.222
RMS PSI Viol : 0.309 0.267 0.223 0.491 0.243 0.419 0.253 0.348 0.304 0.378 0.232 0.336 0.254 0.330 0.186 0.358 0.236 0.275 0.454 0.280 0.319
Final --global-- Summary for 20 models, 273 ACOs/model, 5460 ACOs total
~~~~~~~~~~~~~~~~~~~~~~~~~~~~~~~~~~~~~~~~~~~~~~~~~~~~~~~~~~~~~~~~~~~~~~~~~~~
Summ. Viol. : 269.87
Summ. Sq. Viol. : 402.38
Max. Viol. : 4.297
Avg. Viol. : 0.04943
RMS Viol. : 0.27147
Std. Dev. Viol. : 0.26693
JPEG image for inter-residue distance constraints per residue plot
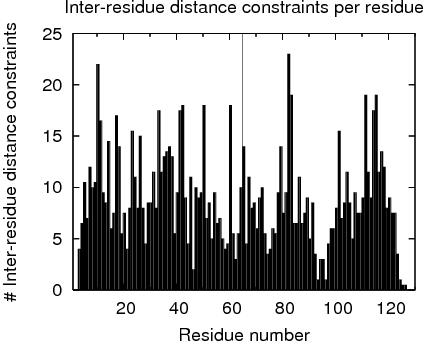
S(phi)|S(psi) V/S Residue number
Text output from PDBStat of phi psi order
# CHAIN .GT. SUM.GT.
# RES ID DIH S(phi) S(psi) S(chi1) S(chi2) S(chi3) S(chi4) S(chi5) 0.90 1.6
# -----------------------------------------------------------------------------------
MET A 1 0.326 0.421 0.148 0.785
SER A 2 0.882 0.310 0.663
ILE A 3 0.853 0.987 0.863 0.888 3
SER A 4 0.166 0.951 0.535
THR A 5 0.799 0.990 0.815
SER A 6 0.998 0.995 0.620 6 6
ALA A 7 0.999 0.998 7 7
GLU A 8 0.995 0.993 0.700 0.613 0.368 8 8
VAL A 9 0.998 0.999 0.999 9 9
TYR A 10 0.997 0.997 0.992 0.880 10 10
TYR A 11 0.999 0.999 0.991 0.995 11 11
GLU A 12 0.997 0.996 0.949 0.795 0.258 12 12
GLU A 13 0.999 0.997 0.997 0.782 0.490 13 13
ALA A 14 0.999 0.999 14 14
GLU A 15 0.999 0.997 0.993 0.786 0.419 15 15
GLU A 16 0.999 0.996 0.888 0.926 0.379 16 16
PHE A 17 0.998 0.998 0.996 0.910 17 17
LEU A 18 0.998 0.998 0.989 0.921 18 18
SER A 19 0.993 0.987 0.608 19 19
LYS A 20 0.987 0.975 0.991 0.481 0.590 0.451 20 20
GLY A 21 0.965 0.984 21 21
ASP A 22 0.983 0.959 0.838 0.537 22 22
LEU A 23 0.974 0.993 0.998 0.999 23 23
VAL A 24 0.995 0.997 0.999 24 24
GLN A 25 0.997 0.997 0.997 0.865 0.480 25 25
ALA A 26 0.994 0.996 26 26
CYS A 27 0.998 0.992 0.437 27 27
GLU A 28 0.997 0.996 0.597 0.338 0.467 28 28
LYS A 29 0.993 0.977 0.860 0.433 0.756 0.227 29 29
TYR A 30 0.993 0.978 0.287 0.455 30 30
TYR A 31 0.999 0.997 0.990 0.898 31 31
LYS A 32 0.997 0.992 0.952 0.502 0.574 0.454 32 32
ALA A 33 0.997 0.998 33 33
ALA A 34 0.996 0.998 34 34
GLU A 35 0.998 0.995 0.388 0.980 0.306 35 35
GLU A 36 0.998 0.994 0.929 0.872 0.691 36 36
ALA A 37 0.997 0.997 37 37
ILE A 38 0.999 0.999 0.998 1.000 38 38
LYS A 39 0.993 0.990 0.316 0.781 0.394 0.078 39 39
LEU A 40 0.997 0.998 0.987 0.669 40 40
LEU A 41 0.997 0.995 0.982 0.641 41 41
VAL A 42 0.994 0.995 0.998 42 42
ILE A 43 0.997 0.995 0.999 0.515 43 43
GLU A 44 0.995 0.986 0.068 0.317 0.305 44 44
ASN A 45 0.991 0.927 0.928 0.857 45 45
ASN A 46 0.865 0.931 0.994 0.422 46
LEU A 47 0.988 0.976 0.999 0.999 47 47
LYS A 48 0.996 0.985 0.788 0.753 0.679 0.642 48 48
GLU A 49 0.995 0.984 0.628 0.542 0.624 49 49
ILE A 50 0.988 0.995 0.996 0.679 50 50
THR A 51 0.995 0.990 0.644 51 51
ASN A 52 0.994 0.990 0.294 0.543 52 52
ASN A 53 0.992 0.984 0.725 0.554 53 53
VAL A 54 0.982 0.973 0.807 54 54
LYS A 55 0.994 0.990 0.836 0.391 0.778 0.425 55 55
ASN A 56 0.929 0.971 0.947 0.402 56 56
LYS A 57 0.984 0.444 0.918 0.746 0.472 0.568
GLY A 58 0.409 0.907
ARG A 59 0.948 0.979 0.970 0.518 0.485 0.811 0.999 59 59
TRP A 60 0.984 0.836 0.970 0.979 60
LYS A 61 0.798 0.429 0.363 0.666 0.506 0.626
SER A 62 0.658 0.966 0.616
GLU A 63 0.996 0.992 0.325 0.640 0.328 63 63
ASN A 64 0.993 0.994 0.629 0.494 64 64
LEU A 65 0.997 0.996 0.998 0.997 65 65
PHE A 66 0.998 0.995 0.623 0.713 66 66
LYS A 67 0.995 0.997 0.980 0.451 0.602 0.572 67 67
ALA A 68 0.996 0.996 68 68
SER A 69 0.997 0.999 0.520 69 69
LYS A 70 0.999 0.998 0.906 0.711 0.857 0.644 70 70
LEU A 71 0.991 0.660 0.999 0.999
LEU A 72 0.599 0.989 0.998 1.000
ARG A 73 0.998 0.997 0.989 0.756 0.533 0.734 0.998 73 73
SER A 74 0.994 0.989 0.397 74 74
ASN A 75 0.987 0.995 0.789 0.455 75 75
ASN A 76 0.987 0.988 0.940 0.544 76 76
THR A 77 0.993 0.993 0.438 77 77
GLU A 78 0.992 0.987 0.996 0.986 0.289 78 78
ILE A 79 0.999 0.999 0.996 0.509 79 79
PRO A 80 0.999 0.999 0.970 0.928 80 80
ILE A 81 0.998 0.997 0.999 0.924 81 81
LEU A 82 0.995 0.997 0.997 0.998 82 82
TRP A 83 0.999 0.997 0.998 0.999 83 83
LYS A 84 0.996 0.996 0.992 0.634 0.807 0.569 84 84
SER A 85 0.998 0.999 0.931 85 85
ALA A 86 0.999 0.999 86 86
TRP A 87 0.997 0.995 0.990 0.995 87 87
THR A 88 0.999 0.995 0.999 88 88
LEU A 89 0.998 0.993 0.861 0.866 89 89
HIS A 90 0.988 0.987 0.889 0.300 90 90
VAL A 91 0.997 0.986 0.999 91 91
GLU A 92 0.776 0.772 0.610 0.316 0.312
GLY A 93 0.741 0.427
PHE A 94 0.419 0.779 0.607 0.510
HIS A 95 0.417 0.609 0.741 0.185
GLU A 96 0.525 0.318 0.399 0.415 0.233
LEU A 97 0.974 0.977 0.995 0.998 97 97
SER A 98 0.587 0.757 0.648
LEU A 99 0.420 0.874 0.998 0.756
ASN A 100 0.919 0.985 0.785 0.231 100 100
GLU A 101 0.995 0.994 0.942 0.790 0.411 101 101
LYS A 102 0.999 0.997 0.972 0.475 0.772 0.631 102 102
GLU A 103 0.995 0.989 0.666 0.097 0.278 103 103
VAL A 104 0.997 0.995 0.998 104 104
LYS A 105 0.997 0.997 0.738 0.532 0.716 0.287 105 105
LYS A 106 0.995 0.995 0.642 0.800 0.226 0.374 106 106
LEU A 107 0.998 0.999 0.978 0.997 107 107
LYS A 108 0.998 0.994 0.465 0.655 0.305 0.464 108 108
GLU A 109 0.996 0.994 0.527 0.436 0.247 109 109
ASP A 110 0.995 0.993 0.920 0.595 110 110
VAL A 111 0.998 0.999 0.999 111 111
ARG A 112 0.998 0.998 0.954 0.924 0.445 0.726 0.999 112 112
LYS A 113 0.998 0.993 0.952 0.635 0.365 0.093 113 113
LEU A 114 0.995 0.998 0.765 0.861 114 114
VAL A 115 0.999 0.999 0.998 115 115
ILE A 116 0.999 0.999 0.999 0.769 116 116
PHE A 117 0.997 0.997 0.906 0.410 117 117
ALA A 118 0.998 0.998 118 118
VAL A 119 0.998 0.997 0.997 119 119
ASN A 120 0.996 0.991 0.993 0.891 120 120
SER A 121 0.995 0.991 0.647 121 121
LEU A 122 0.985 0.937 0.920 0.719 122 122
GLU A 123 0.640 0.850 0.978 0.776 0.349
HIS A 124 0.611 0.687 0.541 0.289
HIS A 125 0.518 0.673 0.632 0.393
HIS A 126 0.116 0.061 0.756 0.320
HIS A 127 0.519 0.688 0.381 0.429
HIS A 128 0.421 0.337 0.422 0.355
HIS A 129 0.868 0.266 0.113
JPEG image of S(phi)~Residue_number Plot
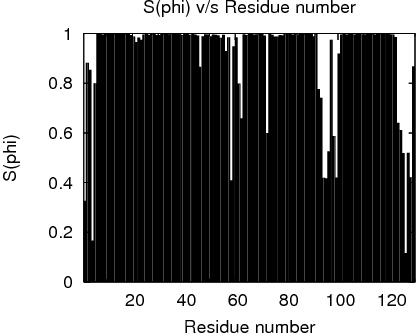
JPEG image of S(psi)~Residue_number Plot
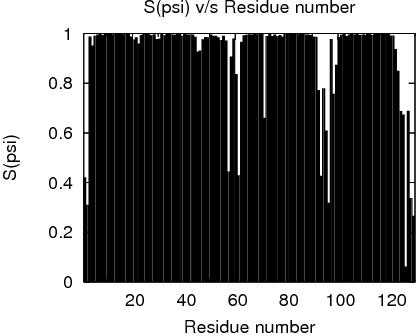
Table of Backbone and Heavy Atom RMSD
Text report of backbone and heavy atom RMSD for ordered regions
>
> Kabsch RMSD data for family `SSR10_NMR_em_bcr3.pdb'
>
> Kabsch RMSD of backbone atoms in res. A[6..45],A[47..56],A[63..70],A[73..91],A[100..122],for model 1 is: 0.636
> Kabsch RMSD of backbone atoms in res. A[6..45],A[47..56],A[63..70],A[73..91],A[100..122],for model 2 is: 0.760
> Kabsch RMSD of backbone atoms in res. A[6..45],A[47..56],A[63..70],A[73..91],A[100..122],for model 3 is: 0.809
> Kabsch RMSD of backbone atoms in res. A[6..45],A[47..56],A[63..70],A[73..91],A[100..122],for model 4 is: 0.764
> Kabsch RMSD of backbone atoms in res. A[6..45],A[47..56],A[63..70],A[73..91],A[100..122],for model 5 is: 0.586
> Kabsch RMSD of backbone atoms in res. A[6..45],A[47..56],A[63..70],A[73..91],A[100..122],for model 6 is: 0.786
> Kabsch RMSD of backbone atoms in res. A[6..45],A[47..56],A[63..70],A[73..91],A[100..122],for model 7 is: 0.926
> Kabsch RMSD of backbone atoms in res. A[6..45],A[47..56],A[63..70],A[73..91],A[100..122],for model 8 is: 0.737
> Kabsch RMSD of backbone atoms in res. A[6..45],A[47..56],A[63..70],A[73..91],A[100..122],for model 9 is: 1.123
> Kabsch RMSD of backbone atoms in res. A[6..45],A[47..56],A[63..70],A[73..91],A[100..122],for model 10 is: 0.779
> Kabsch RMSD of backbone atoms in res. A[6..45],A[47..56],A[63..70],A[73..91],A[100..122],for model 11 is: 0.660
> Kabsch RMSD of backbone atoms in res. A[6..45],A[47..56],A[63..70],A[73..91],A[100..122],for model 12 is: 0.663
> Kabsch RMSD of backbone atoms in res. A[6..45],A[47..56],A[63..70],A[73..91],A[100..122],for model 13 is: 0.656
> Kabsch RMSD of backbone atoms in res. A[6..45],A[47..56],A[63..70],A[73..91],A[100..122],for model 14 is: 0.689
> Kabsch RMSD of backbone atoms in res. A[6..45],A[47..56],A[63..70],A[73..91],A[100..122],for model 15 is: 0.584 (*)
> Kabsch RMSD of backbone atoms in res. A[6..45],A[47..56],A[63..70],A[73..91],A[100..122],for model 16 is: 0.686
> Kabsch RMSD of backbone atoms in res. A[6..45],A[47..56],A[63..70],A[73..91],A[100..122],for model 17 is: 0.774
> Kabsch RMSD of backbone atoms in res. A[6..45],A[47..56],A[63..70],A[73..91],A[100..122],for model 18 is: 0.738
> Kabsch RMSD of backbone atoms in res. A[6..45],A[47..56],A[63..70],A[73..91],A[100..122],for model 19 is: 0.819
> Kabsch RMSD of backbone atoms in res. A[6..45],A[47..56],A[63..70],A[73..91],A[100..122],for model 20 is: 0.618
>
> Kabsch RMSD statistics for 20 structures:
> Mean RMSD using as refer. str. `average' for res.[6..45],[47..56],[63..70],[73..91],[100..122], is: 0.740
> Range of RMSD values to reference struct. is 0.584 to 1.123
> Kabsch RMSD of heavy atoms in res. A[6..45],A[47..56],A[63..70],A[73..91],A[100..122],for model 1 is: 1.046
> Kabsch RMSD of heavy atoms in res. A[6..45],A[47..56],A[63..70],A[73..91],A[100..122],for model 2 is: 1.241
> Kabsch RMSD of heavy atoms in res. A[6..45],A[47..56],A[63..70],A[73..91],A[100..122],for model 3 is: 1.213
> Kabsch RMSD of heavy atoms in res. A[6..45],A[47..56],A[63..70],A[73..91],A[100..122],for model 4 is: 1.207
> Kabsch RMSD of heavy atoms in res. A[6..45],A[47..56],A[63..70],A[73..91],A[100..122],for model 5 is: 0.997
> Kabsch RMSD of heavy atoms in res. A[6..45],A[47..56],A[63..70],A[73..91],A[100..122],for model 6 is: 1.165
> Kabsch RMSD of heavy atoms in res. A[6..45],A[47..56],A[63..70],A[73..91],A[100..122],for model 7 is: 1.369
> Kabsch RMSD of heavy atoms in res. A[6..45],A[47..56],A[63..70],A[73..91],A[100..122],for model 8 is: 1.172
> Kabsch RMSD of heavy atoms in res. A[6..45],A[47..56],A[63..70],A[73..91],A[100..122],for model 9 is: 1.457
> Kabsch RMSD of heavy atoms in res. A[6..45],A[47..56],A[63..70],A[73..91],A[100..122],for model 10 is: 1.211
> Kabsch RMSD of heavy atoms in res. A[6..45],A[47..56],A[63..70],A[73..91],A[100..122],for model 11 is: 1.109
> Kabsch RMSD of heavy atoms in res. A[6..45],A[47..56],A[63..70],A[73..91],A[100..122],for model 12 is: 1.014
> Kabsch RMSD of heavy atoms in res. A[6..45],A[47..56],A[63..70],A[73..91],A[100..122],for model 13 is: 0.997 (*)
> Kabsch RMSD of heavy atoms in res. A[6..45],A[47..56],A[63..70],A[73..91],A[100..122],for model 14 is: 1.138
> Kabsch RMSD of heavy atoms in res. A[6..45],A[47..56],A[63..70],A[73..91],A[100..122],for model 15 is: 1.072
> Kabsch RMSD of heavy atoms in res. A[6..45],A[47..56],A[63..70],A[73..91],A[100..122],for model 16 is: 1.092
> Kabsch RMSD of heavy atoms in res. A[6..45],A[47..56],A[63..70],A[73..91],A[100..122],for model 17 is: 1.245
> Kabsch RMSD of heavy atoms in res. A[6..45],A[47..56],A[63..70],A[73..91],A[100..122],for model 18 is: 1.174
> Kabsch RMSD of heavy atoms in res. A[6..45],A[47..56],A[63..70],A[73..91],A[100..122],for model 19 is: 1.265
> Kabsch RMSD of heavy atoms in res. A[6..45],A[47..56],A[63..70],A[73..91],A[100..122],for model 20 is: 1.077
>
> Kabsch RMSD statistics for 20 structures:
> Mean RMSD using as refer. str. `average' for res.[6..45],[47..56],[63..70],[73..91],[100..122], is: 1.163
> Range of RMSD values to reference struct. is 0.997 to 1.457
Text report of backbone RMSD for entire protein
> Kabsch RMSD of backb atoms in res. *[1..129],for model 1 is: 1.783
> Kabsch RMSD of backb atoms in res. *[1..129],for model 2 is: 1.636
> Kabsch RMSD of backb atoms in res. *[1..129],for model 3 is: 2.399
> Kabsch RMSD of backb atoms in res. *[1..129],for model 4 is: 1.403
> Kabsch RMSD of backb atoms in res. *[1..129],for model 5 is: 1.453
> Kabsch RMSD of backb atoms in res. *[1..129],for model 6 is: 1.750
> Kabsch RMSD of backb atoms in res. *[1..129],for model 7 is: 1.790
> Kabsch RMSD of backb atoms in res. *[1..129],for model 8 is: 1.610
> Kabsch RMSD of backb atoms in res. *[1..129],for model 9 is: 1.928
> Kabsch RMSD of backb atoms in res. *[1..129],for model 10 is: 2.183
> Kabsch RMSD of backb atoms in res. *[1..129],for model 11 is: 1.924
> Kabsch RMSD of backb atoms in res. *[1..129],for model 12 is: 1.309 (*)
> Kabsch RMSD of backb atoms in res. *[1..129],for model 13 is: 1.484
> Kabsch RMSD of backb atoms in res. *[1..129],for model 14 is: 2.071
> Kabsch RMSD of backb atoms in res. *[1..129],for model 15 is: 1.594
> Kabsch RMSD of backb atoms in res. *[1..129],for model 16 is: 1.555
> Kabsch RMSD of backb atoms in res. *[1..129],for model 17 is: 2.199
> Kabsch RMSD of backb atoms in res. *[1..129],for model 18 is: 1.666
> Kabsch RMSD of backb atoms in res. *[1..129],for model 19 is: 1.690
> Kabsch RMSD of backb atoms in res. *[1..129],for model 20 is: 1.349
>
> Kabsch RMSD statistics for 20 structures:
> Mean RMSD using as refer. str. `average' for res.[1..129], is: 1.739
> Range of RMSD values to reference struct. is 1.309 to 2.399
Text report of heavy atom RMSD for entire protein
> Kabsch RMSD of heavy atoms in res. *[1..129],for model 1 is: 2.431
> Kabsch RMSD of heavy atoms in res. *[1..129],for model 2 is: 2.206
> Kabsch RMSD of heavy atoms in res. *[1..129],for model 3 is: 3.004
> Kabsch RMSD of heavy atoms in res. *[1..129],for model 4 is: 2.012
> Kabsch RMSD of heavy atoms in res. *[1..129],for model 5 is: 2.018
> Kabsch RMSD of heavy atoms in res. *[1..129],for model 6 is: 2.364
> Kabsch RMSD of heavy atoms in res. *[1..129],for model 7 is: 2.395
> Kabsch RMSD of heavy atoms in res. *[1..129],for model 8 is: 2.216
> Kabsch RMSD of heavy atoms in res. *[1..129],for model 9 is: 2.498
> Kabsch RMSD of heavy atoms in res. *[1..129],for model 10 is: 2.973
> Kabsch RMSD of heavy atoms in res. *[1..129],for model 11 is: 2.579
> Kabsch RMSD of heavy atoms in res. *[1..129],for model 12 is: 1.846 (*)
> Kabsch RMSD of heavy atoms in res. *[1..129],for model 13 is: 2.072
> Kabsch RMSD of heavy atoms in res. *[1..129],for model 14 is: 2.727
> Kabsch RMSD of heavy atoms in res. *[1..129],for model 15 is: 2.166
> Kabsch RMSD of heavy atoms in res. *[1..129],for model 16 is: 2.222
> Kabsch RMSD of heavy atoms in res. *[1..129],for model 17 is: 2.749
> Kabsch RMSD of heavy atoms in res. *[1..129],for model 18 is: 2.274
> Kabsch RMSD of heavy atoms in res. *[1..129],for model 19 is: 2.400
> Kabsch RMSD of heavy atoms in res. *[1..129],for model 20 is: 2.104
>
> Kabsch RMSD statistics for 20 structures:
> Mean RMSD using as refer. str. `average' for res.[1..129], is: 2.363
> Range of RMSD values to reference struct. is 1.846 to 3.004
Summary of heavy atom and backbone RMSDs over the whole protein and ordered residues
RMSD Values
all residues ordered residues selected residues
All backbone atoms 1.7 0.7 0.7
All heavy atoms 2.4 1.2 1.2
Contact Map (constraints list and 3D Coordinates)
JPEG image of Contact Map for Constraints
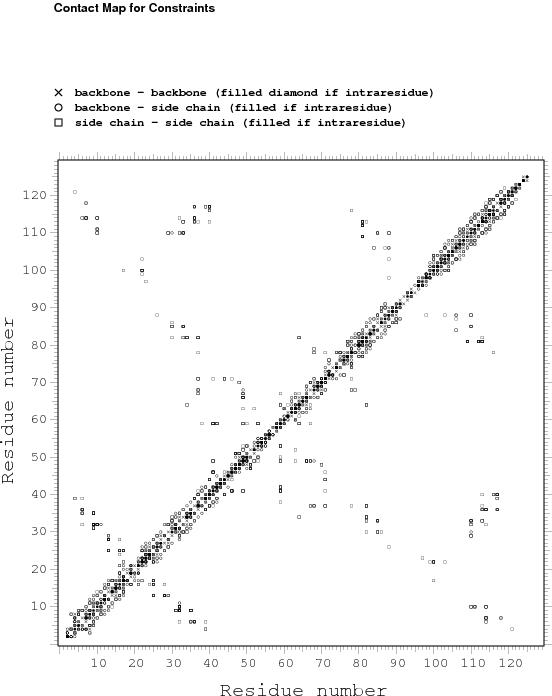
JPEG image of Contact Map for Coordinates
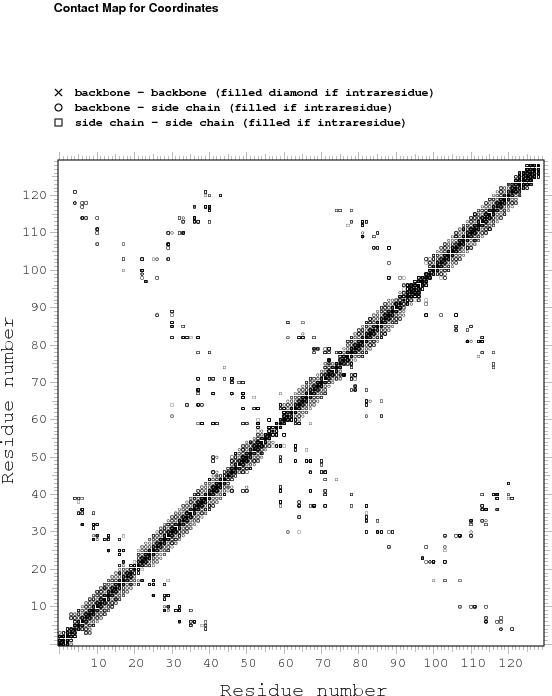
Output from PROCHECK
Ramachandran Plot for all models
Text summary of Ramachandran Plot
+----------<<< P R O C H E C K S U M M A R Y >>>----------+
| |
| SSR10_NMR_em_bcr3_020.rin 0.0 2000 residues |
| |
+| Ramachandran plot: 98.1% core 1.8% allow 0.1% gener 0.0% disall |
| |
+| All Ramachandrans: 6 labelled residues (out of2000) |
+| Chi1-chi2 plots: 14 labelled residues (out of1400) |
JPEG image for all model Ramachandran Plot
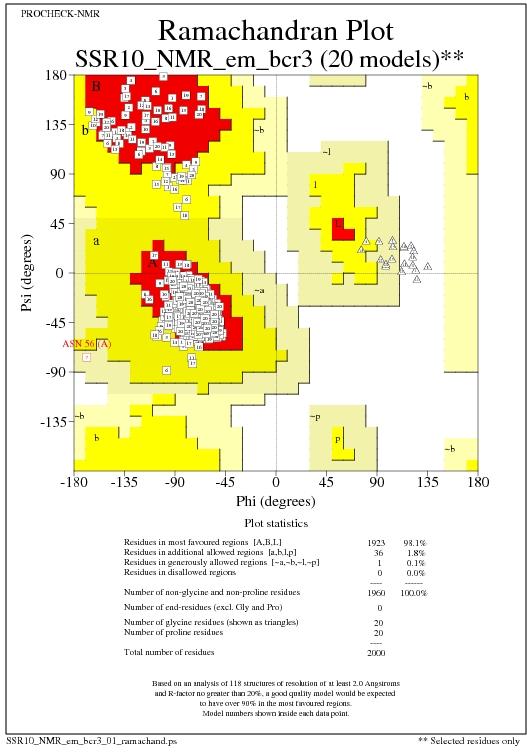
Residue Properties for all models
JPEG for all model Residue Properties - page $num_n

JPEG for all model Residue Properties - page $num_n
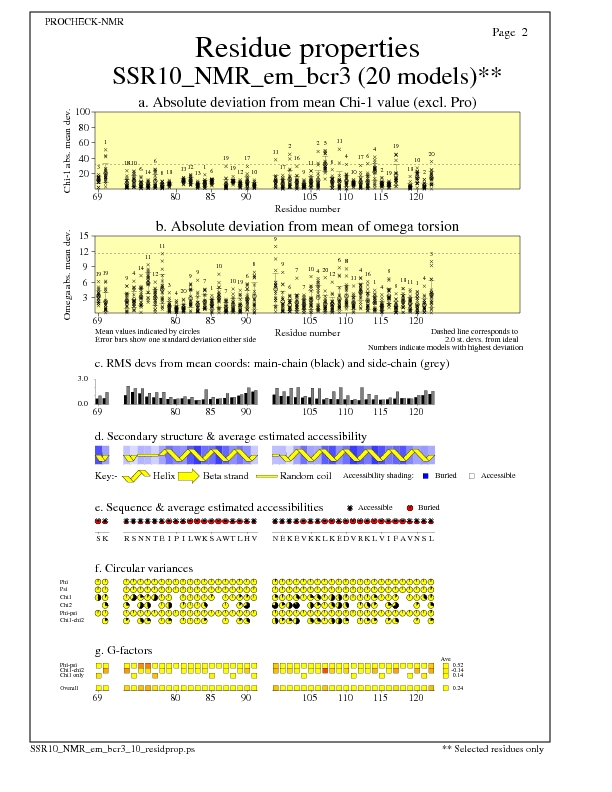

Model Secondary Structures from Procheck
JPEG for Model Secondary Structures - page $num_n

JPEG for Model Secondary Structures - page $num_n

JPEG for Model Secondary Structures - page $num_n

JPEG for Model Secondary Structures - page $num_n

JPEG for Model Secondary Structures - page $num_n

JPEG for Model Secondary Structures - page $num_n

Ramachandran Plots for each residue
JPEG for residue Ramachandran Plots - page $num_n
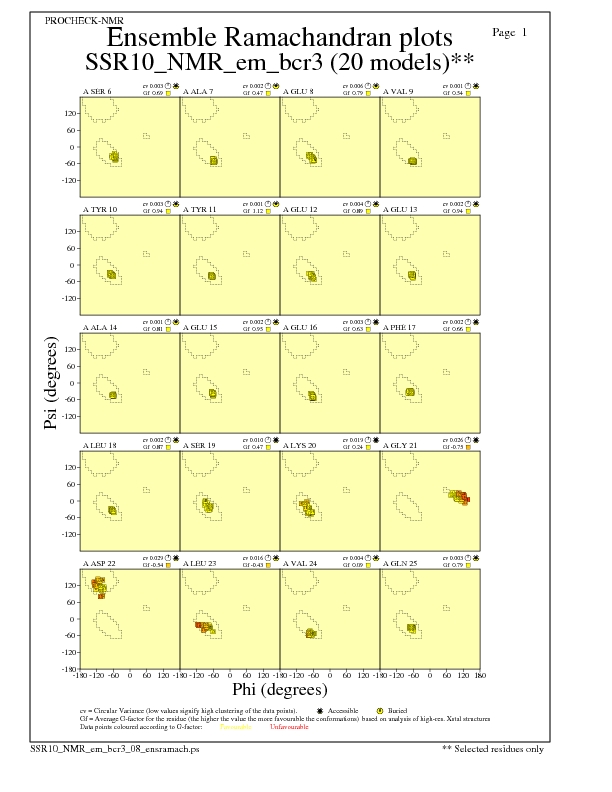
JPEG for residue Ramachandran Plots - page $num_n
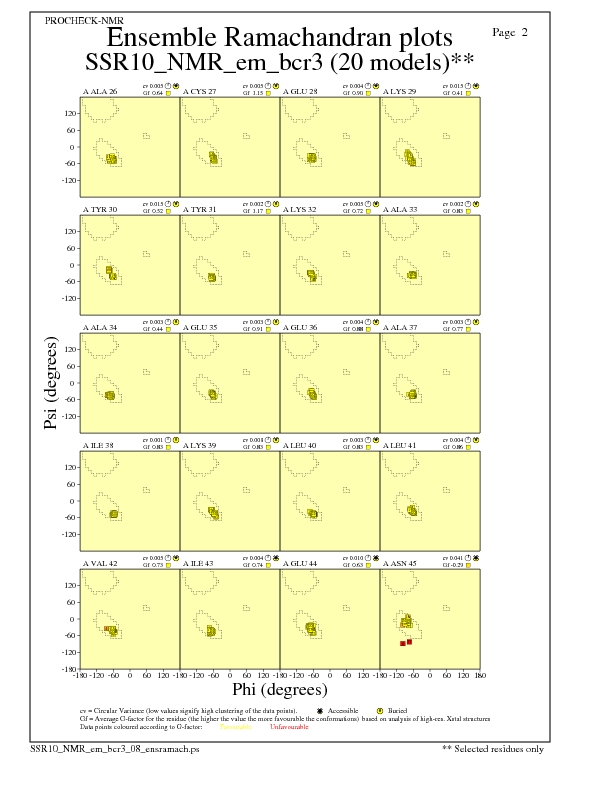
JPEG for residue Ramachandran Plots - page $num_n
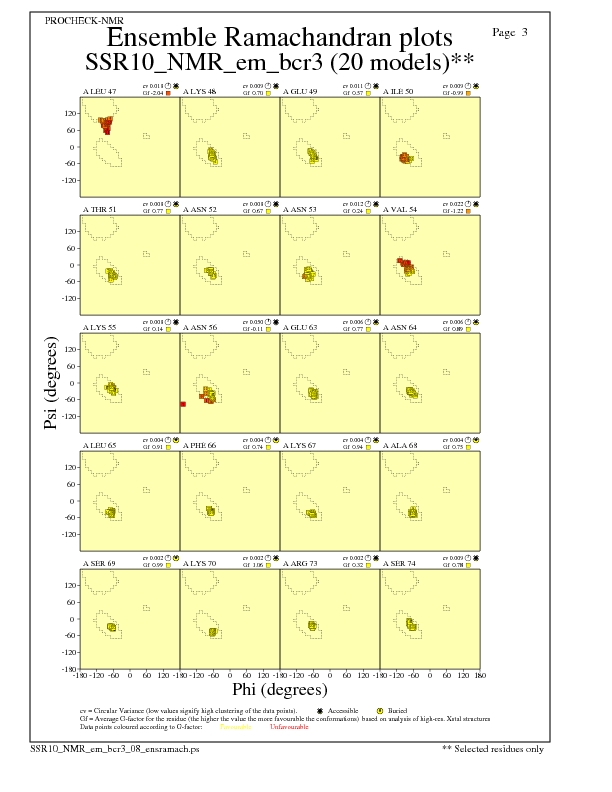
JPEG for residue Ramachandran Plots - page $num_n
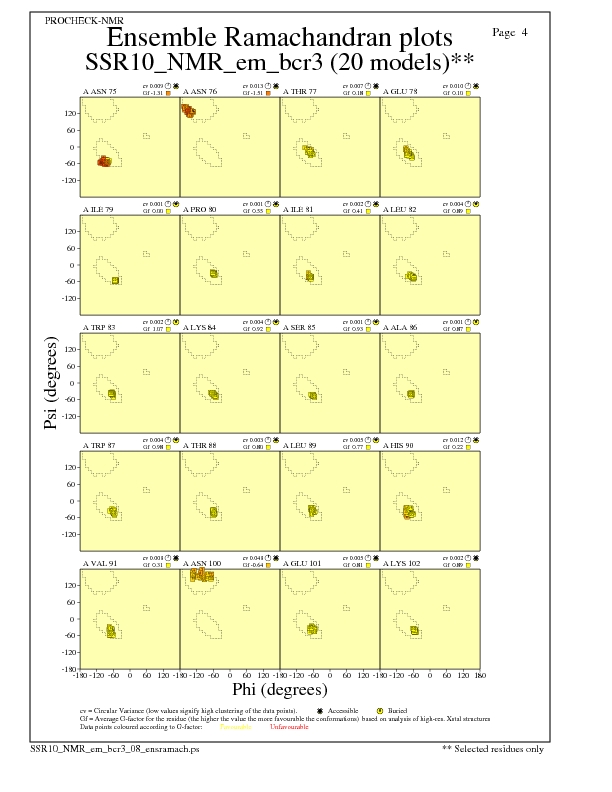
JPEG for residue Ramachandran Plots - page $num_n
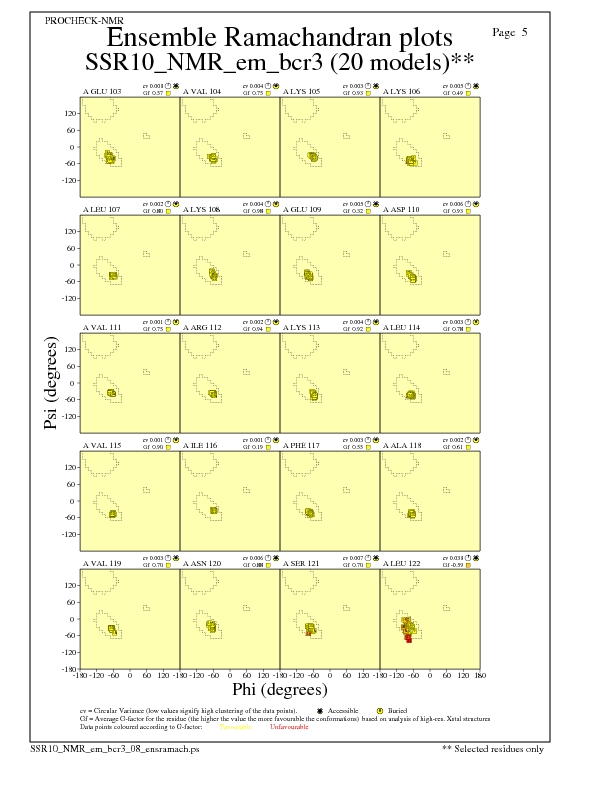
Ramachandran analysis for each residue from Molprobity
Chi1-Chi2 Plots for each residue
JPEG for residue Chi1-Chi2 Plots - page $num_n
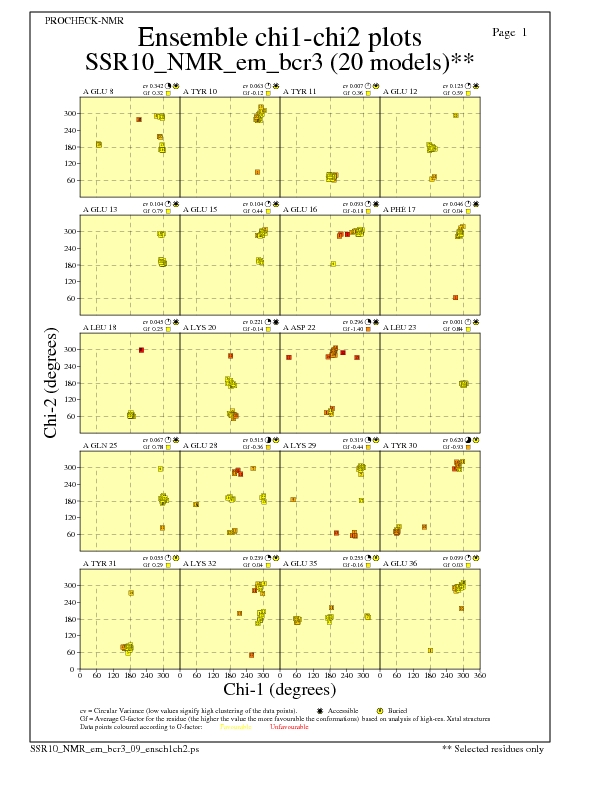
JPEG for residue Chi1-Chi2 Plots - page $num_n
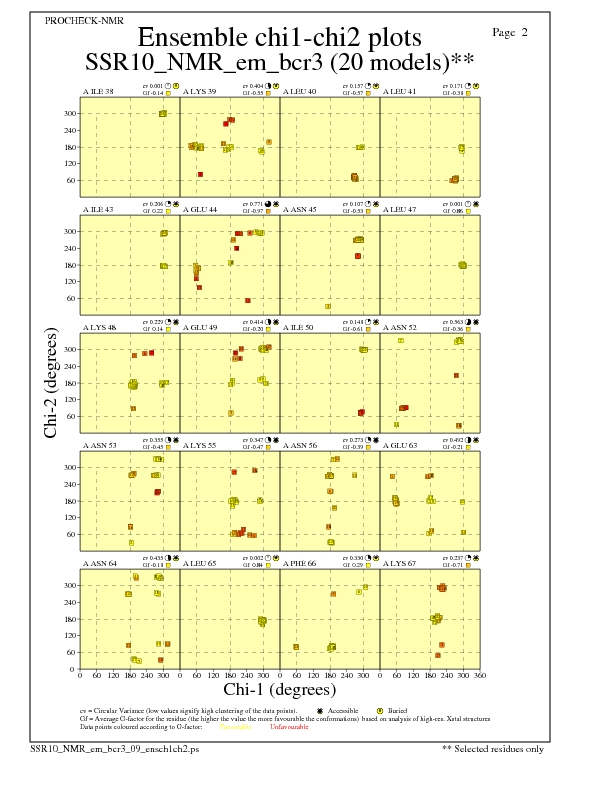
JPEG for residue Chi1-Chi2 Plots - page $num_n
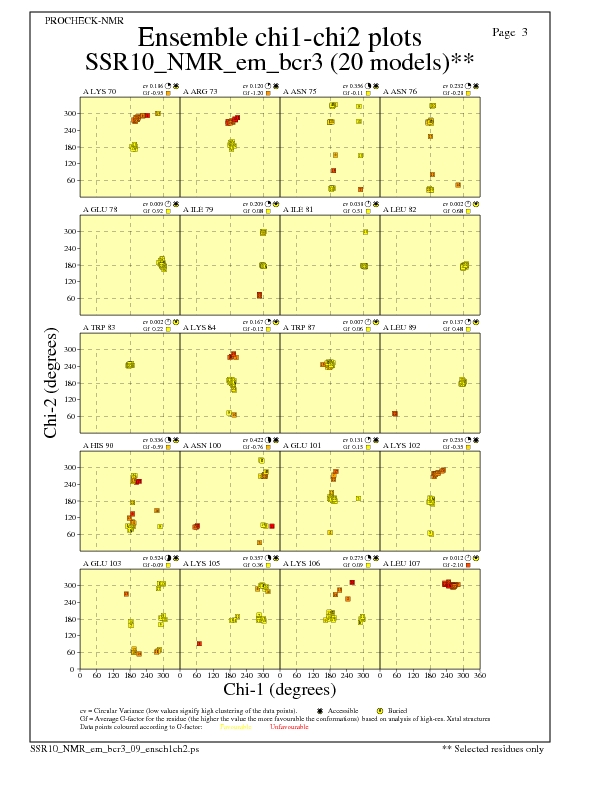
JPEG for residue Chi1-Chi2 Plots - page $num_n
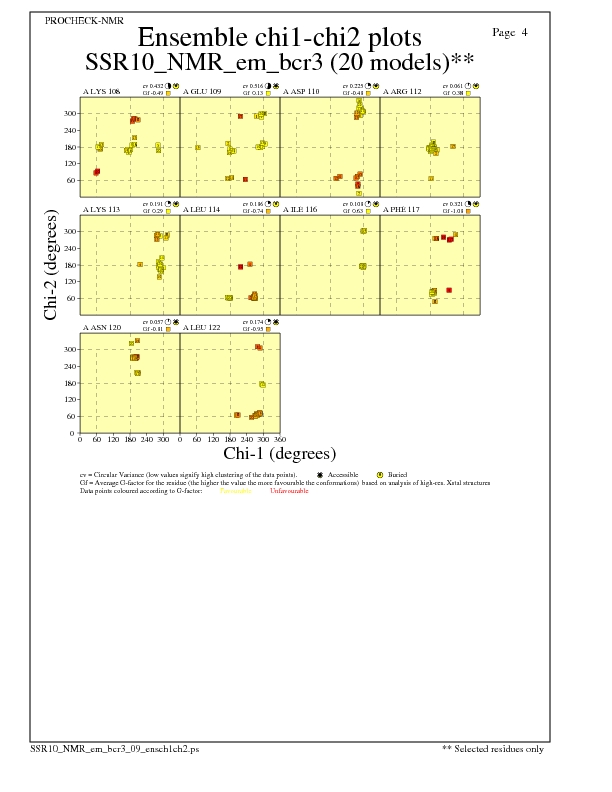
Procheck G-factors for phi-psi for each residue
JPEG image for residue phi-psi G-factors
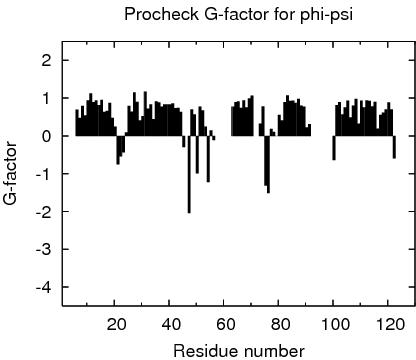
Table of Procheck G-factors for phi-psi for ordered residues
#phipsi_gfactor
#Residue\Model average
6 0.69
7 0.47
8 0.79
9 0.54
10 0.94
11 1.12
12 0.89
13 0.94
14 0.81
15 0.95
16 0.63
17 0.66
18 0.87
19 0.47
20 0.24
21 -0.75
22 -0.54
23 -0.43
24 0.09
25 0.79
26 0.64
27 1.15
28 0.90
29 0.41
30 0.52
31 1.17
32 0.72
33 0.83
34 0.44
35 0.91
36 0.88
37 0.77
38 0.83
39 0.83
40 0.83
41 0.86
42 0.73
43 0.74
44 0.63
45 -0.29
47 -2.04
48 0.70
49 0.57
50 -0.99
51 0.77
52 0.67
53 0.24
54 -1.22
55 0.14
56 -0.11
63 0.77
64 0.89
65 0.91
66 0.74
67 0.94
68 0.75
69 0.99
70 1.06
73 0.32
74 0.78
75 -1.31
76 -1.51
77 0.18
78 0.10
79 0.00
80 0.55
81 0.41
82 0.89
83 1.07
84 0.92
85 0.93
86 0.87
87 0.98
88 0.80
89 0.77
90 0.22
91 0.31
100 -0.64
101 0.81
102 0.89
103 0.57
104 0.75
105 0.93
106 0.49
107 0.80
108 0.98
109 0.32
110 0.93
111 0.75
112 0.94
113 0.92
114 0.78
115 0.90
116 0.19
117 0.55
118 0.61
119 0.70
120 0.88
121 0.70
122 -0.59
#Reported_Model_Average 0.519
#Overall_Average_Reported 0.519
Procheck G-factors for all dihedral angles for each residue
JPEG image for residue all dihedral G-factors
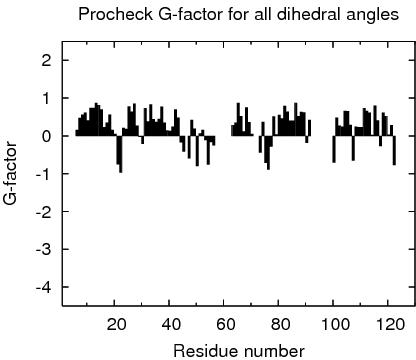
Table of Procheck G-factors for all dihedrals for ordered residues
#alldih_gfactor
#Residue\Model average
6 0.16
7 0.47
8 0.56
9 0.61
10 0.41
11 0.74
12 0.74
13 0.87
14 0.81
15 0.70
16 0.23
17 0.35
18 0.56
19 0.16
20 0.05
21 -0.75
22 -0.97
23 0.21
24 0.18
25 0.78
26 0.64
27 0.85
28 0.27
29 -0.02
30 -0.21
31 0.73
32 0.38
33 0.83
34 0.44
35 0.37
36 0.45
37 0.77
38 0.35
39 0.14
40 0.13
41 0.24
42 0.70
43 0.48
44 -0.17
45 -0.41
47 -0.59
48 0.42
49 0.19
50 -0.80
51 0.06
52 0.16
53 -0.11
54 -0.76
55 -0.16
56 -0.25
63 0.28
64 0.35
65 0.87
66 0.52
67 0.12
68 0.75
69 0.36
70 0.05
73 -0.44
74 0.37
75 -0.71
76 -0.89
77 -0.28
78 0.51
79 0.04
80 0.55
81 0.46
82 0.79
83 0.64
84 0.40
85 0.40
86 0.87
87 0.52
88 0.63
89 0.62
90 -0.18
91 0.42
100 -0.70
101 0.48
102 0.27
103 0.24
104 0.66
105 0.65
106 0.29
107 -0.65
108 0.24
109 0.23
110 0.23
111 0.73
112 0.66
113 0.61
114 0.02
115 0.80
116 0.41
117 -0.27
118 0.61
119 0.52
120 0.03
121 0.28
122 -0.77
#Reported_Model_Average 0.256
#Overall_Average_Reported 0.256
Output from Verify3D
Verify3D Score over a window of $winsize_s residues
JPEG image for Verify3D Score
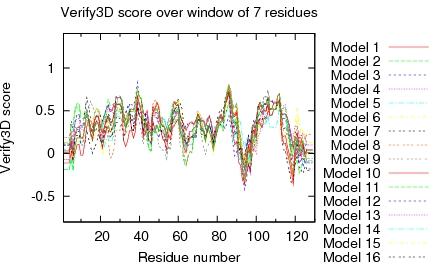
Table of Verify3D scores for ordered residues across all models
#verify3d
#Residue\Model 1 2 3 4 5 6 7 8 9 10 11 12 13 14 15 16 17 18 19 20
6 -0.38 0.47 0.47 0.47 -0.38 0.47 0.47 -0.38 0.47 0.47 -0.38 0.47 0.47 0.47 -0.38 0.47 0.47 0.47 0.47 0.47
7 0.76 0.76 0.76 0.76 0.76 0.76 0.76 0.76 0.76 0.76 0.76 0.76 0.76 0.76 0.76 0.76 0.76 0.76 0.76 0.76
8 -0.43 0.62 -0.43 0.62 -0.43 -0.43 -0.58 -0.43 -0.43 -0.43 -0.43 0.62 -0.43 -0.43 -0.43 -0.43 0.09 0.62 -0.43 0.62
9 -0.62 0.30 -0.62 -0.62 -0.62 -0.62 -0.62 -0.62 -0.62 -0.62 -0.62 -0.62 -0.62 -0.62 -0.62 -0.62 -0.62 -0.62 -0.62 0.30
10 0.50 0.50 0.50 0.50 0.50 0.50 0.50 0.50 0.50 0.50 0.50 0.50 0.50 0.50 0.50 0.50 0.50 0.50 -0.88 0.50
11 0.86 0.86 0.50 0.50 0.50 0.50 0.86 0.86 0.86 0.86 0.86 0.86 0.86 0.86 0.86 0.27 0.50 0.86 0.86 0.50
12 0.60 0.60 0.62 0.62 0.62 0.60 0.60 0.60 0.60 0.60 0.62 0.62 0.60 0.60 0.60 0.62 0.62 0.62 0.62 0.62
13 -0.43 -0.43 -0.43 -0.43 -0.43 -0.43 0.62 -0.43 0.62 -0.43 -0.43 -0.43 -0.43 -0.43 -0.43 -0.43 -0.43 -0.43 -0.43 -0.43
14 0.76 0.76 0.76 0.76 0.76 0.76 0.76 0.76 0.76 0.76 0.76 0.76 0.76 0.76 0.76 0.76 0.76 0.76 0.76 0.76
15 0.62 0.62 0.62 0.62 0.62 0.62 0.62 0.62 0.62 0.62 0.62 0.62 0.62 0.62 0.62 -0.43 0.62 0.62 0.62 0.62
16 0.62 0.62 0.62 0.62 0.62 0.62 0.62 0.62 0.62 0.62 0.62 0.62 0.62 0.62 0.62 0.62 0.62 0.62 0.62 0.60
17 -0.22 -0.22 -0.22 -0.22 -0.22 -0.22 -0.22 -0.22 -0.22 -0.22 -0.22 -0.22 -0.22 -0.22 -0.22 -0.22 -0.22 -0.22 -0.22 -0.22
18 0.16 -0.46 -0.46 -0.30 -0.30 -0.46 -0.30 0.71 1.30 -0.30 0.71 0.16 0.71 0.16 0.71 -0.46 -0.46 0.16 0.16 0.16
19 0.16 0.16 0.16 0.16 0.16 0.16 0.16 0.16 0.16 0.16 0.16 0.16 0.16 0.16 0.16 0.16 0.16 0.16 0.16 0.16
20 0.66 0.66 0.66 0.66 0.66 0.07 0.07 0.66 0.07 0.66 0.66 0.66 0.07 0.66 0.66 0.07 0.66 0.66 0.66 0.07
21 1.10 1.10 1.10 1.10 1.10 1.10 1.10 1.10 1.10 1.10 1.10 1.10 1.10 1.10 1.10 1.10 1.10 1.10 1.10 1.10
22 0.51 0.51 0.51 0.51 0.34 0.51 0.51 0.34 0.51 0.34 0.51 0.51 0.51 0.51 0.51 0.34 0.34 0.51 0.51 0.51
23 -0.30 0.71 -0.30 -0.30 0.71 -0.30 -0.30 -0.30 0.16 -0.30 0.16 0.71 -0.30 -0.30 -0.30 0.16 -0.30 -0.30 -0.30 0.16
24 0.30 -0.62 -0.62 -0.62 -0.62 -1.25 -0.62 -1.25 -1.25 -1.25 -0.62 -0.62 0.30 -1.25 -0.62 -0.62 0.30 -0.62 -0.62 0.30
25 0.62 -0.32 0.62 0.62 0.62 0.62 0.62 0.62 0.62 0.62 0.62 0.62 0.62 -0.32 0.62 0.62 -0.32 0.62 0.62 0.62
26 0.76 0.76 0.76 0.76 0.76 0.76 0.76 0.76 0.76 0.76 0.76 0.76 0.76 0.76 0.76 0.76 0.76 0.76 0.76 0.76
27 0.95 0.95 0.95 0.95 0.95 0.95 -0.17 -0.17 -0.17 0.95 -0.17 0.95 0.95 0.95 0.95 0.95 0.95 0.95 0.95 0.95
28 0.62 0.62 0.62 0.62 -0.43 0.62 0.62 -0.43 0.62 0.62 0.62 0.62 0.62 0.62 0.62 0.62 0.62 0.62 0.62 0.62
29 0.56 0.56 0.56 0.56 0.56 0.56 0.56 0.56 0.56 0.56 0.56 0.56 0.56 0.56 0.56 -0.94 0.56 0.56 0.56 0.56
30 0.27 0.27 0.27 0.27 0.27 0.27 0.27 0.27 0.27 0.27 0.27 0.27 0.27 0.27 0.27 0.27 0.27 0.27 0.27 0.27
31 0.27 0.27 0.86 0.86 0.27 0.27 0.27 0.27 0.86 0.86 0.86 0.86 0.27 0.86 0.86 0.27 0.86 0.86 0.27 0.27
32 0.56 0.56 0.56 0.56 0.66 0.66 0.56 0.66 0.66 0.56 0.56 0.66 0.56 0.56 0.66 0.56 0.56 0.56 0.56 0.66
33 0.76 0.76 0.76 0.76 0.76 0.76 0.76 0.76 0.76 0.76 0.76 0.76 0.76 0.76 0.76 0.76 0.76 0.76 0.76 0.76
34 0.76 0.76 0.76 0.76 0.76 0.76 0.76 0.76 0.76 0.76 0.76 0.76 0.76 0.76 0.76 0.76 0.76 0.76 0.76 0.76
35 -0.43 -0.58 -0.58 -0.58 -0.58 -0.58 -0.58 -0.58 -0.58 -0.58 -0.58 -0.58 -0.58 -0.58 -0.58 -0.58 -0.58 -0.58 -2.15 -0.58
36 0.09 0.09 0.62 0.09 0.09 0.09 0.09 -0.58 0.09 0.09 -0.58 0.62 0.09 0.09 0.09 0.09 -0.43 -0.58 0.09 0.09
37 0.76 0.76 0.76 0.76 0.76 0.76 0.76 0.76 0.76 0.76 0.76 0.76 0.76 0.76 0.76 0.76 0.76 0.76 0.76 0.76
38 1.11 1.11 1.11 1.11 1.11 1.11 1.11 1.11 1.11 1.11 1.11 1.11 1.11 1.11 1.11 1.11 1.11 1.11 1.11 1.11
39 0.56 0.56 0.56 0.56 -0.94 0.56 0.56 0.66 -0.94 -0.94 0.56 -0.94 0.56 0.56 0.56 -0.94 0.56 0.56 0.56 0.56
40 0.71 0.71 0.71 0.71 0.71 0.71 -0.46 0.71 0.71 0.71 0.71 0.71 0.71 0.71 0.71 0.71 0.71 0.71 0.16 0.71
41 1.30 1.30 1.30 0.71 1.30 1.30 1.30 0.71 0.71 1.30 1.30 1.30 1.30 0.71 1.30 1.30 0.71 1.30 1.30 1.30
42 0.74 0.74 0.74 0.74 0.74 0.74 0.74 0.74 0.74 0.74 0.74 0.74 0.74 0.74 0.74 0.74 0.74 0.74 0.74 0.74
43 -0.59 -0.59 -0.59 -0.59 -0.59 -0.59 -0.59 -0.59 -0.59 -0.59 -0.59 -0.59 -0.59 -0.59 -0.59 -0.59 -0.59 -0.59 -0.59 -0.59
44 0.62 0.62 0.62 0.62 0.62 0.09 0.62 0.09 0.09 0.62 0.62 0.62 0.62 0.09 0.62 0.62 0.62 0.62 0.09 0.62
45 -0.02 -0.58 -0.02 -0.02 -0.58 -0.58 -0.02 -0.02 -0.02 -0.48 -0.58 -0.02 -0.02 -0.02 -0.58 -0.02 -0.02 -0.58 -0.58 -0.58
47 1.06 1.06 -0.33 0.77 -0.33 1.06 -0.33 1.06 -0.33 -0.33 -0.33 0.77 0.77 1.06 0.77 -0.33 0.77 0.77 1.06 -0.33
48 0.07 0.07 0.07 0.07 0.66 0.66 0.66 0.66 0.07 0.66 0.07 0.07 0.07 0.07 0.66 0.07 0.66 0.66 0.66 0.07
49 0.60 0.62 0.60 0.62 0.60 0.62 0.60 0.62 0.60 0.62 0.60 0.60 0.60 0.60 0.60 0.62 0.62 0.60 0.60 0.62
50 1.11 1.11 0.55 1.11 1.11 1.11 1.11 1.11 1.11 1.11 1.11 1.11 1.11 1.11 1.11 1.11 0.55 1.11 1.11 1.11
51 0.39 0.39 0.39 -0.13 -0.13 0.39 0.39 0.39 -0.13 -0.13 0.39 0.39 0.39 0.39 0.39 0.39 0.39 0.39 0.39 0.39
52 -0.02 -0.02 -0.02 -0.02 -0.02 -0.02 -0.02 -0.02 -0.02 -0.02 -0.02 -0.02 -0.02 -0.02 -0.02 -0.02 -0.02 -0.02 -0.02 -0.02
53 -0.02 -0.02 -0.02 -0.02 -0.02 -0.02 -0.02 -0.02 -0.58 -0.02 -0.02 -0.02 -0.02 -0.02 -0.02 -0.02 -0.02 -0.02 -0.02 -0.02
54 0.30 0.41 0.30 0.30 0.41 0.30 -0.62 -0.62 -0.62 -0.29 0.30 0.30 0.41 0.41 0.30 -0.29 -0.29 0.41 -0.29 0.30
55 -0.10 -0.10 -0.10 -0.10 -0.10 -0.10 -0.10 0.47 -0.10 -0.10 0.47 -0.10 0.47 0.47 -0.10 -0.10 -0.10 -0.10 0.47 -0.10
56 0.41 0.41 0.41 0.41 0.41 0.41 0.41 0.41 0.41 0.41 0.41 0.41 0.41 0.41 0.41 0.41 0.41 0.41 0.51 0.51
63 0.62 0.62 0.62 0.62 0.62 0.60 0.62 0.62 0.62 0.60 0.62 0.62 0.62 0.60 0.62 0.62 0.60 0.62 0.62 0.62
64 -0.58 -0.48 -0.58 -0.58 -1.76 -0.58 -0.48 -1.76 -1.76 -0.58 -1.76 -0.48 -0.48 -0.58 -0.58 -0.48 -0.58 -0.58 -0.48 -1.76
65 0.71 0.71 0.71 0.71 1.30 0.71 0.71 0.71 0.71 0.71 0.71 0.71 0.71 0.71 0.71 0.71 0.71 0.71 0.71 0.71
66 -0.85 -1.35 -0.22 -0.85 -1.35 -1.35 -1.35 -1.35 -1.35 -1.35 -0.22 -1.35 -1.35 -1.35 -1.35 -1.35 -1.35 -1.35 -0.22 -1.35
67 0.66 0.66 0.66 0.66 0.66 0.66 0.66 0.56 0.66 0.66 0.66 0.66 0.66 0.66 0.66 0.66 0.66 0.66 0.66 0.66
68 0.76 0.76 0.76 0.76 0.76 0.76 0.76 0.76 0.76 0.76 0.76 0.76 0.76 0.76 0.76 0.76 0.76 0.76 0.76 0.76
69 0.47 0.47 0.47 0.47 0.47 0.47 0.47 0.47 0.47 0.47 0.47 0.47 0.47 0.47 0.47 0.47 0.47 0.47 0.47 0.47
70 0.66 0.66 0.66 0.66 0.66 0.66 0.66 0.66 0.66 0.66 0.66 0.66 0.66 0.66 0.66 0.66 0.66 0.66 0.66 0.66
73 0.24 0.24 0.24 0.24 0.24 0.24 0.24 0.24 -0.41 0.24 0.24 0.24 0.24 0.24 0.24 0.24 0.24 0.24 0.24 0.24
74 0.34 0.34 0.34 0.34 0.34 0.34 0.34 0.34 0.34 0.34 0.34 0.34 0.34 0.34 0.34 0.34 0.34 0.34 0.34 0.34
75 0.51 -0.26 0.51 0.51 0.51 0.51 0.51 0.41 -0.26 -0.26 0.51 -0.26 -0.26 0.51 -0.26 -0.26 0.51 0.41 -0.26 -0.26
76 0.51 0.51 0.51 -0.26 0.51 -0.26 -0.26 0.51 -0.26 0.51 -0.26 0.51 0.51 -0.26 -0.26 0.51 0.51 0.51 0.51 0.51
77 0.08 0.08 0.08 0.08 0.08 0.08 0.08 0.08 0.08 0.08 0.08 0.08 0.08 0.08 0.08 0.08 0.08 0.08 0.08 0.08
78 0.28 0.28 0.28 0.28 0.28 0.28 0.28 0.28 0.28 0.28 0.28 0.28 0.28 0.28 0.28 0.28 0.28 0.28 0.28 0.28
79 1.11 1.11 1.11 1.11 0.55 1.11 1.11 0.55 1.11 1.11 1.11 1.11 1.11 1.11 1.11 1.11 1.11 1.11 0.55 0.55
80 -0.41 -0.41 -0.41 -0.41 -0.41 -0.41 -0.41 -0.41 -0.25 -0.25 -0.41 -0.41 -0.41 -0.41 -0.41 -0.41 -0.41 -0.41 -0.25 -0.25
81 -0.59 -0.59 -0.59 -0.59 -0.59 -0.59 -0.59 -0.59 -0.59 -0.59 -0.59 -0.59 -0.59 -0.59 -0.59 -0.59 -0.59 -0.59 -0.59 -0.59
82 0.71 0.71 0.71 0.71 0.71 0.71 0.71 0.71 0.71 0.71 0.71 0.71 0.71 0.71 0.71 1.30 0.71 0.71 0.71 0.71
83 1.01 1.11 1.11 1.11 1.11 1.11 1.11 1.11 1.01 1.11 1.11 1.11 1.11 1.11 1.11 1.01 1.11 1.11 1.11 1.11
84 0.66 0.07 0.66 0.66 0.66 0.66 0.66 0.66 0.66 0.66 0.66 0.66 0.66 0.66 0.66 0.66 0.66 0.66 0.66 0.66
85 0.47 0.47 0.47 0.47 0.47 0.47 -0.38 0.47 0.47 0.47 0.47 0.47 0.47 -0.38 -0.38 0.47 0.47 0.47 0.47 0.47
86 0.76 0.76 0.76 0.76 0.76 0.76 0.76 0.76 0.76 0.76 0.76 0.76 0.76 0.76 0.76 0.76 0.76 0.76 0.76 0.76
87 0.86 1.01 0.86 1.01 0.86 1.01 0.86 0.86 0.86 0.86 0.86 -1.09 1.01 0.86 0.86 0.86 0.86 0.86 0.86 0.86
88 0.39 0.39 -0.13 0.39 -0.13 0.39 -0.13 0.39 0.39 -0.20 -0.13 -0.13 -0.13 -0.13 -0.13 -0.13 0.39 0.39 -0.13 0.39
89 1.30 1.30 1.30 1.30 1.30 1.30 1.30 1.30 1.30 1.30 1.30 1.30 0.71 1.30 1.30 1.30 1.30 1.30 1.30 1.30
90 0.17 0.61 0.17 -0.91 0.17 0.17 0.17 0.61 0.82 0.61 0.17 0.17 0.61 -0.91 0.17 -0.91 0.17 0.17 -0.91 0.17
91 -0.62 -0.62 -0.62 -0.62 -0.62 -0.62 -0.62 -0.62 -0.62 -0.62 -0.62 -0.62 -0.62 -0.62 -0.62 -0.62 -0.62 -0.62 -0.62 -0.62
100 0.41 0.41 0.41 0.41 0.51 0.51 0.41 0.41 0.41 0.41 0.41 0.51 0.41 0.51 0.51 0.41 0.41 0.41 0.51 0.41
101 0.62 0.62 0.62 0.62 0.60 0.62 0.62 0.62 0.62 0.62 0.62 0.62 0.62 0.60 0.62 0.62 0.62 0.62 0.60 0.62
102 0.07 0.07 0.07 0.07 0.07 0.66 0.07 0.07 0.07 0.07 0.07 0.07 0.07 0.07 0.07 0.07 0.07 0.07 0.07 0.07
103 0.62 0.62 -0.43 -0.43 0.62 -0.43 -0.43 0.62 0.62 0.62 0.62 0.09 0.62 -0.43 -0.58 0.62 0.62 0.62 -0.58 0.62
104 0.74 0.74 0.74 0.74 0.74 0.74 0.74 0.74 0.74 0.74 0.74 0.74 0.74 0.74 0.74 0.74 0.74 0.74 0.74 0.74
105 0.07 0.66 0.66 0.66 0.66 0.66 0.66 0.66 0.66 0.66 0.66 0.66 0.66 0.07 0.66 0.66 0.66 0.66 0.07 0.66
106 0.66 0.66 0.66 0.66 0.66 0.66 0.66 0.66 0.66 0.66 0.66 0.66 0.66 0.66 0.66 0.66 0.66 0.66 0.66 0.66
107 0.71 0.71 1.30 1.30 0.71 0.71 0.71 0.71 1.30 0.16 0.71 1.30 0.71 0.71 0.71 0.71 0.71 1.30 1.30 0.71
108 0.56 0.56 0.56 0.66 -0.94 0.56 0.56 0.56 0.56 0.56 0.56 0.56 0.56 0.56 0.56 0.56 0.66 0.56 0.56 0.56
109 0.62 0.62 0.62 0.62 0.62 0.62 0.62 0.62 0.60 0.62 0.62 0.62 0.60 0.62 0.62 0.62 0.62 0.62 0.60 0.62
110 -0.28 -0.28 -0.28 -0.28 -0.28 -0.28 -0.28 -0.28 -0.28 -0.28 -0.28 -0.28 -0.28 -0.28 -0.28 -0.28 -0.28 -0.28 -0.28 -0.28
111 0.74 0.74 0.74 0.74 0.74 0.74 0.74 0.74 0.74 0.74 0.74 0.74 0.74 0.74 0.74 0.74 0.74 0.74 0.74 0.74
112 0.56 0.56 0.56 0.56 0.56 0.56 0.56 0.56 0.56 1.10 1.10 0.56 0.56 0.56 0.56 0.56 0.56 0.56 0.56 0.56
113 0.66 0.66 0.66 0.66 0.66 0.66 0.66 0.66 0.66 0.66 0.66 0.66 0.66 0.66 0.66 0.66 0.66 0.66 0.66 0.66
114 1.30 1.30 1.30 1.30 1.30 1.30 1.30 1.30 1.30 1.30 1.30 1.30 1.30 1.30 1.30 1.30 1.30 1.30 1.30 1.30
115 0.74 0.74 0.41 0.41 0.74 0.41 0.74 0.74 0.41 0.74 0.74 0.41 0.41 0.74 0.74 0.74 0.74 0.74 0.74 0.41
116 -2.36 -0.59 -0.59 -0.59 -0.59 -0.59 -0.59 -0.59 -0.59 -2.36 -0.59 -0.59 -0.59 -0.59 -2.36 -2.36 -2.36 -0.59 -0.59 -0.59
117 -0.22 -0.22 -0.22 0.87 -0.22 -1.35 -1.35 -1.35 -1.35 -1.35 -0.22 -0.22 -0.22 -0.22 -0.22 -1.35 -0.85 -1.35 -0.22 -0.22
118 0.76 0.76 0.76 0.76 0.76 0.76 0.76 0.76 0.76 0.76 0.76 0.76 0.76 0.76 0.76 0.76 0.76 0.76 0.76 0.76
119 -0.29 -0.29 -0.29 -0.62 0.30 0.30 0.41 0.30 0.30 0.30 -0.62 0.30 0.30 -0.62 0.41 -0.62 -0.62 0.30 0.30 0.30
120 -0.02 -0.58 0.32 -0.58 -0.07 0.32 -0.02 -0.02 0.32 -0.02 -0.02 0.32 -0.02 -0.02 -0.02 -0.02 0.32 -0.02 -0.02 -0.58
121 0.47 -0.38 -0.38 0.47 0.47 0.47 -0.38 0.47 -0.38 0.47 0.47 -0.38 0.47 0.47 0.47 0.47 -0.38 0.47 0.47 0.47
122 -0.30 -0.46 0.16 0.16 -0.30 0.71 0.16 -0.30 0.71 -0.30 0.71 0.71 0.71 0.16 0.71 0.71 0.71 -0.30 -0.30 0.16
#Reported_Model_Average 0.371 0.376 0.368 0.370 0.312 0.357 0.307 0.322 0.314 0.312 0.363 0.381 0.402 0.327 0.349 0.273 0.347 0.396 0.328 0.373
#Overall_Average_Reported 0.347
Output from ProsaII
ProsaII Score over a window of $winsize_s residues
JPEG image for ProsaII Score
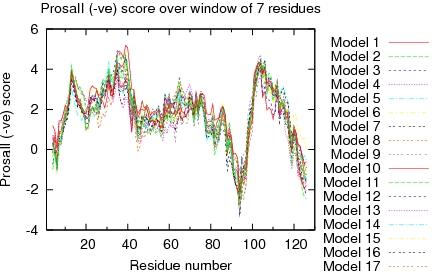
Table of Verify3D scores for ordered residues across all models
#verify3d
#Residue\Model 1 2 3 4 5 6 7 8 9 10 11 12 13 14 15 16 17 18 19 20
6 -0.38 0.47 0.47 0.47 -0.38 0.47 0.47 -0.38 0.47 0.47 -0.38 0.47 0.47 0.47 -0.38 0.47 0.47 0.47 0.47 0.47
7 0.76 0.76 0.76 0.76 0.76 0.76 0.76 0.76 0.76 0.76 0.76 0.76 0.76 0.76 0.76 0.76 0.76 0.76 0.76 0.76
8 -0.43 0.62 -0.43 0.62 -0.43 -0.43 -0.58 -0.43 -0.43 -0.43 -0.43 0.62 -0.43 -0.43 -0.43 -0.43 0.09 0.62 -0.43 0.62
9 -0.62 0.30 -0.62 -0.62 -0.62 -0.62 -0.62 -0.62 -0.62 -0.62 -0.62 -0.62 -0.62 -0.62 -0.62 -0.62 -0.62 -0.62 -0.62 0.30
10 0.50 0.50 0.50 0.50 0.50 0.50 0.50 0.50 0.50 0.50 0.50 0.50 0.50 0.50 0.50 0.50 0.50 0.50 -0.88 0.50
11 0.86 0.86 0.50 0.50 0.50 0.50 0.86 0.86 0.86 0.86 0.86 0.86 0.86 0.86 0.86 0.27 0.50 0.86 0.86 0.50
12 0.60 0.60 0.62 0.62 0.62 0.60 0.60 0.60 0.60 0.60 0.62 0.62 0.60 0.60 0.60 0.62 0.62 0.62 0.62 0.62
13 -0.43 -0.43 -0.43 -0.43 -0.43 -0.43 0.62 -0.43 0.62 -0.43 -0.43 -0.43 -0.43 -0.43 -0.43 -0.43 -0.43 -0.43 -0.43 -0.43
14 0.76 0.76 0.76 0.76 0.76 0.76 0.76 0.76 0.76 0.76 0.76 0.76 0.76 0.76 0.76 0.76 0.76 0.76 0.76 0.76
15 0.62 0.62 0.62 0.62 0.62 0.62 0.62 0.62 0.62 0.62 0.62 0.62 0.62 0.62 0.62 -0.43 0.62 0.62 0.62 0.62
16 0.62 0.62 0.62 0.62 0.62 0.62 0.62 0.62 0.62 0.62 0.62 0.62 0.62 0.62 0.62 0.62 0.62 0.62 0.62 0.60
17 -0.22 -0.22 -0.22 -0.22 -0.22 -0.22 -0.22 -0.22 -0.22 -0.22 -0.22 -0.22 -0.22 -0.22 -0.22 -0.22 -0.22 -0.22 -0.22 -0.22
18 0.16 -0.46 -0.46 -0.30 -0.30 -0.46 -0.30 0.71 1.30 -0.30 0.71 0.16 0.71 0.16 0.71 -0.46 -0.46 0.16 0.16 0.16
19 0.16 0.16 0.16 0.16 0.16 0.16 0.16 0.16 0.16 0.16 0.16 0.16 0.16 0.16 0.16 0.16 0.16 0.16 0.16 0.16
20 0.66 0.66 0.66 0.66 0.66 0.07 0.07 0.66 0.07 0.66 0.66 0.66 0.07 0.66 0.66 0.07 0.66 0.66 0.66 0.07
21 1.10 1.10 1.10 1.10 1.10 1.10 1.10 1.10 1.10 1.10 1.10 1.10 1.10 1.10 1.10 1.10 1.10 1.10 1.10 1.10
22 0.51 0.51 0.51 0.51 0.34 0.51 0.51 0.34 0.51 0.34 0.51 0.51 0.51 0.51 0.51 0.34 0.34 0.51 0.51 0.51
23 -0.30 0.71 -0.30 -0.30 0.71 -0.30 -0.30 -0.30 0.16 -0.30 0.16 0.71 -0.30 -0.30 -0.30 0.16 -0.30 -0.30 -0.30 0.16
24 0.30 -0.62 -0.62 -0.62 -0.62 -1.25 -0.62 -1.25 -1.25 -1.25 -0.62 -0.62 0.30 -1.25 -0.62 -0.62 0.30 -0.62 -0.62 0.30
25 0.62 -0.32 0.62 0.62 0.62 0.62 0.62 0.62 0.62 0.62 0.62 0.62 0.62 -0.32 0.62 0.62 -0.32 0.62 0.62 0.62
26 0.76 0.76 0.76 0.76 0.76 0.76 0.76 0.76 0.76 0.76 0.76 0.76 0.76 0.76 0.76 0.76 0.76 0.76 0.76 0.76
27 0.95 0.95 0.95 0.95 0.95 0.95 -0.17 -0.17 -0.17 0.95 -0.17 0.95 0.95 0.95 0.95 0.95 0.95 0.95 0.95 0.95
28 0.62 0.62 0.62 0.62 -0.43 0.62 0.62 -0.43 0.62 0.62 0.62 0.62 0.62 0.62 0.62 0.62 0.62 0.62 0.62 0.62
29 0.56 0.56 0.56 0.56 0.56 0.56 0.56 0.56 0.56 0.56 0.56 0.56 0.56 0.56 0.56 -0.94 0.56 0.56 0.56 0.56
30 0.27 0.27 0.27 0.27 0.27 0.27 0.27 0.27 0.27 0.27 0.27 0.27 0.27 0.27 0.27 0.27 0.27 0.27 0.27 0.27
31 0.27 0.27 0.86 0.86 0.27 0.27 0.27 0.27 0.86 0.86 0.86 0.86 0.27 0.86 0.86 0.27 0.86 0.86 0.27 0.27
32 0.56 0.56 0.56 0.56 0.66 0.66 0.56 0.66 0.66 0.56 0.56 0.66 0.56 0.56 0.66 0.56 0.56 0.56 0.56 0.66
33 0.76 0.76 0.76 0.76 0.76 0.76 0.76 0.76 0.76 0.76 0.76 0.76 0.76 0.76 0.76 0.76 0.76 0.76 0.76 0.76
34 0.76 0.76 0.76 0.76 0.76 0.76 0.76 0.76 0.76 0.76 0.76 0.76 0.76 0.76 0.76 0.76 0.76 0.76 0.76 0.76
35 -0.43 -0.58 -0.58 -0.58 -0.58 -0.58 -0.58 -0.58 -0.58 -0.58 -0.58 -0.58 -0.58 -0.58 -0.58 -0.58 -0.58 -0.58 -2.15 -0.58
36 0.09 0.09 0.62 0.09 0.09 0.09 0.09 -0.58 0.09 0.09 -0.58 0.62 0.09 0.09 0.09 0.09 -0.43 -0.58 0.09 0.09
37 0.76 0.76 0.76 0.76 0.76 0.76 0.76 0.76 0.76 0.76 0.76 0.76 0.76 0.76 0.76 0.76 0.76 0.76 0.76 0.76
38 1.11 1.11 1.11 1.11 1.11 1.11 1.11 1.11 1.11 1.11 1.11 1.11 1.11 1.11 1.11 1.11 1.11 1.11 1.11 1.11
39 0.56 0.56 0.56 0.56 -0.94 0.56 0.56 0.66 -0.94 -0.94 0.56 -0.94 0.56 0.56 0.56 -0.94 0.56 0.56 0.56 0.56
40 0.71 0.71 0.71 0.71 0.71 0.71 -0.46 0.71 0.71 0.71 0.71 0.71 0.71 0.71 0.71 0.71 0.71 0.71 0.16 0.71
41 1.30 1.30 1.30 0.71 1.30 1.30 1.30 0.71 0.71 1.30 1.30 1.30 1.30 0.71 1.30 1.30 0.71 1.30 1.30 1.30
42 0.74 0.74 0.74 0.74 0.74 0.74 0.74 0.74 0.74 0.74 0.74 0.74 0.74 0.74 0.74 0.74 0.74 0.74 0.74 0.74
43 -0.59 -0.59 -0.59 -0.59 -0.59 -0.59 -0.59 -0.59 -0.59 -0.59 -0.59 -0.59 -0.59 -0.59 -0.59 -0.59 -0.59 -0.59 -0.59 -0.59
44 0.62 0.62 0.62 0.62 0.62 0.09 0.62 0.09 0.09 0.62 0.62 0.62 0.62 0.09 0.62 0.62 0.62 0.62 0.09 0.62
45 -0.02 -0.58 -0.02 -0.02 -0.58 -0.58 -0.02 -0.02 -0.02 -0.48 -0.58 -0.02 -0.02 -0.02 -0.58 -0.02 -0.02 -0.58 -0.58 -0.58
47 1.06 1.06 -0.33 0.77 -0.33 1.06 -0.33 1.06 -0.33 -0.33 -0.33 0.77 0.77 1.06 0.77 -0.33 0.77 0.77 1.06 -0.33
48 0.07 0.07 0.07 0.07 0.66 0.66 0.66 0.66 0.07 0.66 0.07 0.07 0.07 0.07 0.66 0.07 0.66 0.66 0.66 0.07
49 0.60 0.62 0.60 0.62 0.60 0.62 0.60 0.62 0.60 0.62 0.60 0.60 0.60 0.60 0.60 0.62 0.62 0.60 0.60 0.62
50 1.11 1.11 0.55 1.11 1.11 1.11 1.11 1.11 1.11 1.11 1.11 1.11 1.11 1.11 1.11 1.11 0.55 1.11 1.11 1.11
51 0.39 0.39 0.39 -0.13 -0.13 0.39 0.39 0.39 -0.13 -0.13 0.39 0.39 0.39 0.39 0.39 0.39 0.39 0.39 0.39 0.39
52 -0.02 -0.02 -0.02 -0.02 -0.02 -0.02 -0.02 -0.02 -0.02 -0.02 -0.02 -0.02 -0.02 -0.02 -0.02 -0.02 -0.02 -0.02 -0.02 -0.02
53 -0.02 -0.02 -0.02 -0.02 -0.02 -0.02 -0.02 -0.02 -0.58 -0.02 -0.02 -0.02 -0.02 -0.02 -0.02 -0.02 -0.02 -0.02 -0.02 -0.02
54 0.30 0.41 0.30 0.30 0.41 0.30 -0.62 -0.62 -0.62 -0.29 0.30 0.30 0.41 0.41 0.30 -0.29 -0.29 0.41 -0.29 0.30
55 -0.10 -0.10 -0.10 -0.10 -0.10 -0.10 -0.10 0.47 -0.10 -0.10 0.47 -0.10 0.47 0.47 -0.10 -0.10 -0.10 -0.10 0.47 -0.10
56 0.41 0.41 0.41 0.41 0.41 0.41 0.41 0.41 0.41 0.41 0.41 0.41 0.41 0.41 0.41 0.41 0.41 0.41 0.51 0.51
63 0.62 0.62 0.62 0.62 0.62 0.60 0.62 0.62 0.62 0.60 0.62 0.62 0.62 0.60 0.62 0.62 0.60 0.62 0.62 0.62
64 -0.58 -0.48 -0.58 -0.58 -1.76 -0.58 -0.48 -1.76 -1.76 -0.58 -1.76 -0.48 -0.48 -0.58 -0.58 -0.48 -0.58 -0.58 -0.48 -1.76
65 0.71 0.71 0.71 0.71 1.30 0.71 0.71 0.71 0.71 0.71 0.71 0.71 0.71 0.71 0.71 0.71 0.71 0.71 0.71 0.71
66 -0.85 -1.35 -0.22 -0.85 -1.35 -1.35 -1.35 -1.35 -1.35 -1.35 -0.22 -1.35 -1.35 -1.35 -1.35 -1.35 -1.35 -1.35 -0.22 -1.35
67 0.66 0.66 0.66 0.66 0.66 0.66 0.66 0.56 0.66 0.66 0.66 0.66 0.66 0.66 0.66 0.66 0.66 0.66 0.66 0.66
68 0.76 0.76 0.76 0.76 0.76 0.76 0.76 0.76 0.76 0.76 0.76 0.76 0.76 0.76 0.76 0.76 0.76 0.76 0.76 0.76
69 0.47 0.47 0.47 0.47 0.47 0.47 0.47 0.47 0.47 0.47 0.47 0.47 0.47 0.47 0.47 0.47 0.47 0.47 0.47 0.47
70 0.66 0.66 0.66 0.66 0.66 0.66 0.66 0.66 0.66 0.66 0.66 0.66 0.66 0.66 0.66 0.66 0.66 0.66 0.66 0.66
73 0.24 0.24 0.24 0.24 0.24 0.24 0.24 0.24 -0.41 0.24 0.24 0.24 0.24 0.24 0.24 0.24 0.24 0.24 0.24 0.24
74 0.34 0.34 0.34 0.34 0.34 0.34 0.34 0.34 0.34 0.34 0.34 0.34 0.34 0.34 0.34 0.34 0.34 0.34 0.34 0.34
75 0.51 -0.26 0.51 0.51 0.51 0.51 0.51 0.41 -0.26 -0.26 0.51 -0.26 -0.26 0.51 -0.26 -0.26 0.51 0.41 -0.26 -0.26
76 0.51 0.51 0.51 -0.26 0.51 -0.26 -0.26 0.51 -0.26 0.51 -0.26 0.51 0.51 -0.26 -0.26 0.51 0.51 0.51 0.51 0.51
77 0.08 0.08 0.08 0.08 0.08 0.08 0.08 0.08 0.08 0.08 0.08 0.08 0.08 0.08 0.08 0.08 0.08 0.08 0.08 0.08
78 0.28 0.28 0.28 0.28 0.28 0.28 0.28 0.28 0.28 0.28 0.28 0.28 0.28 0.28 0.28 0.28 0.28 0.28 0.28 0.28
79 1.11 1.11 1.11 1.11 0.55 1.11 1.11 0.55 1.11 1.11 1.11 1.11 1.11 1.11 1.11 1.11 1.11 1.11 0.55 0.55
80 -0.41 -0.41 -0.41 -0.41 -0.41 -0.41 -0.41 -0.41 -0.25 -0.25 -0.41 -0.41 -0.41 -0.41 -0.41 -0.41 -0.41 -0.41 -0.25 -0.25
81 -0.59 -0.59 -0.59 -0.59 -0.59 -0.59 -0.59 -0.59 -0.59 -0.59 -0.59 -0.59 -0.59 -0.59 -0.59 -0.59 -0.59 -0.59 -0.59 -0.59
82 0.71 0.71 0.71 0.71 0.71 0.71 0.71 0.71 0.71 0.71 0.71 0.71 0.71 0.71 0.71 1.30 0.71 0.71 0.71 0.71
83 1.01 1.11 1.11 1.11 1.11 1.11 1.11 1.11 1.01 1.11 1.11 1.11 1.11 1.11 1.11 1.01 1.11 1.11 1.11 1.11
84 0.66 0.07 0.66 0.66 0.66 0.66 0.66 0.66 0.66 0.66 0.66 0.66 0.66 0.66 0.66 0.66 0.66 0.66 0.66 0.66
85 0.47 0.47 0.47 0.47 0.47 0.47 -0.38 0.47 0.47 0.47 0.47 0.47 0.47 -0.38 -0.38 0.47 0.47 0.47 0.47 0.47
86 0.76 0.76 0.76 0.76 0.76 0.76 0.76 0.76 0.76 0.76 0.76 0.76 0.76 0.76 0.76 0.76 0.76 0.76 0.76 0.76
87 0.86 1.01 0.86 1.01 0.86 1.01 0.86 0.86 0.86 0.86 0.86 -1.09 1.01 0.86 0.86 0.86 0.86 0.86 0.86 0.86
88 0.39 0.39 -0.13 0.39 -0.13 0.39 -0.13 0.39 0.39 -0.20 -0.13 -0.13 -0.13 -0.13 -0.13 -0.13 0.39 0.39 -0.13 0.39
89 1.30 1.30 1.30 1.30 1.30 1.30 1.30 1.30 1.30 1.30 1.30 1.30 0.71 1.30 1.30 1.30 1.30 1.30 1.30 1.30
90 0.17 0.61 0.17 -0.91 0.17 0.17 0.17 0.61 0.82 0.61 0.17 0.17 0.61 -0.91 0.17 -0.91 0.17 0.17 -0.91 0.17
91 -0.62 -0.62 -0.62 -0.62 -0.62 -0.62 -0.62 -0.62 -0.62 -0.62 -0.62 -0.62 -0.62 -0.62 -0.62 -0.62 -0.62 -0.62 -0.62 -0.62
100 0.41 0.41 0.41 0.41 0.51 0.51 0.41 0.41 0.41 0.41 0.41 0.51 0.41 0.51 0.51 0.41 0.41 0.41 0.51 0.41
101 0.62 0.62 0.62 0.62 0.60 0.62 0.62 0.62 0.62 0.62 0.62 0.62 0.62 0.60 0.62 0.62 0.62 0.62 0.60 0.62
102 0.07 0.07 0.07 0.07 0.07 0.66 0.07 0.07 0.07 0.07 0.07 0.07 0.07 0.07 0.07 0.07 0.07 0.07 0.07 0.07
103 0.62 0.62 -0.43 -0.43 0.62 -0.43 -0.43 0.62 0.62 0.62 0.62 0.09 0.62 -0.43 -0.58 0.62 0.62 0.62 -0.58 0.62
104 0.74 0.74 0.74 0.74 0.74 0.74 0.74 0.74 0.74 0.74 0.74 0.74 0.74 0.74 0.74 0.74 0.74 0.74 0.74 0.74
105 0.07 0.66 0.66 0.66 0.66 0.66 0.66 0.66 0.66 0.66 0.66 0.66 0.66 0.07 0.66 0.66 0.66 0.66 0.07 0.66
106 0.66 0.66 0.66 0.66 0.66 0.66 0.66 0.66 0.66 0.66 0.66 0.66 0.66 0.66 0.66 0.66 0.66 0.66 0.66 0.66
107 0.71 0.71 1.30 1.30 0.71 0.71 0.71 0.71 1.30 0.16 0.71 1.30 0.71 0.71 0.71 0.71 0.71 1.30 1.30 0.71
108 0.56 0.56 0.56 0.66 -0.94 0.56 0.56 0.56 0.56 0.56 0.56 0.56 0.56 0.56 0.56 0.56 0.66 0.56 0.56 0.56
109 0.62 0.62 0.62 0.62 0.62 0.62 0.62 0.62 0.60 0.62 0.62 0.62 0.60 0.62 0.62 0.62 0.62 0.62 0.60 0.62
110 -0.28 -0.28 -0.28 -0.28 -0.28 -0.28 -0.28 -0.28 -0.28 -0.28 -0.28 -0.28 -0.28 -0.28 -0.28 -0.28 -0.28 -0.28 -0.28 -0.28
111 0.74 0.74 0.74 0.74 0.74 0.74 0.74 0.74 0.74 0.74 0.74 0.74 0.74 0.74 0.74 0.74 0.74 0.74 0.74 0.74
112 0.56 0.56 0.56 0.56 0.56 0.56 0.56 0.56 0.56 1.10 1.10 0.56 0.56 0.56 0.56 0.56 0.56 0.56 0.56 0.56
113 0.66 0.66 0.66 0.66 0.66 0.66 0.66 0.66 0.66 0.66 0.66 0.66 0.66 0.66 0.66 0.66 0.66 0.66 0.66 0.66
114 1.30 1.30 1.30 1.30 1.30 1.30 1.30 1.30 1.30 1.30 1.30 1.30 1.30 1.30 1.30 1.30 1.30 1.30 1.30 1.30
115 0.74 0.74 0.41 0.41 0.74 0.41 0.74 0.74 0.41 0.74 0.74 0.41 0.41 0.74 0.74 0.74 0.74 0.74 0.74 0.41
116 -2.36 -0.59 -0.59 -0.59 -0.59 -0.59 -0.59 -0.59 -0.59 -2.36 -0.59 -0.59 -0.59 -0.59 -2.36 -2.36 -2.36 -0.59 -0.59 -0.59
117 -0.22 -0.22 -0.22 0.87 -0.22 -1.35 -1.35 -1.35 -1.35 -1.35 -0.22 -0.22 -0.22 -0.22 -0.22 -1.35 -0.85 -1.35 -0.22 -0.22
118 0.76 0.76 0.76 0.76 0.76 0.76 0.76 0.76 0.76 0.76 0.76 0.76 0.76 0.76 0.76 0.76 0.76 0.76 0.76 0.76
119 -0.29 -0.29 -0.29 -0.62 0.30 0.30 0.41 0.30 0.30 0.30 -0.62 0.30 0.30 -0.62 0.41 -0.62 -0.62 0.30 0.30 0.30
120 -0.02 -0.58 0.32 -0.58 -0.07 0.32 -0.02 -0.02 0.32 -0.02 -0.02 0.32 -0.02 -0.02 -0.02 -0.02 0.32 -0.02 -0.02 -0.58
121 0.47 -0.38 -0.38 0.47 0.47 0.47 -0.38 0.47 -0.38 0.47 0.47 -0.38 0.47 0.47 0.47 0.47 -0.38 0.47 0.47 0.47
122 -0.30 -0.46 0.16 0.16 -0.30 0.71 0.16 -0.30 0.71 -0.30 0.71 0.71 0.71 0.16 0.71 0.71 0.71 -0.30 -0.30 0.16
#Reported_Model_Average 0.371 0.376 0.368 0.370 0.312 0.357 0.307 0.322 0.314 0.312 0.363 0.381 0.402 0.327 0.349 0.273 0.347 0.396 0.328 0.373
#Overall_Average_Reported 0.347
Output from MolProbity
VdW violations from MAGE
JPEG image for MAGE VdW violation
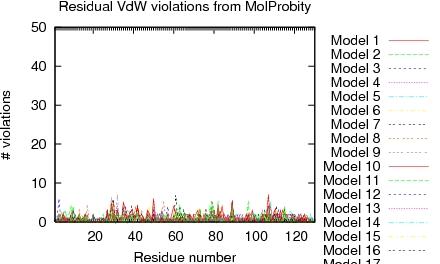
Table of MAGE VdW violations for ordered residues across all models
#mage_clash
#Residue\Model 1 2 3 4 5 6 7 8 9 10 11 12 13 14 15 16 17 18 19 20
6.000 0 1 0 0 0 0 1 1 1 1 0 0 1 0 0 0 0 0 0 3
7.000 0 2 0 0 0 0 1 0 0 0 0 1 0 0 0 0 0 1 0 1
8.000 0 2 0 0 0 0 0 1 0 1 0 0 0 0 0 0 0 0 1 1
9.000 2 4 1 0 1 1 1 1 1 1 0 0 1 1 1 2 1 0 1 0
10.000 3 0 2 0 1 1 2 1 0 1 0 3 0 0 2 1 0 0 0 4
11.000 1 1 0 0 1 1 2 0 1 0 1 0 0 1 0 0 1 0 0 1
12.000 0 0 1 0 0 0 0 0 1 0 1 1 0 0 0 1 0 0 1 0
13.000 2 2 0 0 0 1 1 0 2 0 0 1 0 0 0 1 3 0 1 0
14.000 0 0 0 0 0 0 0 1 2 0 0 0 0 0 0 0 0 1 0 1
15.000 1 1 0 0 0 1 1 0 1 0 0 0 0 1 0 1 1 0 0 1
16.000 0 2 1 0 0 0 0 0 1 0 1 2 0 0 0 0 0 0 2 0
17.000 0 1 1 0 1 1 0 4 0 0 0 2 0 0 0 0 2 0 0 1
18.000 0 0 0 0 0 0 0 1 0 0 0 0 0 0 0 0 0 1 0 0
19.000 0 0 0 0 0 0 0 0 0 0 0 0 0 0 0 0 0 0 0 0
20.000 0 2 0 0 0 0 0 2 0 0 0 2 0 0 0 0 0 2 0 0
21.000 0 0 0 0 0 0 0 0 0 0 0 0 0 0 0 0 0 0 0 0
22.000 0 0 0 0 0 0 0 1 0 0 0 0 0 0 0 0 0 0 0 0
23.000 0 0 0 1 0 0 0 0 0 0 0 1 0 0 0 0 1 1 0 0
24.000 1 0 0 1 0 0 0 0 0 0 1 1 0 0 1 0 1 0 0 0
25.000 0 0 0 0 1 0 0 1 1 0 2 0 1 0 0 1 0 0 0 0
26.000 0 0 0 0 0 0 0 1 0 0 0 1 0 0 0 0 0 0 0 0
27.000 1 0 2 1 2 3 2 1 0 1 0 2 3 1 3 3 0 1 3 1
28.000 0 0 0 0 2 0 1 0 1 0 0 0 0 0 0 0 0 0 0 0
29.000 4 3 2 0 3 2 5 4 6 0 1 2 1 0 0 4 5 0 0 2
30.000 1 0 2 2 3 3 2 3 1 5 1 1 2 3 4 6 1 3 2 2
31.000 0 0 0 1 0 0 3 0 1 0 0 0 0 0 0 0 0 0 0 0
32.000 0 3 1 4 2 2 3 3 3 3 2 2 2 0 2 2 1 7 0 1
33.000 0 0 1 0 0 0 1 0 0 0 0 1 0 0 0 1 0 0 0 2
34.000 0 1 0 0 0 1 1 0 0 0 0 1 1 0 1 1 0 1 1 1
35.000 1 0 3 4 1 0 3 3 1 5 2 1 2 0 2 4 0 0 1 1
36.000 1 3 0 1 2 2 2 2 1 3 0 1 3 0 3 4 0 3 2 1
37.000 1 1 0 2 1 1 1 2 0 1 0 0 0 0 1 0 1 3 0 0
38.000 1 1 1 1 1 2 2 1 1 2 2 2 2 1 2 2 1 2 2 2
39.000 1 2 1 2 2 3 2 2 1 1 0 3 1 1 2 4 0 1 3 2
40.000 0 1 0 0 2 2 1 0 2 1 1 1 1 2 0 2 0 2 2 0
41.000 1 1 0 1 1 0 1 4 0 1 1 1 1 1 1 1 4 1 1 0
42.000 2 2 3 3 1 4 3 4 1 4 2 2 2 3 3 4 4 2 4 2
43.000 0 1 1 1 1 0 1 1 1 0 0 1 0 1 1 1 0 0 1 1
44.000 1 0 0 0 0 0 1 0 0 1 1 2 1 0 0 2 3 0 0 0
45.000 0 0 0 1 0 1 1 0 0 0 0 1 0 0 0 1 1 0 1 0
47.000 1 1 2 3 0 3 2 3 0 1 2 2 1 1 2 1 2 1 3 1
48.000 0 0 1 0 1 0 0 1 1 0 1 1 2 1 4 0 2 2 2 0
49.000 0 0 0 0 0 0 0 0 0 0 0 1 0 0 0 0 1 1 0 0
50.000 1 1 1 3 0 5 2 2 1 2 0 3 1 1 0 2 1 4 6 0
51.000 1 0 1 0 2 0 0 0 1 0 1 1 1 2 0 0 0 2 0 0
52.000 0 0 0 0 0 0 0 2 0 0 2 1 2 0 0 0 2 1 0 0
53.000 0 0 0 0 0 0 2 0 1 0 0 1 0 0 0 0 0 0 0 0
54.000 1 1 3 1 0 1 0 1 0 2 0 1 3 2 0 1 3 3 1 0
55.000 0 0 0 0 1 0 0 1 0 0 2 0 1 1 0 0 5 1 0 0
56.000 0 0 0 0 0 0 0 0 0 0 0 0 0 0 0 0 0 0 0 2
63.000 1 3 0 0 2 0 4 2 0 0 0 0 0 0 2 2 0 0 1 4
64.000 0 0 2 1 1 1 1 1 0 1 0 0 0 1 0 0 0 1 0 0
65.000 1 1 1 0 1 0 1 0 1 2 2 1 0 1 0 3 1 1 0 0
66.000 0 2 0 0 0 1 2 0 0 0 1 0 0 0 0 0 0 0 2 2
67.000 0 1 2 0 0 0 1 0 0 0 0 0 0 1 0 1 2 1 2 1
68.000 1 1 1 1 1 2 1 0 0 2 0 1 1 0 0 2 2 1 0 0
69.000 0 0 0 0 0 0 0 0 1 0 1 0 1 0 0 1 0 0 0 0
70.000 1 1 0 0 0 1 0 2 2 0 1 0 0 1 0 0 0 0 2 0
73.000 3 0 2 3 3 2 2 3 0 2 1 0 1 1 3 2 2 3 0 3
74.000 0 0 0 0 0 0 0 0 0 0 0 0 0 0 0 0 0 0 0 0
75.000 0 0 0 0 0 0 0 0 0 0 0 0 1 0 0 0 0 0 2 0
76.000 1 2 1 1 2 1 1 2 0 1 1 2 1 0 1 3 3 0 0 2
77.000 1 0 0 1 0 1 1 1 0 1 0 1 1 2 1 1 1 1 0 1
78.000 1 1 2 2 1 1 1 2 1 1 4 1 1 1 1 1 2 1 1 1
79.000 2 3 1 3 2 1 3 3 1 2 1 3 3 2 3 5 3 4 2 5
80.000 2 2 1 3 1 1 3 2 2 1 3 2 3 2 2 3 2 3 1 1
81.000 0 0 0 1 0 1 0 0 1 0 0 0 0 0 0 0 1 0 0 0
82.000 3 3 2 2 2 2 1 2 1 1 5 2 1 1 1 1 1 1 4 1
83.000 0 0 2 0 1 0 3 0 0 0 1 0 1 1 0 0 0 0 1 0
84.000 0 0 0 2 0 2 0 2 1 0 0 0 0 0 0 0 1 1 4 0
85.000 0 1 0 1 1 0 0 1 0 2 0 1 0 1 1 0 1 0 1 0
86.000 0 0 0 1 0 0 0 0 1 1 0 1 0 0 0 0 0 0 0 0
87.000 0 1 2 0 1 0 1 0 0 1 1 1 2 0 1 0 2 2 1 0
88.000 0 0 1 0 0 0 0 0 0 0 0 0 0 0 0 0 0 0 0 0
89.000 2 2 1 1 1 0 1 0 1 5 1 5 0 3 1 2 3 2 5 4
90.000 0 0 0 1 0 0 2 0 0 1 0 1 0 0 0 0 0 0 0 0
91.000 0 0 1 0 2 0 0 0 0 1 1 1 1 0 1 0 2 1 1 0
100.000 1 1 0 0 2 1 1 0 1 1 0 0 1 2 2 1 0 2 1 2
101.000 0 1 1 0 0 0 0 0 0 0 1 1 0 0 1 2 0 1 0 2
102.000 0 0 0 0 2 2 1 0 0 1 0 0 0 0 1 2 0 0 0 0
103.000 1 0 0 0 0 1 1 0 0 0 0 0 0 2 1 0 0 1 0 1
104.000 1 1 0 1 1 1 1 0 2 1 0 0 2 0 3 1 1 3 2 1
105.000 0 1 1 0 2 1 3 0 0 2 1 1 0 0 2 2 2 1 0 2
106.000 1 0 0 0 3 0 1 0 0 1 0 0 0 1 0 1 0 0 1 0
107.000 4 3 1 4 2 0 1 4 2 4 2 3 2 3 3 2 1 1 7 2
108.000 1 0 0 1 5 2 2 0 0 4 0 0 0 5 1 0 2 1 0 0
109.000 1 1 1 1 0 2 1 0 0 1 2 1 0 0 0 0 0 0 1 0
110.000 2 2 0 2 1 0 0 1 0 3 1 2 2 1 1 2 0 0 0 0
111.000 0 2 1 2 2 2 2 4 0 1 2 1 1 2 1 4 2 1 2 1
112.000 1 1 3 1 0 1 0 0 0 1 2 2 1 0 0 0 1 1 1 1
113.000 1 1 0 1 2 2 0 0 0 3 2 1 2 0 1 2 1 0 0 0
114.000 0 0 1 1 0 0 0 0 0 0 2 1 0 0 0 0 0 1 3 0
115.000 1 4 1 2 2 1 3 1 1 2 1 2 2 2 2 1 1 2 2 3
116.000 1 1 0 1 1 1 0 1 0 2 1 1 2 0 1 0 2 2 2 1
117.000 0 0 0 0 0 0 0 2 0 0 0 0 1 1 0 0 1 0 2 0
118.000 0 0 0 2 0 0 0 2 1 0 1 0 1 3 1 0 0 1 2 0
119.000 1 0 0 2 2 0 1 0 1 2 1 2 2 1 1 0 0 1 1 1
120.000 1 0 0 0 1 1 0 1 0 2 2 0 1 0 1 0 1 1 1 0
121.000 1 0 0 1 0 0 0 0 0 1 1 0 0 1 1 0 0 0 0 0
122.000 0 0 0 1 1 0 1 1 1 1 1 2 2 0 0 1 0 2 0 0
#Reported_Model_Average 0.670 0.830 0.630 0.800 0.840 0.760 0.990 0.950 0.570 0.950 0.740 0.930 0.760 0.660 0.770 1.010 0.910 0.920 0.990 0.760
#Overall_Average_Reported 0.822
List of bad contacts calculated by MAGE for model $num_n
/farm/software/bin/probe
: 2123:A 107 LEU HG :A 89 LEU 1HD1 : -0.763: 0
: 2123:A 107 LEU HG :A 89 LEU CD1 : -0.496: 0
: 2123:A 107 LEU HA :A 107 LEU 3HD1 : -0.410: 0
: 2123:A 82 LEU HG :A 78 GLU O : -0.740: 0
: 2123:A 82 LEU 2HD2 :A 110 ASP CG : -0.474: 0
: 2123:A 82 LEU 2HD2 :A 110 ASP 2HB : -0.403: 0
: 2123:A 29 LYS 2HG :A 10 TYR HD2 : -0.649: 0
: 2123:A 9 VAL 3HG2 :A 5 THR O : -0.475: 0
: 2123:A 9 VAL O :A 13 GLU 2HG : -0.463: 0
: 2123:A 29 LYS 1HG :A 13 GLU 1HB : -0.455: 0
: 2123:A 10 TYR O :A 29 LYS 2HB : -0.452: 0
: 2123:A 29 LYS 2HG :A 10 TYR CD2 : -0.438: 0
: 2123:A 44 GLU 2HG :A 121 SER 1HB : -0.621: 0
: 2123:A 47 LEU 1HB :A 42 VAL 3HG1 : -0.609: 0
: 2123:A 42 VAL 3HG2 :A 38 ILE O : -0.586: 0
: 2123:A 73 ARG O :A 73 ARG 2HD : -0.583: 0
: 2123:A 77 THR HA :A 73 ARG 1HG : -0.481: 0
: 2123:A 106 LYS 1HE :A 103 GLU HA : -0.582: 0
: 2123:A 108 LYS O :A 112 ARG 1HB : -0.574: 0
: 2123:A 50 ILE 1HG2 :A 68 ALA 2HB : -0.573: 0
: 2123:A 36 GLU HA :A 39 LYS 2HD : -0.570: 0
: 2123:A 35 GLU 2HG :A 65 LEU 3HD1 : -0.557: 0
: 2123:A 3 ILE CG2 :A 125 HIS HA : -0.556: 0
: 2123:A 125 HIS HA :A 3 ILE 1HG2 : -0.451: 0
: 2123:A 15 GLU 1HG :A 11 TYR O : -0.541: 0
: 2123:A 54 VAL 2HG1 :A 51 THR HA : -0.529: 0
: 2123:A 61 LYS 2HG :A 63 GLU OE1 : -0.521: 0
: 2123:A 76 ASN 2HB :A 79 ILE 2HG1 : -0.512: 0
: 2123:A 80 PRO 1HD :A 79 ILE N : -0.444: 0
: 2123:A 80 PRO 2HB :A 70 LYS HA : -0.426: 0
: 2123:A 109 GLU O :A 113 LYS 2HG : -0.502: 0
: 2123:A 95 HIS 2HB :A 94 PHE O : -0.464: 0
: 2123:A 41 LEU HG :A 37 ALA O : -0.458: 0
: 2123:A 104 VAL 3HG2 :A 100 ASN O : -0.447: 0
: 2123:A 115 VAL O :A 119 VAL 3HG2 : -0.444: 0
: 2123:A 27 CYS HA :A 30 TYR HD2 : -0.429: 0
: 2123:A 97 LEU 2HB :A 24 VAL CG1 : -0.418: 0
: 2123:A 116 ILE O :A 120 ASN 1HB : -0.402: 0
#sum2 ::17.90 clashscore : 17.90 clashscore B<40
#summary::2123 atoms:2123 atoms B<40:236977 potential dots:14810.0 A^2:38 bumps:38 bumps B<40:586 score
List of bad contacts calculated by MAGE for model $num_n
/farm/software/bin/probe
: 2123:A 76 ASN 2HB :A 79 ILE 2HG1 : -0.792: 0
: 2123:A 76 ASN 2HB :A 79 ILE CG1 : -0.493: 0
: 2123:A 80 PRO 1HD :A 79 ILE N : -0.490: 0
: 2123:A 80 PRO 2HB :A 70 LYS HA : -0.401: 0
: 2123:A 97 LEU 1HB :A 93 GLY 1HA : -0.790: 0
: 2123:A 92 GLU O :A 97 LEU HG : -0.518: 0
: 2123:A 97 LEU 2HD2 :A 96 GLU C : -0.485: 0
: 2123:A 97 LEU N :A 97 LEU 2HD2 : -0.406: 0
: 2123:A 107 LEU HG :A 89 LEU 1HD1 : -0.742: 0
: 2123:A 107 LEU HG :A 89 LEU CD1 : -0.526: 0
: 2123:A 107 LEU 1HD2 :A 85 SER 1HB : -0.460: 0
: 2123:A 39 LYS 1HE :A 36 GLU HA : -0.735: 0
: 2123:A 43 ILE 1HG1 :A 39 LYS O : -0.612: 0
: 2123:A 36 GLU O :A 40 LEU 3HD1 : -0.514: 0
: 2123:A 32 LYS O :A 36 GLU 1HG : -0.505: 0
: 2123:A 32 LYS HA :A 32 LYS 2HD : -0.431: 0
: 2123:A 82 LEU HG :A 78 GLU O : -0.698: 0
: 2123:A 110 ASP CA :A 82 LEU 2HD2 : -0.434: 0
: 2123:A 82 LEU 2HD2 :A 110 ASP HA : -0.426: 0
: 2123:A 63 GLU HA :A 66 PHE CE2 : -0.628: 0
: 2123:A 63 GLU O :A 67 LYS 1HB : -0.503: 0
: 2123:A 63 GLU HA :A 66 PHE CZ : -0.412: 0
: 2123:A 126 HIS CE1 :A 123 GLU 2HB : -0.590: 0
: 2123:A 17 PHE HE2 :A 29 LYS 2HE : -0.585: 0
: 2123:A 29 LYS 1HG :A 13 GLU 1HB : -0.582: 0
: 2123:A 29 LYS 1HG :A 13 GLU CB : -0.492: 0
: 2123:A 112 ARG O :A 116 ILE 2HG1 : -0.572: 0
: 2123:A 42 VAL 3HG2 :A 38 ILE O : -0.560: 0
: 2123:A 47 LEU 1HB :A 42 VAL 3HG1 : -0.536: 0
: 2123:A 7 ALA 1HB :A 115 VAL 3HG1 : -0.517: 0
: 2123:A 7 ALA CB :A 115 VAL 3HG1 : -0.453: 0
: 2123:A 8 GLU 1HG :A 115 VAL 1HG1 : -0.425: 0
: 2123:A 4 SER 2HB :A 8 GLU 1HB : -0.420: 0
: 2123:A 111 VAL 3HG1 :A 34 ALA 2HB : -0.413: 0
: 2123:A 115 VAL 3HG2 :A 111 VAL O : -0.410: 0
: 2123:A 127 HIS C :A 129 HIS H : -0.504: 0
: 2123:A 105 LYS 2HG :A 101 GLU O : -0.498: 0
: 2123:A 104 VAL 3HG2 :A 100 ASN O : -0.491: 0
: 2123:A 72 LEU HG :A 68 ALA O : -0.487: 0
: 2123:A 65 LEU 2HD1 :A 62 SER HA : -0.481: 0
: 2123:A 87 TRP CZ2 :A 62 SER 1HB : -0.469: 0
: 2123:A 3 ILE 2HD1 :A 9 VAL 1HG2 : -0.472: 0
: 2123:A 9 VAL 3HG2 :A 5 THR O : -0.471: 0
: 2123:A 3 ILE 2HD1 :A 9 VAL CG2 : -0.448: 0
: 2123:A 9 VAL HB :A 6 SER HA : -0.426: 0
: 2123:A 16 GLU HA :A 16 GLU OE1 : -0.471: 0
: 2123:A 15 GLU 1HG :A 11 TYR O : -0.452: 0
: 2123:A 54 VAL 3HG2 :A 50 ILE O : -0.442: 0
: 2123:A 113 LYS 1HE :A 109 GLU 1HG : -0.440: 0
: 2123:A 37 ALA O :A 41 LEU 3HD1 : -0.433: 0
: 2123:A 20 LYS 1HE :A 20 LYS 1HB : -0.425: 0
#sum2 ::24.02 clashscore : 24.02 clashscore B<40
#summary::2123 atoms:2123 atoms B<40:237180 potential dots:14820.0 A^2:51 bumps:51 bumps B<40:580.2 score
List of bad contacts calculated by MAGE for model $num_n
/farm/software/bin/probe
: 2123:A 107 LEU HG :A 89 LEU 1HD1 : -0.720: 0
: 2123:A 82 LEU HG :A 78 GLU O : -0.648: 0
: 2123:A 82 LEU 3HD1 :A 114 LEU 1HB : -0.536: 0
: 2123:A 78 GLU 1HB :A 76 ASN ND2 : -0.440: 0
: 2123:A 16 GLU 1HG :A 12 GLU O : -0.613: 0
: 2123:A 109 GLU 2HG :A 112 ARG 1HH2 : -0.596: 0
: 2123:A 112 ARG 1HH1 :A 112 ARG 1HD : -0.401: 0
: 2123:A 30 TYR CE2 :A 27 CYS HA : -0.592: 0
: 2123:A 27 CYS HA :A 30 TYR CZ : -0.584: 0
: 2123:A 51 THR OG1 :A 48 LYS HA : -0.552: 0
: 2123:A 101 GLU 2HB :A 105 LYS 1HE : -0.522: 0
: 2123:A 54 VAL 3HG2 :A 50 ILE O : -0.497: 0
: 2123:A 54 VAL 2HG2 :A 64 ASN ND2 : -0.439: 0
: 2123:A 64 ASN 1HD2 :A 54 VAL 2HG2 : -0.424: 0
: 2123:A 35 GLU 2HG :A 32 LYS HA : -0.493: 0
: 2123:A 35 GLU HA :A 83 TRP CZ3 : -0.465: 0
: 2123:A 35 GLU 1HG :A 83 TRP HH2 : -0.435: 0
: 2123:A 47 LEU 1HD1 :A 72 LEU 1HD2 : -0.490: 0
: 2123:A 47 LEU 1HB :A 42 VAL 3HG1 : -0.454: 0
: 2123:A 42 VAL 3HG2 :A 38 ILE O : -0.453: 0
: 2123:A 68 ALA 1HB :A 42 VAL 1HG2 : -0.436: 0
: 2123:A 87 TRP HZ3 :A 62 SER 2HB : -0.488: 0
: 2123:A 65 LEU 2HD1 :A 62 SER HA : -0.481: 0
: 2123:A 91 VAL 3HG2 :A 87 TRP O : -0.477: 0
: 2123:A 2 SER 2HB :A 124 HIS 2HB : -0.487: 0
: 2123:A 39 LYS O :A 43 ILE 2HG1 : -0.481: 0
: 2123:A 73 ARG O :A 73 ARG 1HD : -0.479: 0
: 2123:A 10 TYR 2HB :A 29 LYS O : -0.467: 0
: 2123:A 33 ALA 2HB :A 10 TYR 1HB : -0.465: 0
: 2123:A 17 PHE HE2 :A 29 LYS 2HE : -0.436: 0
: 2123:A 88 THR O :A 92 GLU 1HG : -0.453: 0
: 2123:A 94 PHE CE2 :A 92 GLU 1HB : -0.441: 0
: 2123:A 123 GLU OE1 :A 126 HIS 2HB : -0.452: 0
: 2123:A 79 ILE HB :A 80 PRO CD : -0.449: 0
: 2123:A 111 VAL O :A 115 VAL 3HG2 : -0.444: 0
: 2123:A 95 HIS O :A 96 GLU 1HB : -0.439: 0
: 2123:A 9 VAL 3HG2 :A 5 THR O : -0.429: 0
: 2123:A 67 LYS 2HE :A 67 LYS CA : -0.420: 0
: 2123:A 129 HIS 1HB :A 128 HIS O : -0.401: 0
#sum2 ::18.37 clashscore : 18.37 clashscore B<40
#summary::2123 atoms:2123 atoms B<40:237358 potential dots:14830.0 A^2:39 bumps:39 bumps B<40:574 score
List of bad contacts calculated by MAGE for model $num_n
/farm/software/bin/probe
: 2123:A 32 LYS HA :A 35 GLU 1HG : -0.908: 0
: 2123:A 35 GLU 1HG :A 32 LYS CA : -0.574: 0
: 2123:A 35 GLU OE1 :A 32 LYS 1HG : -0.487: 0
: 2123:A 32 LYS HA :A 35 GLU CG : -0.439: 0
: 2123:A 104 VAL 1HG2 :A 23 LEU 2HD2 : -0.848: 0
: 2123:A 42 VAL 2HG2 :A 47 LEU 2HD1 : -0.786: 0
: 2123:A 42 VAL 3HG2 :A 38 ILE O : -0.582: 0
: 2123:A 54 VAL 3HG2 :A 50 ILE O : -0.528: 0
: 2123:A 64 ASN OD1 :A 50 ILE 3HG2 : -0.527: 0
: 2123:A 47 LEU 1HB :A 42 VAL 3HG1 : -0.483: 0
: 2123:A 50 ILE HB :A 47 LEU 1HB : -0.412: 0
: 2123:A 97 LEU 1HB :A 93 GLY 1HA : -0.726: 0
: 2123:A 93 GLY 1HA :A 97 LEU CB : -0.446: 0
: 2123:A 45 ASN 2HB :A 41 LEU O : -0.689: 0
: 2123:A 107 LEU HG :A 89 LEU 1HD2 : -0.675: 0
: 2123:A 85 SER 1HB :A 107 LEU CD1 : -0.445: 0
: 2123:A 107 LEU 3HD1 :A 107 LEU HA : -0.432: 0
: 2123:A 82 LEU HG :A 78 GLU O : -0.657: 0
: 2123:A 113 LYS 1HB :A 110 ASP O : -0.443: 0
: 2123:A 82 LEU 2HD2 :A 110 ASP O : -0.434: 0
: 2123:A 81 ILE 2HD1 :A 78 GLU HA : -0.433: 0
: 2123:A 80 PRO 2HG :A 73 ARG 1HB : -0.624: 0
: 2123:A 77 THR HA :A 73 ARG 2HG : -0.602: 0
: 2123:A 76 ASN 2HB :A 79 ILE 2HG1 : -0.478: 0
: 2123:A 80 PRO 1HD :A 79 ILE N : -0.451: 0
: 2123:A 80 PRO 2HG :A 73 ARG CB : -0.441: 0
: 2123:A 114 LEU 1HD2 :A 79 ILE 3HG2 : -0.414: 0
: 2123:A 124 HIS 1HB :A 121 SER O : -0.566: 0
: 2123:A 30 TYR 2HB :A 111 VAL 1HG2 : -0.565: 0
: 2123:A 119 VAL HA :A 122 LEU 3HD2 : -0.511: 0
: 2123:A 115 VAL O :A 119 VAL 3HG2 : -0.467: 0
: 2123:A 27 CYS HA :A 30 TYR HD2 : -0.429: 0
: 2123:A 111 VAL O :A 115 VAL 3HG2 : -0.425: 0
: 2123:A 108 LYS 1HD :A 109 GLU N : -0.554: 0
: 2123:A 118 ALA 2HB :A 37 ALA CB : -0.536: 0
: 2123:A 118 ALA 2HB :A 37 ALA HA : -0.469: 0
: 2123:A 39 LYS O :A 43 ILE 2HG1 : -0.518: 0
: 2123:A 39 LYS 2HB :A 36 GLU O : -0.445: 0
: 2123:A 31 TYR HA :A 86 ALA 1HB : -0.480: 0
: 2123:A 98 SER HA :A 24 VAL 2HG2 : -0.479: 0
: 2123:A 112 ARG O :A 116 ILE 2HG1 : -0.476: 0
: 2123:A 84 LYS 2HB :A 84 LYS NZ : -0.469: 0
: 2123:A 94 PHE 1HB :A 90 HIS O : -0.447: 0
: 2123:A 72 LEU HG :A 68 ALA O : -0.447: 0
#sum2 ::20.73 clashscore : 20.73 clashscore B<40
#summary::2123 atoms:2123 atoms B<40:237293 potential dots:14830.0 A^2:44 bumps:44 bumps B<40:603.1 score
List of bad contacts calculated by MAGE for model $num_n
/farm/software/bin/probe
: 2123:A 61 LYS 1HE :A 63 GLU 2HB : -0.773: 0
: 2123:A 64 ASN H :A 61 LYS 2HG : -0.537: 0
: 2123:A 61 LYS 1HE :A 63 GLU CB : -0.481: 0
: 2123:A 128 HIS HD2 :A 126 HIS 2HB : -0.733: 0
: 2123:A 126 HIS 2HB :A 128 HIS CD2 : -0.429: 0
: 2123:A 30 TYR CE2 :A 27 CYS HA : -0.695: 0
: 2123:A 27 CYS HA :A 30 TYR CZ : -0.522: 0
: 2123:A 111 VAL 1HG1 :A 30 TYR O : -0.511: 0
: 2123:A 115 VAL O :A 119 VAL 3HG2 : -0.463: 0
: 2123:A 9 VAL 3HG2 :A 5 THR O : -0.452: 0
: 2123:A 111 VAL O :A 115 VAL 3HG2 : -0.452: 0
: 2123:A 119 VAL 2HG2 :A 5 THR OG1 : -0.412: 0
: 2123:A 65 LEU 2HD1 :A 62 SER HA : -0.684: 0
: 2123:A 39 LYS O :A 43 ILE 2HG1 : -0.582: 0
: 2123:A 40 LEU HG :A 36 GLU O : -0.476: 0
: 2123:A 122 LEU 1HD1 :A 40 LEU 2HD2 : -0.468: 0
: 2123:A 39 LYS 2HD :A 36 GLU OE2 : -0.410: 0
: 2123:A 82 LEU HG :A 78 GLU O : -0.573: 0
: 2123:A 113 LYS 2HE :A 82 LEU 1HD1 : -0.533: 0
: 2123:A 113 LYS 2HD :A 110 ASP HA : -0.415: 0
: 2123:A 102 LYS 2HB :A 100 ASN OD1 : -0.552: 0
: 2123:A 106 LYS 1HB :A 106 LYS 2HE : -0.424: 0
: 2123:A 104 VAL 3HG2 :A 100 ASN O : -0.415: 0
: 2123:A 102 LYS O :A 106 LYS 2HG : -0.400: 0
: 2123:A 42 VAL 3HG2 :A 38 ILE O : -0.549: 0
: 2123:A 28 GLU 2HB :A 94 PHE CE1 : -0.539: 0
: 2123:A 28 GLU 2HB :A 94 PHE CZ : -0.525: 0
: 2123:A 120 ASN 1HB :A 116 ILE O : -0.536: 0
: 2123:A 25 GLN O :A 29 LYS 1HG : -0.533: 0
: 2123:A 10 TYR 2HB :A 29 LYS O : -0.504: 0
: 2123:A 17 PHE HE2 :A 29 LYS 2HE : -0.471: 0
: 2123:A 79 ILE CD1 :A 76 ASN 2HB : -0.508: 0
: 2123:A 76 ASN 2HB :A 79 ILE 3HD1 : -0.424: 0
: 2123:A 91 VAL 3HG2 :A 87 TRP O : -0.493: 0
: 2123:A 92 GLU 2HG :A 91 VAL 2HG1 : -0.418: 0
: 2123:A 80 PRO 2HG :A 73 ARG 1HB : -0.469: 0
: 2123:A 73 ARG 2HD :A 73 ARG C : -0.411: 0
: 2123:A 105 LYS 1HE :A 105 LYS 1HB : -0.467: 0
: 2123:A 51 THR OG1 :A 48 LYS HA : -0.451: 0
: 2123:A 55 LYS 1HB :A 51 THR O : -0.401: 0
: 2123:A 107 LEU HG :A 89 LEU 1HD1 : -0.449: 0
: 2123:A 107 LEU 1HD2 :A 85 SER 1HB : -0.434: 0
: 2123:A 35 GLU 1HG :A 83 TRP HH2 : -0.446: 0
: 2123:A 72 LEU HG :A 68 ALA O : -0.442: 0
: 2123:A 108 LYS O :A 108 LYS 1HD : -0.438: 0
: 2123:A 108 LYS 1HD :A 108 LYS C : -0.437: 0
: 2123:A 11 TYR CE1 :A 108 LYS 2HD : -0.433: 0
: 2123:A 37 ALA O :A 41 LEU 3HD1 : -0.436: 0
: 2123:A 32 LYS HA :A 32 LYS 2HD : -0.422: 0
#sum2 ::23.08 clashscore : 23.08 clashscore B<40
#summary::2123 atoms:2123 atoms B<40:237337 potential dots:14830.0 A^2:49 bumps:49 bumps B<40:610.5 score
List of bad contacts calculated by MAGE for model $num_n
/farm/software/bin/probe
: 2123:A 27 CYS HA :A 30 TYR CZ : -0.718: 0
: 2123:A 30 TYR CE2 :A 27 CYS HA : -0.544: 0
: 2123:A 27 CYS SG :A 104 VAL 2HG2 : -0.454: 0
: 2123:A 111 VAL 1HG2 :A 30 TYR HD1 : -0.449: 0
: 2123:A 111 VAL O :A 115 VAL 3HG2 : -0.428: 0
: 2123:A 50 ILE 3HD1 :A 47 LEU 2HB : -0.709: 0
: 2123:A 47 LEU 1HB :A 42 VAL 3HG1 : -0.618: 0
: 2123:A 72 LEU HG :A 68 ALA O : -0.545: 0
: 2123:A 54 VAL 3HG2 :A 50 ILE O : -0.528: 0
: 2123:A 68 ALA 1HB :A 42 VAL 2HG2 : -0.506: 0
: 2123:A 42 VAL 3HG2 :A 38 ILE O : -0.502: 0
: 2123:A 64 ASN 2HB :A 50 ILE 3HG2 : -0.478: 0
: 2123:A 42 VAL CG1 :A 47 LEU 1HB : -0.475: 0
: 2123:A 38 ILE 1HG1 :A 34 ALA O : -0.433: 0
: 2123:A 50 ILE N :A 50 ILE 2HD1 : -0.425: 0
: 2123:A 39 LYS 2HG :A 36 GLU O : -0.566: 0
: 2123:A 39 LYS 1HB :A 60 TRP CZ2 : -0.553: 0
: 2123:A 40 LEU HG :A 36 GLU O : -0.523: 0
: 2123:A 60 TRP HD1 :A 59 ARG 1HH1 : -0.503: 0
: 2123:A 40 LEU 1HB :A 37 ALA O : -0.473: 0
: 2123:A 39 LYS 1HB :A 60 TRP CE2 : -0.404: 0
: 2123:A 80 PRO 2HG :A 73 ARG 1HB : -0.547: 0
: 2123:A 77 THR HA :A 73 ARG 1HG : -0.537: 0
: 2123:A 120 ASN 1HB :A 116 ILE O : -0.528: 0
: 2123:A 9 VAL O :A 13 GLU 1HG : -0.482: 0
: 2123:A 105 LYS O :A 108 LYS 1HG : -0.471: 0
: 2123:A 109 GLU 2HG :A 108 LYS 2HG : -0.445: 0
: 2123:A 109 GLU HA :A 112 ARG CB : -0.429: 0
: 2123:A 113 LYS 2HD :A 82 LEU CD1 : -0.469: 0
: 2123:A 113 LYS 2HD :A 82 LEU 1HD1 : -0.451: 0
: 2123:A 100 ASN 1HB :A 102 LYS 2HB : -0.456: 0
: 2123:A 103 GLU OE2 :A 102 LYS 2HE : -0.400: 0
: 2123:A 10 TYR 2HB :A 29 LYS O : -0.449: 0
: 2123:A 29 LYS 1HG :A 17 PHE HE2 : -0.414: 0
: 2123:A 57 LYS O :A 57 LYS 2HD : -0.448: 0
: 2123:A 76 ASN 2HB :A 79 ILE 2HG1 : -0.437: 0
: 2123:A 81 ILE 2HD1 :A 78 GLU HA : -0.434: 0
: 2123:A 45 ASN O :A 46 ASN 1HB : -0.433: 0
: 2123:A 70 LYS 1HB :A 66 PHE O : -0.427: 0
: 2123:A 32 LYS 2HD :A 32 LYS HA : -0.419: 0
: 2123:A 15 GLU 1HG :A 11 TYR O : -0.409: 0
: 2123:A 84 LYS 2HD :A 84 LYS C : -0.409: 0
#sum2 ::19.78 clashscore : 19.78 clashscore B<40
#summary::2123 atoms:2123 atoms B<40:237132 potential dots:14820.0 A^2:42 bumps:42 bumps B<40:592.2 score
List of bad contacts calculated by MAGE for model $num_n
/farm/software/bin/probe
: 2123:A 107 LEU HG :A 89 LEU 1HD1 : -0.760: 0
: 2123:A 29 LYS 1HE :A 29 LYS HA : -0.701: 0
: 2123:A 29 LYS CE :A 29 LYS HA : -0.514: 0
: 2123:A 13 GLU CB :A 29 LYS 2HD : -0.450: 0
: 2123:A 30 TYR CE2 :A 27 CYS HA : -0.694: 0
: 2123:A 27 CYS HA :A 30 TYR CZ : -0.578: 0
: 2123:A 63 GLU HA :A 66 PHE CE2 : -0.688: 0
: 2123:A 63 GLU 2HG :A 64 ASN N : -0.456: 0
: 2123:A 63 GLU HA :A 66 PHE CZ : -0.423: 0
: 2123:A 63 GLU O :A 67 LYS 1HB : -0.403: 0
: 2123:A 32 LYS HA :A 32 LYS 2HE : -0.654: 0
: 2123:A 28 GLU O :A 32 LYS 1HG : -0.536: 0
: 2123:A 36 GLU HA :A 39 LYS 2HE : -0.612: 0
: 2123:A 39 LYS O :A 43 ILE 2HG1 : -0.508: 0
: 2123:A 40 LEU HG :A 36 GLU O : -0.500: 0
: 2123:A 42 VAL 3HG2 :A 38 ILE O : -0.585: 0
: 2123:A 50 ILE 1HG2 :A 68 ALA 2HB : -0.585: 0
: 2123:A 47 LEU 1HB :A 42 VAL 3HG1 : -0.539: 0
: 2123:A 38 ILE 2HG1 :A 83 TRP CE3 : -0.533: 0
: 2123:A 53 ASN 2HB :A 58 GLY 2HA : -0.504: 0
: 2123:A 53 ASN 1HB :A 50 ILE O : -0.471: 0
: 2123:A 47 LEU 2HD1 :A 42 VAL 3HG1 : -0.468: 0
: 2123:A 31 TYR O :A 35 GLU 2HG : -0.464: 0
: 2123:A 35 GLU 1HG :A 83 TRP HH2 : -0.439: 0
: 2123:A 31 TYR OH :A 90 HIS 1HB : -0.420: 0
: 2123:A 31 TYR HE2 :A 90 HIS HD2 : -0.404: 0
: 2123:A 83 TRP CH2 :A 35 GLU 1HG : -0.404: 0
: 2123:A 15 GLU 1HG :A 11 TYR O : -0.582: 0
: 2123:A 105 LYS O :A 109 GLU 2HG : -0.527: 0
: 2123:A 11 TYR OH :A 108 LYS 2HG : -0.490: 0
: 2123:A 108 LYS 2HB :A 105 LYS HA : -0.462: 0
: 2123:A 105 LYS 1HB :A 102 LYS O : -0.431: 0
: 2123:A 7 ALA 1HB :A 115 VAL 2HG2 : -0.572: 0
: 2123:A 111 VAL O :A 115 VAL 3HG2 : -0.515: 0
: 2123:A 115 VAL O :A 119 VAL 3HG2 : -0.501: 0
: 2123:A 34 ALA 2HB :A 111 VAL 2HG2 : -0.487: 0
: 2123:A 87 TRP HH2 :A 62 SER 2HB : -0.555: 0
: 2123:A 65 LEU 2HD1 :A 62 SER HA : -0.416: 0
: 2123:A 80 PRO 2HG :A 73 ARG 1HB : -0.554: 0
: 2123:A 76 ASN 2HB :A 79 ILE 2HG1 : -0.529: 0
: 2123:A 77 THR HA :A 73 ARG 1HG : -0.509: 0
: 2123:A 79 ILE HB :A 80 PRO CD : -0.462: 0
: 2123:A 79 ILE HB :A 80 PRO 2HD : -0.429: 0
: 2123:A 104 VAL 3HG2 :A 100 ASN O : -0.516: 0
: 2123:A 103 GLU 2HG :A 106 LYS 2HD : -0.503: 0
: 2123:A 125 HIS 2HB :A 124 HIS O : -0.485: 0
: 2123:A 124 HIS HD2 :A 122 LEU HA : -0.417: 0
: 2123:A 127 HIS 1HB :A 126 HIS O : -0.472: 0
: 2123:A 6 SER O :A 10 TYR HD2 : -0.470: 0
: 2123:A 33 ALA 2HB :A 10 TYR 1HB : -0.440: 0
: 2123:A 9 VAL 3HG2 :A 5 THR O : -0.451: 0
: 2123:A 82 LEU HG :A 78 GLU O : -0.438: 0
: 2123:A 44 GLU 2HG :A 45 ASN 2HD2 : -0.410: 0
: 2123:A 37 ALA O :A 41 LEU 3HD1 : -0.410: 0
: 2123:A 97 LEU 3HD1 :A 97 LEU HA : -0.407: 0
#sum2 ::25.91 clashscore : 25.91 clashscore B<40
#summary::2123 atoms:2123 atoms B<40:237192 potential dots:14820.0 A^2:55 bumps:55 bumps B<40:514.8 score
List of bad contacts calculated by MAGE for model $num_n
/farm/software/bin/probe
: 2123:A 32 LYS HA :A 35 GLU 1HG : -0.861: 0
: 2123:A 35 GLU 1HG :A 32 LYS CA : -0.525: 0
: 2123:A 35 GLU OE2 :A 32 LYS 1HG : -0.424: 0
: 2123:A 30 TYR CE2 :A 27 CYS HA : -0.756: 0
: 2123:A 111 VAL 1HG2 :A 30 TYR HD1 : -0.592: 0
: 2123:A 111 VAL 1HG2 :A 30 TYR CD1 : -0.468: 0
: 2123:A 111 VAL O :A 115 VAL 3HG2 : -0.424: 0
: 2123:A 107 LEU O :A 111 VAL 3HG2 : -0.409: 0
: 2123:A 107 LEU 3HD1 :A 107 LEU HA : -0.409: 0
: 2123:A 107 LEU 1HD2 :A 85 SER 1HB : -0.402: 0
: 2123:A 96 GLU 2HG :A 97 LEU 2HD2 : -0.716: 0
: 2123:A 61 LYS 2HB :A 63 GLU 1HG : -0.649: 0
: 2123:A 61 LYS 2HB :A 63 GLU CG : -0.551: 0
: 2123:A 64 ASN ND2 :A 61 LYS 1HB : -0.483: 0
: 2123:A 50 ILE 3HD1 :A 47 LEU 2HB : -0.624: 0
: 2123:A 42 VAL 2HG2 :A 47 LEU 2HD1 : -0.623: 0
: 2123:A 47 LEU 1HB :A 42 VAL 3HG1 : -0.617: 0
: 2123:A 42 VAL 3HG2 :A 38 ILE O : -0.453: 0
: 2123:A 54 VAL 3HG2 :A 50 ILE O : -0.431: 0
: 2123:A 42 VAL HB :A 60 TRP HH2 : -0.417: 0
: 2123:A 17 PHE HE2 :A 29 LYS 2HE : -0.613: 0
: 2123:A 25 GLN O :A 29 LYS 2HD : -0.572: 0
: 2123:A 10 TYR 2HB :A 29 LYS O : -0.525: 0
: 2123:A 22 ASP 2HB :A 17 PHE 2HB : -0.512: 0
: 2123:A 17 PHE 1HB :A 26 ALA 2HB : -0.500: 0
: 2123:A 29 LYS 2HE :A 17 PHE CE2 : -0.435: 0
: 2123:A 18 LEU 1HB :A 14 ALA O : -0.608: 0
: 2123:A 36 GLU 1HG :A 6 SER 2HB : -0.602: 0
: 2123:A 43 ILE 1HG1 :A 39 LYS O : -0.571: 0
: 2123:A 39 LYS 2HB :A 36 GLU O : -0.416: 0
: 2123:A 122 LEU 3HD2 :A 3 ILE 2HD1 : -0.572: 0
: 2123:A 82 LEU HG :A 78 GLU O : -0.547: 0
: 2123:A 82 LEU 2HD2 :A 110 ASP 2HB : -0.514: 0
: 2123:A 79 ILE HB :A 80 PRO CD : -0.430: 0
: 2123:A 80 PRO 1HD :A 79 ILE N : -0.429: 0
: 2123:A 78 GLU 1HB :A 76 ASN OD1 : -0.416: 0
: 2123:A 79 ILE 2HG1 :A 76 ASN OD1 : -0.410: 0
: 2123:A 77 THR HA :A 73 ARG 1HG : -0.540: 0
: 2123:A 73 ARG O :A 73 ARG 2HD : -0.404: 0
: 2123:A 5 THR O :A 8 GLU 1HG : -0.512: 0
: 2123:A 9 VAL 3HG2 :A 5 THR O : -0.432: 0
: 2123:A 126 HIS H :A 124 HIS C : -0.498: 0
: 2123:A 52 ASN O :A 55 LYS 1HB : -0.489: 0
: 2123:A 52 ASN ND2 :A 48 LYS 1HG : -0.436: 0
: 2123:A 84 LYS 2HD :A 84 LYS C : -0.484: 0
: 2123:A 118 ALA 2HB :A 37 ALA 1HB : -0.472: 0
: 2123:A 117 PHE CD2 :A 41 LEU 1HD2 : -0.449: 0
: 2123:A 37 ALA O :A 41 LEU 3HD1 : -0.432: 0
: 2123:A 41 LEU CD1 :A 118 ALA HA : -0.420: 0
: 2123:A 117 PHE HD2 :A 41 LEU 1HD2 : -0.411: 0
: 2123:A 99 LEU N :A 99 LEU 2HD2 : -0.463: 0
: 2123:A 120 ASN 1HB :A 116 ILE O : -0.443: 0
: 2123:A 70 LYS 2HD :A 70 LYS C : -0.427: 0
: 2123:A 20 LYS 2HE :A 20 LYS 1HB : -0.406: 0
#sum2 ::25.44 clashscore : 25.44 clashscore B<40
#summary::2123 atoms:2123 atoms B<40:237196 potential dots:14820.0 A^2:54 bumps:54 bumps B<40:515.9 score
List of bad contacts calculated by MAGE for model $num_n
/farm/software/bin/probe
: 2123:A 32 LYS 1HG :A 29 LYS HA : -0.684: 0
: 2123:A 29 LYS 2HD :A 13 GLU 1HB : -0.663: 0
: 2123:A 29 LYS HA :A 32 LYS CG : -0.478: 0
: 2123:A 30 TYR CE2 :A 14 ALA 1HB : -0.471: 0
: 2123:A 25 GLN O :A 29 LYS 1HG : -0.446: 0
: 2123:A 28 GLU O :A 32 LYS 1HG : -0.431: 0
: 2123:A 14 ALA 2HB :A 29 LYS 2HG : -0.429: 0
: 2123:A 29 LYS 2HD :A 13 GLU CB : -0.401: 0
: 2123:A 97 LEU 1HB :A 93 GLY 1HA : -0.682: 0
: 2123:A 97 LEU 2HD2 :A 96 GLU H : -0.432: 0
: 2123:A 51 THR OG1 :A 48 LYS HA : -0.596: 0
: 2123:A 5 THR 1HG2 :A 40 LEU 1HD2 : -0.567: 0
: 2123:A 9 VAL 3HG2 :A 5 THR O : -0.425: 0
: 2123:A 36 GLU O :A 40 LEU 3HD1 : -0.420: 0
: 2123:A 6 SER H :A 5 THR 2HG2 : -0.412: 0
: 2123:A 65 LEU 3HD1 :A 35 GLU 1HB : -0.549: 0
: 2123:A 16 GLU 1HG :A 12 GLU O : -0.532: 0
: 2123:A 82 LEU HG :A 78 GLU O : -0.532: 0
: 2123:A 4 SER N :A 3 ILE 2HG2 : -0.530: 0
: 2123:A 3 ILE 2HG2 :A 4 SER H : -0.437: 0
: 2123:A 107 LEU HG :A 89 LEU 1HD1 : -0.527: 0
: 2123:A 104 VAL 3HG2 :A 100 ASN O : -0.514: 0
: 2123:A 104 VAL O :A 107 LEU 1HB : -0.454: 0
: 2123:A 81 ILE O :A 84 LYS 2HB : -0.504: 0
: 2123:A 15 GLU 1HG :A 11 TYR O : -0.499: 0
: 2123:A 69 SER OG :A 80 PRO HA : -0.494: 0
: 2123:A 80 PRO 1HD :A 79 ILE N : -0.463: 0
: 2123:A 118 ALA O :A 122 LEU HG : -0.493: 0
: 2123:A 39 LYS O :A 43 ILE 2HG1 : -0.446: 0
: 2123:A 42 VAL 3HG2 :A 38 ILE O : -0.443: 0
: 2123:A 53 ASN 1HB :A 50 ILE O : -0.441: 0
: 2123:A 86 ALA 1HB :A 31 TYR HD1 : -0.434: 0
: 2123:A 115 VAL O :A 119 VAL 3HG2 : -0.418: 0
: 2123:A 70 LYS 2HD :A 70 LYS O : -0.407: 0
#sum2 ::16.02 clashscore : 16.02 clashscore B<40
#summary::2123 atoms:2123 atoms B<40:237167 potential dots:14820.0 A^2:34 bumps:34 bumps B<40:605.6 score
List of bad contacts calculated by MAGE for model $num_n
/farm/software/bin/probe
: 2123:A 44 GLU 2HG :A 121 SER 1HB : -1.015: 0
: 2123:A 32 LYS HA :A 35 GLU 1HG : -0.907: 0
: 2123:A 36 GLU HA :A 39 LYS 2HE : -0.767: 0
: 2123:A 54 VAL 3HG2 :A 50 ILE O : -0.640: 0
: 2123:A 35 GLU CG :A 32 LYS HA : -0.613: 0
: 2123:A 35 GLU 1HG :A 32 LYS CA : -0.587: 0
: 2123:A 68 ALA 1HB :A 42 VAL 1HG2 : -0.570: 0
: 2123:A 47 LEU 1HB :A 42 VAL 3HG1 : -0.521: 0
: 2123:A 50 ILE 2HG1 :A 68 ALA HA : -0.491: 0
: 2123:A 38 ILE HB :A 65 LEU 2HD2 : -0.471: 0
: 2123:A 42 VAL 3HG2 :A 38 ILE O : -0.469: 0
: 2123:A 36 GLU N :A 35 GLU 2HG : -0.460: 0
: 2123:A 65 LEU CD1 :A 35 GLU 1HB : -0.413: 0
: 2123:A 60 TRP HH2 :A 42 VAL 1HG1 : -0.409: 0
: 2123:A 40 LEU 3HD1 :A 36 GLU O : -0.408: 0
: 2123:A 64 ASN ND2 :A 54 VAL 3HG1 : -0.408: 0
: 2123:A 113 LYS 2HD :A 110 ASP HA : -0.815: 0
: 2123:A 113 LYS 1HB :A 110 ASP O : -0.509: 0
: 2123:A 110 ASP HA :A 113 LYS CD : -0.423: 0
: 2123:A 89 LEU 1HD1 :A 30 TYR HE2 : -0.686: 0
: 2123:A 109 GLU 2HG :A 108 LYS 1HD : -0.683: 0
: 2123:A 85 SER 2HB :A 107 LEU 1HD1 : -0.631: 0
: 2123:A 89 LEU 2HD1 :A 90 HIS N : -0.572: 0
: 2123:A 107 LEU 1HD2 :A 85 SER 1HB : -0.571: 0
: 2123:A 107 LEU HG :A 89 LEU CD2 : -0.541: 0
: 2123:A 108 LYS HA :A 30 TYR HE1 : -0.534: 0
: 2123:A 105 LYS O :A 108 LYS 2HG : -0.475: 0
: 2123:A 89 LEU 1HD1 :A 30 TYR CE2 : -0.460: 0
: 2123:A 107 LEU 2HB :A 30 TYR OH : -0.455: 0
: 2123:A 27 CYS HA :A 30 TYR 2HB : -0.431: 0
: 2123:A 89 LEU HG :A 86 ALA HA : -0.413: 0
: 2123:A 108 LYS 1HG :A 105 LYS HA : -0.401: 0
: 2123:A 120 ASN 1HB :A 116 ILE O : -0.645: 0
: 2123:A 119 VAL O :A 123 GLU 2HG : -0.550: 0
: 2123:A 111 VAL O :A 115 VAL 3HG2 : -0.438: 0
: 2123:A 112 ARG O :A 116 ILE 2HG1 : -0.431: 0
: 2123:A 123 GLU OE2 :A 120 ASN HA : -0.428: 0
: 2123:A 115 VAL O :A 119 VAL 3HG2 : -0.403: 0
: 2123:A 92 GLU 2HG :A 96 GLU 1HB : -0.637: 0
: 2123:A 94 PHE 2HB :A 96 GLU OE2 : -0.461: 0
: 2123:A 91 VAL 3HG2 :A 87 TRP O : -0.625: 0
: 2123:A 76 ASN 2HB :A 79 ILE 2HG1 : -0.583: 0
: 2123:A 80 PRO 1HD :A 79 ILE N : -0.455: 0
: 2123:A 82 LEU HG :A 78 GLU O : -0.569: 0
: 2123:A 77 THR HA :A 73 ARG 2HG : -0.557: 0
: 2123:A 71 LEU C :A 73 ARG H : -0.454: 0
: 2123:A 6 SER 2HB :A 10 TYR CE2 : -0.536: 0
: 2123:A 106 LYS 2HG :A 102 LYS O : -0.500: 0
: 2123:A 9 VAL 3HG2 :A 5 THR O : -0.454: 0
: 2123:A 5 THR O :A 8 GLU 1HG : -0.412: 0
: 2123:A 41 LEU HG :A 37 ALA O : -0.452: 0
: 2123:A 127 HIS 1HB :A 126 HIS O : -0.428: 0
: 2123:A 104 VAL 3HG2 :A 100 ASN O : -0.426: 0
: 2123:A 122 LEU C :A 124 HIS H : -0.401: 0
#sum2 ::25.44 clashscore : 25.44 clashscore B<40
#summary::2123 atoms:2123 atoms B<40:237390 potential dots:14840.0 A^2:54 bumps:54 bumps B<40:515.8 score
List of bad contacts calculated by MAGE for model $num_n
/farm/software/bin/probe
: 2123:A 57 LYS 2HE :A 61 LYS 1HD : -0.955: 0
: 2123:A 65 LEU 2HD1 :A 62 SER HA : -0.677: 0
: 2123:A 65 LEU HG :A 61 LYS O : -0.486: 0
: 2123:A 107 LEU HG :A 89 LEU 1HD1 : -0.875: 0
: 2123:A 30 TYR 2HB :A 111 VAL 1HG2 : -0.520: 0
: 2123:A 107 LEU O :A 111 VAL 3HG2 : -0.456: 0
: 2123:A 80 PRO 2HG :A 73 ARG 1HB : -0.768: 0
: 2123:A 80 PRO 2HB :A 70 LYS HA : -0.501: 0
: 2123:A 79 ILE HB :A 80 PRO CD : -0.403: 0
: 2123:A 47 LEU 1HD1 :A 72 LEU 1HD2 : -0.711: 0
: 2123:A 47 LEU 1HB :A 42 VAL 3HG1 : -0.626: 0
: 2123:A 42 VAL 3HG2 :A 38 ILE O : -0.537: 0
: 2123:A 38 ILE 2HD1 :A 69 SER 1HB : -0.482: 0
: 2123:A 72 LEU 3HD1 :A 41 LEU 3HD2 : -0.426: 0
: 2123:A 32 LYS HA :A 35 GLU 1HG : -0.667: 0
: 2123:A 35 GLU 1HG :A 32 LYS CA : -0.439: 0
: 2123:A 5 THR 1HG2 :A 40 LEU 1HD2 : -0.663: 0
: 2123:A 52 ASN HA :A 55 LYS 2HD : -0.653: 0
: 2123:A 52 ASN O :A 55 LYS 1HB : -0.586: 0
: 2123:A 82 LEU HG :A 78 GLU O : -0.631: 0
: 2123:A 78 GLU 1HB :A 76 ASN ND2 : -0.540: 0
: 2123:A 82 LEU 1HD1 :A 78 GLU 2HB : -0.458: 0
: 2123:A 82 LEU CB :A 114 LEU 1HD1 : -0.419: 0
: 2123:A 82 LEU 2HB :A 114 LEU 1HD1 : -0.418: 0
: 2123:A 78 GLU 2HB :A 82 LEU CD1 : -0.407: 0
: 2123:A 44 GLU 2HG :A 122 LEU 2HD1 : -0.563: 0
: 2123:A 25 GLN OE1 :A 24 VAL HB : -0.550: 0
: 2123:A 25 GLN O :A 29 LYS 1HG : -0.492: 0
: 2123:A 51 THR OG1 :A 48 LYS HA : -0.546: 0
: 2123:A 11 TYR HE2 :A 112 ARG 1HH1 : -0.544: 0
: 2123:A 113 LYS 1HG :A 109 GLU O : -0.436: 0
: 2123:A 113 LYS CG :A 110 ASP HA : -0.415: 0
: 2123:A 109 GLU O :A 112 ARG 2HB : -0.410: 0
: 2123:A 101 GLU O :A 105 LYS 1HB : -0.530: 0
: 2123:A 120 ASN 1HB :A 116 ILE O : -0.519: 0
: 2123:A 123 GLU 2HG :A 120 ASN O : -0.410: 0
: 2123:A 16 GLU 1HG :A 12 GLU O : -0.487: 0
: 2123:A 91 VAL 3HG2 :A 87 TRP O : -0.480: 0
: 2123:A 115 VAL O :A 119 VAL 3HG2 : -0.454: 0
: 2123:A 118 ALA O :A 121 SER 2HB : -0.417: 0
: 2123:A 66 PHE HA :A 83 TRP CD1 : -0.416: 0
#sum2 ::19.31 clashscore : 19.31 clashscore B<40
#summary::2123 atoms:2123 atoms B<40:237214 potential dots:14830.0 A^2:41 bumps:41 bumps B<40:522.3 score
List of bad contacts calculated by MAGE for model $num_n
/farm/software/bin/probe
: 2123:A 89 LEU 3HD2 :A 107 LEU HG : -0.988: 0
: 2123:A 89 LEU 3HD2 :A 107 LEU CG : -0.696: 0
: 2123:A 30 TYR CE2 :A 27 CYS HA : -0.664: 0
: 2123:A 89 LEU 2HD1 :A 90 HIS N : -0.558: 0
: 2123:A 85 SER 1HB :A 107 LEU CD1 : -0.424: 0
: 2123:A 89 LEU HG :A 86 ALA HA : -0.411: 0
: 2123:A 27 CYS SG :A 89 LEU 3HD1 : -0.407: 0
: 2123:A 122 LEU 3HD1 :A 3 ILE 2HG1 : -0.850: 0
: 2123:A 122 LEU 2HD1 :A 119 VAL HA : -0.542: 0
: 2123:A 7 ALA 1HB :A 115 VAL 3HG1 : -0.498: 0
: 2123:A 3 ILE N :A 3 ILE 3HD1 : -0.451: 0
: 2123:A 3 ILE H :A 3 ILE 3HD1 : -0.427: 0
: 2123:A 2 SER 2HB :A 3 ILE 3HD1 : -0.421: 0
: 2123:A 115 VAL O :A 119 VAL 3HG2 : -0.414: 0
: 2123:A 57 LYS 1HE :A 57 LYS HA : -0.830: 0
: 2123:A 57 LYS 1HE :A 57 LYS CA : -0.416: 0
: 2123:A 96 GLU 1HG :A 99 LEU 3HD2 : -0.822: 0
: 2123:A 82 LEU HG :A 78 GLU O : -0.756: 0
: 2123:A 113 LYS 1HB :A 110 ASP O : -0.493: 0
: 2123:A 82 LEU 2HD2 :A 110 ASP O : -0.463: 0
: 2123:A 32 LYS HA :A 35 GLU 1HG : -0.747: 0
: 2123:A 17 PHE HE2 :A 29 LYS 2HE : -0.603: 0
: 2123:A 10 TYR CB :A 33 ALA 2HB : -0.471: 0
: 2123:A 17 PHE 1HB :A 26 ALA 2HB : -0.468: 0
: 2123:A 10 TYR 2HB :A 29 LYS O : -0.426: 0
: 2123:A 10 TYR CE1 :A 32 LYS 2HE : -0.425: 0
: 2123:A 44 GLU 1HG :A 41 LEU HA : -0.614: 0
: 2123:A 44 GLU 2HG :A 45 ASN N : -0.436: 0
: 2123:A 39 LYS 1HE :A 40 LEU 2HD1 : -0.611: 0
: 2123:A 39 LYS 2HB :A 36 GLU O : -0.446: 0
: 2123:A 43 ILE 2HG1 :A 39 LYS O : -0.408: 0
: 2123:A 112 ARG O :A 116 ILE 1HG1 : -0.597: 0
: 2123:A 112 ARG 1HH2 :A 109 GLU CD : -0.480: 0
: 2123:A 79 ILE HB :A 80 PRO 2HD : -0.590: 0
: 2123:A 79 ILE CD1 :A 76 ASN 2HB : -0.555: 0
: 2123:A 77 THR O :A 80 PRO 1HD : -0.482: 0
: 2123:A 76 ASN 2HB :A 79 ILE 3HD1 : -0.455: 0
: 2123:A 105 LYS 1HG :A 101 GLU O : -0.587: 0
: 2123:A 42 VAL 3HG2 :A 38 ILE O : -0.547: 0
: 2123:A 47 LEU 1HB :A 42 VAL 3HG1 : -0.521: 0
: 2123:A 114 LEU 1HD1 :A 38 ILE 1HD1 : -0.510: 0
: 2123:A 47 LEU 1HD1 :A 72 LEU 1HD2 : -0.408: 0
: 2123:A 20 LYS 1HE :A 20 LYS HA : -0.519: 0
: 2123:A 51 THR OG1 :A 48 LYS HA : -0.515: 0
: 2123:A 53 ASN 2HB :A 50 ILE O : -0.506: 0
: 2123:A 50 ILE 1HG2 :A 68 ALA 2HB : -0.450: 0
: 2123:A 54 VAL 3HG2 :A 50 ILE O : -0.440: 0
: 2123:A 16 GLU 1HG :A 12 GLU O : -0.474: 0
: 2123:A 13 GLU O :A 16 GLU 1HB : -0.421: 0
: 2123:A 23 LEU 1HB :A 98 SER 1HB : -0.473: 0
: 2123:A 98 SER HA :A 24 VAL CG2 : -0.450: 0
: 2123:A 91 VAL 3HG2 :A 87 TRP O : -0.460: 0
: 2123:A 111 VAL 3HG1 :A 34 ALA 2HB : -0.452: 0
: 2123:A 49 GLU HA :A 52 ASN ND2 : -0.442: 0
: 2123:A 65 LEU 2HD1 :A 62 SER HA : -0.433: 0
#sum2 ::25.91 clashscore : 25.91 clashscore B<40
#summary::2123 atoms:2123 atoms B<40:237421 potential dots:14840.0 A^2:55 bumps:55 bumps B<40:519.8 score
List of bad contacts calculated by MAGE for model $num_n
/farm/software/bin/probe
: 2123:A 113 LYS 2HD :A 110 ASP HA : -0.939: 0
: 2123:A 113 LYS 1HB :A 110 ASP O : -0.420: 0
: 2123:A 32 LYS HA :A 35 GLU 1HG : -0.782: 0
: 2123:A 39 LYS 1HG :A 36 GLU O : -0.605: 0
: 2123:A 32 LYS O :A 36 GLU 1HG : -0.590: 0
: 2123:A 83 TRP CH2 :A 35 GLU 2HB : -0.484: 0
: 2123:A 36 GLU 2HB :A 6 SER 2HB : -0.435: 0
: 2123:A 118 ALA HA :A 41 LEU 1HD2 : -0.732: 0
: 2123:A 51 THR OG1 :A 48 LYS HA : -0.665: 0
: 2123:A 52 ASN ND2 :A 48 LYS 1HG : -0.490: 0
: 2123:A 52 ASN HA :A 55 LYS 2HD : -0.410: 0
: 2123:A 30 TYR CE2 :A 27 CYS HA : -0.652: 0
: 2123:A 27 CYS SG :A 104 VAL 2HG2 : -0.618: 0
: 2123:A 104 VAL 3HG2 :A 100 ASN O : -0.571: 0
: 2123:A 27 CYS HA :A 30 TYR CZ : -0.516: 0
: 2123:A 77 THR HA :A 73 ARG 2HG : -0.610: 0
: 2123:A 54 VAL 3HG2 :A 50 ILE O : -0.573: 0
: 2123:A 54 VAL O :A 54 VAL 2HG1 : -0.410: 0
: 2123:A 82 LEU HG :A 78 GLU O : -0.572: 0
: 2123:A 38 ILE 1HG1 :A 34 ALA O : -0.552: 0
: 2123:A 42 VAL 3HG2 :A 38 ILE O : -0.549: 0
: 2123:A 47 LEU 1HB :A 42 VAL 3HG1 : -0.537: 0
: 2123:A 112 ARG O :A 116 ILE 2HG1 : -0.517: 0
: 2123:A 120 ASN 1HB :A 116 ILE O : -0.445: 0
: 2123:A 25 GLN O :A 29 LYS 1HG : -0.507: 0
: 2123:A 79 ILE 1HG1 :A 76 ASN 2HB : -0.500: 0
: 2123:A 79 ILE HB :A 80 PRO CD : -0.483: 0
: 2123:A 79 ILE HB :A 80 PRO 2HD : -0.443: 0
: 2123:A 69 SER OG :A 80 PRO HA : -0.442: 0
: 2123:A 44 GLU 1HG :A 40 LEU O : -0.499: 0
: 2123:A 91 VAL 3HG2 :A 87 TRP O : -0.497: 0
: 2123:A 87 TRP HH2 :A 62 SER 1HB : -0.400: 0
: 2123:A 72 LEU HG :A 68 ALA O : -0.496: 0
: 2123:A 59 ARG H :A 57 LYS 2HB : -0.486: 0
: 2123:A 119 VAL HA :A 122 LEU 3HD1 : -0.475: 0
: 2123:A 111 VAL O :A 115 VAL 3HG2 : -0.454: 0
: 2123:A 115 VAL O :A 119 VAL 3HG2 : -0.449: 0
: 2123:A 9 VAL 3HG2 :A 5 THR O : -0.420: 0
: 2123:A 122 LEU 1HD2 :A 5 THR 1HG2 : -0.414: 0
: 2123:A 117 PHE CZ :A 75 ASN 2HB : -0.474: 0
: 2123:A 99 LEU N :A 99 LEU 2HD2 : -0.461: 0
: 2123:A 107 LEU HA :A 107 LEU 3HD1 : -0.403: 0
#sum2 ::19.78 clashscore : 19.78 clashscore B<40
#summary::2123 atoms:2123 atoms B<40:237216 potential dots:14830.0 A^2:42 bumps:42 bumps B<40:549.1 score
List of bad contacts calculated by MAGE for model $num_n
/farm/software/bin/probe
: 2123:A 82 LEU HG :A 78 GLU O : -0.632: 0
: 2123:A 47 LEU 1HB :A 42 VAL 3HG1 : -0.617: 0
: 2123:A 42 VAL 3HG2 :A 38 ILE O : -0.524: 0
: 2123:A 42 VAL 1HG1 :A 60 TRP HH2 : -0.412: 0
: 2123:A 89 LEU 2HD2 :A 99 LEU 3HD1 : -0.580: 0
: 2123:A 107 LEU HG :A 89 LEU CD1 : -0.551: 0
: 2123:A 107 LEU HG :A 89 LEU 1HD1 : -0.498: 0
: 2123:A 107 LEU 1HD2 :A 85 SER 1HB : -0.436: 0
: 2123:A 79 ILE HB :A 80 PRO 2HD : -0.578: 0
: 2123:A 77 THR HA :A 73 ARG 1HG : -0.463: 0
: 2123:A 72 LEU 2HB :A 79 ILE CD1 : -0.417: 0
: 2123:A 77 THR O :A 80 PRO 1HD : -0.407: 0
: 2123:A 30 TYR 2HB :A 111 VAL 1HG2 : -0.544: 0
: 2123:A 115 VAL O :A 119 VAL 3HG2 : -0.521: 0
: 2123:A 108 LYS 2HD :A 108 LYS O : -0.484: 0
: 2123:A 111 VAL O :A 115 VAL 3HG2 : -0.465: 0
: 2123:A 108 LYS 2HD :A 108 LYS C : -0.451: 0
: 2123:A 30 TYR OH :A 108 LYS 1HG : -0.406: 0
: 2123:A 27 CYS HA :A 30 TYR 1HB : -0.400: 0
: 2123:A 64 ASN 2HB :A 54 VAL 2HG2 : -0.522: 0
: 2123:A 54 VAL 3HG2 :A 50 ILE O : -0.412: 0
: 2123:A 83 TRP CZ2 :A 65 LEU 3HD1 : -0.501: 0
: 2123:A 103 GLU CB :A 100 ASN 1HB : -0.498: 0
: 2123:A 103 GLU 1HB :A 100 ASN 1HB : -0.493: 0
: 2123:A 40 LEU CD1 :A 118 ALA 1HB : -0.482: 0
: 2123:A 40 LEU 2HD1 :A 118 ALA 1HB : -0.446: 0
: 2123:A 118 ALA O :A 121 SER 2HB : -0.419: 0
: 2123:A 15 GLU 1HG :A 11 TYR O : -0.476: 0
: 2123:A 51 THR O :A 55 LYS 2HD : -0.472: 0
: 2123:A 51 THR OG1 :A 48 LYS HA : -0.416: 0
: 2123:A 43 ILE 1HG1 :A 39 LYS O : -0.469: 0
: 2123:A 110 ASP OD2 :A 106 LYS 1HG : -0.441: 0
: 2123:A 41 LEU 1HD1 :A 117 PHE 2HB : -0.437: 0
: 2123:A 70 LYS 1HB :A 67 LYS O : -0.411: 0
: 2123:A 9 VAL 3HG2 :A 5 THR O : -0.409: 0
#sum2 ::16.49 clashscore : 16.49 clashscore B<40
#summary::2123 atoms:2123 atoms B<40:237198 potential dots:14820.0 A^2:35 bumps:35 bumps B<40:668.8 score
List of bad contacts calculated by MAGE for model $num_n
/farm/software/bin/probe
: 2123:A 63 GLU 1HG :A 61 LYS 1HD : -0.911: 0
: 2123:A 61 LYS 1HD :A 63 GLU CG : -0.466: 0
: 2123:A 107 LEU HG :A 89 LEU 1HD1 : -0.768: 0
: 2123:A 107 LEU 3HD2 :A 103 GLU 1HG : -0.515: 0
: 2123:A 85 SER 1HB :A 107 LEU CD1 : -0.402: 0
: 2123:A 30 TYR CE2 :A 27 CYS HA : -0.707: 0
: 2123:A 27 CYS HA :A 30 TYR CZ : -0.694: 0
: 2123:A 104 VAL 3HG2 :A 100 ASN O : -0.500: 0
: 2123:A 102 LYS 2HB :A 100 ASN OD1 : -0.458: 0
: 2123:A 104 VAL 3HG1 :A 30 TYR CE2 : -0.431: 0
: 2123:A 27 CYS SG :A 99 LEU 2HD1 : -0.424: 0
: 2123:A 30 TYR HE2 :A 104 VAL 3HG1 : -0.419: 0
: 2123:A 48 LYS HA :A 48 LYS 2HE : -0.630: 0
: 2123:A 48 LYS HA :A 48 LYS CE : -0.476: 0
: 2123:A 47 LEU 1HB :A 42 VAL 3HG1 : -0.623: 0
: 2123:A 42 VAL 3HG2 :A 38 ILE O : -0.499: 0
: 2123:A 42 VAL 2HG2 :A 47 LEU 2HD1 : -0.446: 0
: 2123:A 38 ILE 1HG1 :A 34 ALA O : -0.426: 0
: 2123:A 39 LYS 1HG :A 36 GLU O : -0.512: 0
: 2123:A 43 ILE 1HG1 :A 39 LYS O : -0.480: 0
: 2123:A 10 TYR CE2 :A 36 GLU 2HG : -0.458: 0
: 2123:A 36 GLU 2HG :A 10 TYR HE2 : -0.401: 0
: 2123:A 82 LEU HG :A 78 GLU O : -0.494: 0
: 2123:A 76 ASN 2HB :A 79 ILE 2HG1 : -0.491: 0
: 2123:A 80 PRO 1HD :A 79 ILE N : -0.422: 0
: 2123:A 79 ILE HB :A 80 PRO CD : -0.420: 0
: 2123:A 77 THR HA :A 73 ARG 1HG : -0.471: 0
: 2123:A 73 ARG 2HD :A 73 ARG C : -0.431: 0
: 2123:A 118 ALA O :A 121 SER 2HB : -0.461: 0
: 2123:A 32 LYS HA :A 35 GLU 1HG : -0.457: 0
: 2123:A 35 GLU CD :A 32 LYS 2HD : -0.450: 0
: 2123:A 9 VAL 3HG2 :A 5 THR O : -0.452: 0
: 2123:A 105 LYS 1HG :A 101 GLU 1HG : -0.446: 0
: 2123:A 105 LYS O :A 108 LYS 1HG : -0.405: 0
: 2123:A 111 VAL O :A 115 VAL 3HG2 : -0.444: 0
: 2123:A 115 VAL O :A 119 VAL 3HG2 : -0.427: 0
: 2123:A 113 LYS 1HB :A 110 ASP O : -0.413: 0
: 2123:A 37 ALA O :A 41 LEU 3HD1 : -0.413: 0
: 2123:A 91 VAL 3HG2 :A 87 TRP O : -0.411: 0
: 2123:A 24 VAL 3HG1 :A 97 LEU 2HB : -0.407: 0
: 2123:A 116 ILE O :A 120 ASN 1HB : -0.404: 0
#sum2 ::19.31 clashscore : 19.31 clashscore B<40
#summary::2123 atoms:2123 atoms B<40:237159 potential dots:14820.0 A^2:41 bumps:41 bumps B<40:630.5 score
List of bad contacts calculated by MAGE for model $num_n
/farm/software/bin/probe
: 2123:A 82 LEU HG :A 78 GLU O : -0.712: 0
: 2123:A 113 LYS 1HD :A 110 ASP HA : -0.696: 0
: 2123:A 113 LYS 1HB :A 110 ASP O : -0.469: 0
: 2123:A 122 LEU 1HD1 :A 40 LEU 2HD2 : -0.687: 0
: 2123:A 40 LEU HG :A 36 GLU O : -0.641: 0
: 2123:A 39 LYS 2HZ :A 36 GLU HA : -0.555: 0
: 2123:A 36 GLU CD :A 39 LYS 1HZ : -0.517: 0
: 2123:A 36 GLU HA :A 39 LYS NZ : -0.511: 0
: 2123:A 43 ILE 1HG1 :A 39 LYS O : -0.424: 0
: 2123:A 76 ASN 2HB :A 79 ILE 2HG1 : -0.651: 0
: 2123:A 80 PRO 2HG :A 73 ARG 1HB : -0.612: 0
: 2123:A 79 ILE CG1 :A 76 ASN 2HB : -0.500: 0
: 2123:A 79 ILE HB :A 80 PRO CD : -0.452: 0
: 2123:A 79 ILE CD1 :A 76 ASN 2HB : -0.413: 0
: 2123:A 77 THR HA :A 73 ARG 1HG : -0.405: 0
: 2123:A 80 PRO 1HD :A 79 ILE N : -0.405: 0
: 2123:A 47 LEU 1HB :A 42 VAL 3HG1 : -0.603: 0
: 2123:A 42 VAL 3HG2 :A 38 ILE O : -0.587: 0
: 2123:A 38 ILE 1HG1 :A 34 ALA O : -0.512: 0
: 2123:A 72 LEU 2HD1 :A 69 SER HA : -0.491: 0
: 2123:A 72 LEU HG :A 68 ALA O : -0.481: 0
: 2123:A 60 TRP HH2 :A 42 VAL 1HG1 : -0.448: 0
: 2123:A 68 ALA 1HB :A 42 VAL CG2 : -0.425: 0
: 2123:A 65 LEU CD1 :A 35 GLU 2HG : -0.600: 0
: 2123:A 61 LYS HA :A 61 LYS 1HE : -0.574: 0
: 2123:A 61 LYS CA :A 61 LYS 1HE : -0.546: 0
: 2123:A 61 LYS O :A 65 LEU 2HD1 : -0.482: 0
: 2123:A 32 LYS HA :A 35 GLU 2HB : -0.469: 0
: 2123:A 61 LYS CE :A 61 LYS HA : -0.445: 0
: 2123:A 32 LYS O :A 35 GLU 2HB : -0.444: 0
: 2123:A 35 GLU 2HG :A 65 LEU 1HD1 : -0.428: 0
: 2123:A 10 TYR O :A 29 LYS 2HB : -0.597: 0
: 2123:A 25 GLN O :A 29 LYS 1HG : -0.584: 0
: 2123:A 29 LYS 2HE :A 29 LYS HA : -0.566: 0
: 2123:A 105 LYS 1HE :A 101 GLU 1HG : -0.596: 0
: 2123:A 105 LYS 1HG :A 101 GLU O : -0.516: 0
: 2123:A 54 VAL 3HG2 :A 50 ILE O : -0.588: 0
: 2123:A 50 ILE 1HD1 :A 67 LYS 1HD : -0.564: 0
: 2123:A 111 VAL 1HG1 :A 30 TYR 1HB : -0.560: 0
: 2123:A 30 TYR CE2 :A 27 CYS HA : -0.545: 0
: 2123:A 107 LEU HG :A 89 LEU 1HD1 : -0.541: 0
: 2123:A 107 LEU 2HD2 :A 106 LYS 1HE : -0.522: 0
: 2123:A 111 VAL 1HG2 :A 30 TYR HD1 : -0.507: 0
: 2123:A 111 VAL 1HG2 :A 30 TYR CD1 : -0.486: 0
: 2123:A 111 VAL O :A 115 VAL 3HG2 : -0.461: 0
: 2123:A 27 CYS HA :A 30 TYR CZ : -0.446: 0
: 2123:A 33 ALA 3HB :A 30 TYR HA : -0.438: 0
: 2123:A 89 LEU 3HD1 :A 27 CYS 2HB : -0.402: 0
: 2123:A 63 GLU CD :A 63 GLU H : -0.537: 0
: 2123:A 9 VAL O :A 13 GLU 2HG : -0.497: 0
: 2123:A 5 THR O :A 9 VAL 3HG2 : -0.400: 0
: 2123:A 4 SER N :A 3 ILE 2HG2 : -0.495: 0
: 2123:A 104 VAL 3HG2 :A 100 ASN O : -0.476: 0
: 2123:A 102 LYS 2HD :A 102 LYS C : -0.440: 0
: 2123:A 44 GLU 1HG :A 41 LEU HA : -0.429: 0
: 2123:A 45 ASN N :A 44 GLU 2HG : -0.402: 0
: 2123:A 12 GLU O :A 15 GLU 1HB : -0.404: 0
#sum2 ::26.85 clashscore : 26.85 clashscore B<40
#summary::2123 atoms:2123 atoms B<40:237314 potential dots:14830.0 A^2:57 bumps:57 bumps B<40:532.7 score
List of bad contacts calculated by MAGE for model $num_n
/farm/software/bin/probe
: 2123:A 104 VAL 1HG2 :A 23 LEU 2HD2 : -0.894: 0
: 2123:A 108 LYS 2HE :A 105 LYS HA : -0.866: 0
: 2123:A 105 LYS O :A 108 LYS 1HG : -0.420: 0
: 2123:A 107 LEU HG :A 89 LEU 1HD2 : -0.711: 0
: 2123:A 93 GLY 2HA :A 89 LEU O : -0.444: 0
: 2123:A 85 SER O :A 89 LEU HG : -0.422: 0
: 2123:A 94 PHE 2HB :A 93 GLY O : -0.418: 0
: 2123:A 24 VAL 3HG1 :A 94 PHE 1HB : -0.411: 0
: 2123:A 54 VAL 1HG1 :A 60 TRP HA : -0.668: 0
: 2123:A 54 VAL O :A 54 VAL 2HG1 : -0.434: 0
: 2123:A 126 HIS HA :A 123 GLU OE1 : -0.637: 0
: 2123:A 123 GLU CD :A 126 HIS HA : -0.540: 0
: 2123:A 77 THR HA :A 73 ARG 1HG : -0.619: 0
: 2123:A 80 PRO 1HD :A 79 ILE N : -0.446: 0
: 2123:A 79 ILE HB :A 80 PRO CD : -0.427: 0
: 2123:A 79 ILE 1HG1 :A 76 ASN 2HB : -0.420: 0
: 2123:A 78 GLU 1HB :A 76 ASN OD1 : -0.417: 0
: 2123:A 73 ARG HA :A 76 ASN O : -0.410: 0
: 2123:A 82 LEU HG :A 78 GLU O : -0.407: 0
: 2123:A 65 LEU 2HD1 :A 62 SER HA : -0.609: 0
: 2123:A 92 GLU 2HG :A 91 VAL 2HG1 : -0.518: 0
: 2123:A 91 VAL 3HG2 :A 87 TRP O : -0.467: 0
: 2123:A 62 SER 1HB :A 87 TRP CZ3 : -0.455: 0
: 2123:A 47 LEU 1HB :A 42 VAL 3HG1 : -0.592: 0
: 2123:A 42 VAL 3HG2 :A 38 ILE O : -0.562: 0
: 2123:A 50 ILE 1HG2 :A 68 ALA 2HB : -0.488: 0
: 2123:A 42 VAL HA :A 72 LEU 1HD2 : -0.461: 0
: 2123:A 72 LEU HG :A 68 ALA O : -0.420: 0
: 2123:A 42 VAL CG1 :A 47 LEU 1HB : -0.407: 0
: 2123:A 13 GLU CB :A 29 LYS 2HD : -0.565: 0
: 2123:A 29 LYS 2HD :A 13 GLU 1HB : -0.492: 0
: 2123:A 32 LYS 1HB :A 29 LYS HA : -0.432: 0
: 2123:A 9 VAL O :A 13 GLU 2HG : -0.412: 0
: 2123:A 17 PHE HE2 :A 29 LYS CE : -0.410: 0
: 2123:A 17 PHE HE2 :A 29 LYS 2HE : -0.408: 0
: 2123:A 55 LYS 2HD :A 55 LYS O : -0.560: 0
: 2123:A 52 ASN HA :A 55 LYS 1HB : -0.524: 0
: 2123:A 49 GLU HA :A 52 ASN ND2 : -0.460: 0
: 2123:A 55 LYS 2HD :A 55 LYS C : -0.431: 0
: 2123:A 120 ASN 1HB :A 116 ILE O : -0.547: 0
: 2123:A 112 ARG O :A 116 ILE 2HG1 : -0.422: 0
: 2123:A 44 GLU CG :A 41 LEU HA : -0.538: 0
: 2123:A 41 LEU HG :A 37 ALA O : -0.423: 0
: 2123:A 41 LEU O :A 44 GLU 2HG : -0.417: 0
: 2123:A 44 GLU 1HG :A 41 LEU HA : -0.411: 0
: 2123:A 48 LYS 1HB :A 48 LYS NZ : -0.508: 0
: 2123:A 46 ASN 1HB :A 45 ASN O : -0.504: 0
: 2123:A 111 VAL O :A 115 VAL 3HG2 : -0.493: 0
: 2123:A 30 TYR 2HB :A 111 VAL 1HG2 : -0.401: 0
: 2123:A 81 ILE O :A 84 LYS 2HB : -0.476: 0
: 2123:A 99 LEU H :A 97 LEU C : -0.468: 0
: 2123:A 67 LYS 1HD :A 67 LYS HA : -0.426: 0
: 2123:A 113 LYS O :A 117 PHE HD2 : -0.413: 0
: 2123:A 11 TYR O :A 15 GLU 2HG : -0.404: 0
#sum2 ::25.44 clashscore : 25.44 clashscore B<40
#summary::2123 atoms:2123 atoms B<40:237291 potential dots:14830.0 A^2:54 bumps:54 bumps B<40:514.2 score
List of bad contacts calculated by MAGE for model $num_n
/farm/software/bin/probe
: 2123:A 89 LEU 2HD2 :A 99 LEU 3HD1 : -0.894: 0
: 2123:A 107 LEU HG :A 89 LEU 1HD1 : -0.861: 0
: 2123:A 32 LYS HA :A 32 LYS 1HE : -0.883: 0
: 2123:A 32 LYS HA :A 32 LYS CE : -0.627: 0
: 2123:A 32 LYS O :A 36 GLU 1HG : -0.611: 0
: 2123:A 32 LYS 1HE :A 32 LYS CA : -0.589: 0
: 2123:A 36 GLU O :A 40 LEU 3HD1 : -0.558: 0
: 2123:A 39 LYS 3HZ :A 36 GLU HA : -0.490: 0
: 2123:A 5 THR 1HG2 :A 122 LEU 1HB : -0.443: 0
: 2123:A 40 LEU 3HD2 :A 122 LEU 3HD1 : -0.441: 0
: 2123:A 48 LYS HA :A 51 THR 2HG2 : -0.742: 0
: 2123:A 48 LYS HA :A 51 THR CG2 : -0.475: 0
: 2123:A 73 ARG O :A 73 ARG 1HD : -0.638: 0
: 2123:A 77 THR HA :A 73 ARG 1HG : -0.524: 0
: 2123:A 112 ARG O :A 116 ILE 1HG1 : -0.606: 0
: 2123:A 120 ASN 1HB :A 116 ILE O : -0.400: 0
: 2123:A 50 ILE 1HD1 :A 71 LEU 2HD1 : -0.593: 0
: 2123:A 64 ASN 2HB :A 50 ILE 3HG2 : -0.589: 0
: 2123:A 54 VAL 3HG2 :A 50 ILE O : -0.496: 0
: 2123:A 54 VAL CG1 :A 60 TRP HA : -0.484: 0
: 2123:A 71 LEU HG :A 67 LYS O : -0.463: 0
: 2123:A 50 ILE 2HD1 :A 49 GLU 1HB : -0.437: 0
: 2123:A 54 VAL 1HG1 :A 60 TRP HA : -0.435: 0
: 2123:A 30 TYR HE2 :A 104 VAL 3HG1 : -0.586: 0
: 2123:A 27 CYS HA :A 30 TYR CD2 : -0.471: 0
: 2123:A 100 ASN 1HB :A 103 GLU OE1 : -0.446: 0
: 2123:A 23 LEU 2HD2 :A 104 VAL 1HG1 : -0.428: 0
: 2123:A 14 ALA 1HB :A 30 TYR CE1 : -0.416: 0
: 2123:A 104 VAL 3HG2 :A 100 ASN O : -0.413: 0
: 2123:A 82 LEU HG :A 78 GLU O : -0.583: 0
: 2123:A 47 LEU 1HB :A 42 VAL 3HG1 : -0.582: 0
: 2123:A 42 VAL 3HG2 :A 38 ILE O : -0.580: 0
: 2123:A 38 ILE 1HG1 :A 34 ALA O : -0.519: 0
: 2123:A 7 ALA 1HB :A 115 VAL 3HG1 : -0.568: 0
: 2123:A 111 VAL O :A 115 VAL 3HG2 : -0.426: 0
: 2123:A 65 LEU 2HD1 :A 62 SER HA : -0.547: 0
: 2123:A 123 GLU OE2 :A 119 VAL 3HG1 : -0.496: 0
: 2123:A 79 ILE HB :A 80 PRO CD : -0.488: 0
: 2123:A 79 ILE 1HG2 :A 72 LEU 3HD1 : -0.443: 0
: 2123:A 80 PRO 1HD :A 79 ILE N : -0.441: 0
: 2123:A 72 LEU HG :A 68 ALA O : -0.440: 0
: 2123:A 79 ILE HB :A 80 PRO 2HD : -0.416: 0
: 2123:A 52 ASN O :A 55 LYS 1HB : -0.483: 0
: 2123:A 18 LEU 2HD2 :A 108 LYS 2HD : -0.454: 0
: 2123:A 37 ALA O :A 41 LEU 3HD1 : -0.451: 0
: 2123:A 114 LEU HG :A 37 ALA CB : -0.443: 0
: 2123:A 37 ALA HA :A 118 ALA CB : -0.408: 0
: 2123:A 105 LYS 1HG :A 101 GLU O : -0.447: 0
: 2123:A 20 LYS 1HD :A 20 LYS HA : -0.434: 0
: 2123:A 87 TRP 2HB :A 84 LYS HA : -0.431: 0
: 2123:A 91 VAL 3HG2 :A 87 TRP O : -0.428: 0
: 2123:A 128 HIS O :A 129 HIS 2HB : -0.405: 0
#sum2 ::24.49 clashscore : 24.49 clashscore B<40
#summary::2123 atoms:2123 atoms B<40:237214 potential dots:14830.0 A^2:52 bumps:52 bumps B<40:560.8 score
List of bad contacts calculated by MAGE for model $num_n
/farm/software/bin/probe
: 2123:A 82 LEU 2HB :A 114 LEU 1HD1 : -0.846: 0
: 2123:A 82 LEU 2HB :A 114 LEU CD1 : -0.512: 0
: 2123:A 114 LEU 1HD1 :A 82 LEU CB : -0.512: 0
: 2123:A 82 LEU HG :A 78 GLU O : -0.493: 0
: 2123:A 107 LEU HG :A 89 LEU 1HD1 : -0.783: 0
: 2123:A 89 LEU 2HB :A 27 CYS 2HB : -0.614: 0
: 2123:A 107 LEU CG :A 89 LEU 1HD1 : -0.582: 0
: 2123:A 107 LEU HG :A 89 LEU 1HD2 : -0.524: 0
: 2123:A 27 CYS SG :A 104 VAL 2HG2 : -0.517: 0
: 2123:A 30 TYR 2HB :A 111 VAL 1HG2 : -0.505: 0
: 2123:A 85 SER C :A 107 LEU 1HD1 : -0.482: 0
: 2123:A 104 VAL 3HG2 :A 100 ASN O : -0.443: 0
: 2123:A 107 LEU HG :A 89 LEU CD1 : -0.427: 0
: 2123:A 107 LEU 3HD1 :A 107 LEU HA : -0.421: 0
: 2123:A 111 VAL O :A 115 VAL 3HG2 : -0.409: 0
: 2123:A 119 VAL 3HG2 :A 115 VAL O : -0.403: 0
: 2123:A 30 TYR HD2 :A 27 CYS HA : -0.400: 0
: 2123:A 40 LEU 2HD1 :A 118 ALA 1HB : -0.686: 0
: 2123:A 40 LEU CD1 :A 118 ALA 1HB : -0.498: 0
: 2123:A 47 LEU 1HB :A 42 VAL 3HG1 : -0.643: 0
: 2123:A 36 GLU HA :A 39 LYS 2HD : -0.577: 0
: 2123:A 42 VAL 3HG2 :A 38 ILE O : -0.569: 0
: 2123:A 54 VAL 3HG2 :A 50 ILE O : -0.521: 0
: 2123:A 50 ILE 3HD1 :A 47 LEU 2HB : -0.519: 0
: 2123:A 79 ILE 1HG2 :A 72 LEU 2HD1 : -0.514: 0
: 2123:A 80 PRO 1HD :A 79 ILE N : -0.500: 0
: 2123:A 60 TRP CE2 :A 39 LYS 1HG : -0.466: 0
: 2123:A 60 TRP N :A 59 ARG 1HG : -0.456: 0
: 2123:A 60 TRP HH2 :A 42 VAL 1HG1 : -0.449: 0
: 2123:A 38 ILE 1HG1 :A 34 ALA O : -0.444: 0
: 2123:A 42 VAL HA :A 72 LEU 1HD2 : -0.442: 0
: 2123:A 50 ILE N :A 50 ILE 2HD1 : -0.437: 0
: 2123:A 36 GLU N :A 35 GLU 2HG : -0.434: 0
: 2123:A 72 LEU HG :A 47 LEU 1HD1 : -0.429: 0
: 2123:A 50 ILE CD1 :A 71 LEU 2HD1 : -0.423: 0
: 2123:A 50 ILE 1HD1 :A 71 LEU 2HD1 : -0.421: 0
: 2123:A 60 TRP H :A 59 ARG 1HG : -0.420: 0
: 2123:A 43 ILE 1HG1 :A 39 LYS O : -0.419: 0
: 2123:A 83 TRP HE1 :A 66 PHE HD1 : -0.629: 0
: 2123:A 66 PHE HD2 :A 63 GLU HA : -0.433: 0
: 2123:A 16 GLU 1HG :A 12 GLU O : -0.603: 0
: 2123:A 13 GLU O :A 16 GLU 1HB : -0.507: 0
: 2123:A 45 ASN OD1 :A 75 ASN 1HB : -0.587: 0
: 2123:A 41 LEU 1HD1 :A 117 PHE 2HB : -0.565: 0
: 2123:A 117 PHE HZ :A 75 ASN 1HD2 : -0.463: 0
: 2123:A 112 ARG O :A 116 ILE 1HG1 : -0.586: 0
: 2123:A 120 ASN 1HB :A 116 ILE O : -0.479: 0
: 2123:A 99 LEU 1HD2 :A 92 GLU HA : -0.534: 0
: 2123:A 99 LEU N :A 99 LEU 2HD2 : -0.419: 0
: 2123:A 109 GLU 1HG :A 106 LYS O : -0.533: 0
: 2123:A 84 LYS 2HD :A 84 LYS C : -0.514: 0
: 2123:A 84 LYS O :A 84 LYS 2HD : -0.498: 0
: 2123:A 5 THR O :A 8 GLU 1HG : -0.490: 0
: 2123:A 9 VAL 3HG2 :A 5 THR O : -0.477: 0
: 2123:A 91 VAL 3HG2 :A 87 TRP O : -0.487: 0
: 2123:A 57 LYS 2HE :A 57 LYS 2HB : -0.443: 0
: 2123:A 48 LYS 2HE :A 48 LYS 1HB : -0.419: 0
: 2123:A 97 LEU N :A 97 LEU 2HD2 : -0.410: 0
: 2123:A 125 HIS H :A 123 GLU C : -0.409: 0
: 2123:A 61 LYS 1HE :A 61 LYS 2HB : -0.406: 0
: 2123:A 67 LYS 2HE :A 67 LYS 1HB : -0.405: 0
: 2123:A 70 LYS HA :A 70 LYS 1HD : -0.403: 0
#sum2 ::29.20 clashscore : 29.20 clashscore B<40
#summary::2123 atoms:2123 atoms B<40:237202 potential dots:14830.0 A^2:62 bumps:62 bumps B<40:583.3 score
List of bad contacts calculated by MAGE for model $num_n
/farm/software/bin/probe
: 2123:A 17 PHE HE2 :A 29 LYS 2HE : -0.840: 0
: 2123:A 6 SER H :A 5 THR 2HG2 : -0.571: 0
: 2123:A 10 TYR CE2 :A 6 SER 1HB : -0.495: 0
: 2123:A 10 TYR 2HB :A 29 LYS O : -0.495: 0
: 2123:A 6 SER N :A 5 THR 2HG2 : -0.466: 0
: 2123:A 10 TYR CB :A 33 ALA 2HB : -0.454: 0
: 2123:A 33 ALA 2HB :A 10 TYR 1HB : -0.429: 0
: 2123:A 107 LEU HG :A 89 LEU 1HD1 : -0.813: 0
: 2123:A 107 LEU HG :A 89 LEU CD1 : -0.607: 0
: 2123:A 89 LEU 2HD2 :A 99 LEU 2HD2 : -0.458: 0
: 2123:A 89 LEU 3HD1 :A 99 LEU 3HD1 : -0.434: 0
: 2123:A 61 LYS 2HB :A 57 LYS 1HD : -0.806: 0
: 2123:A 63 GLU HA :A 66 PHE CE2 : -0.688: 0
: 2123:A 63 GLU O :A 67 LYS 1HB : -0.506: 0
: 2123:A 61 LYS 2HE :A 63 GLU 1HB : -0.455: 0
: 2123:A 63 GLU HA :A 66 PHE CZ : -0.408: 0
: 2123:A 82 LEU HG :A 78 GLU O : -0.653: 0
: 2123:A 47 LEU 1HB :A 42 VAL 3HG1 : -0.648: 0
: 2123:A 42 VAL 3HG2 :A 38 ILE O : -0.493: 0
: 2123:A 38 ILE 1HG1 :A 34 ALA O : -0.484: 0
: 2123:A 79 ILE 1HG1 :A 76 ASN 2HB : -0.644: 0
: 2123:A 79 ILE HB :A 80 PRO 2HD : -0.594: 0
: 2123:A 79 ILE 1HG2 :A 72 LEU 1HB : -0.459: 0
: 2123:A 79 ILE 1HG2 :A 72 LEU CB : -0.441: 0
: 2123:A 76 ASN 2HB :A 79 ILE CG1 : -0.435: 0
: 2123:A 4 SER 2HB :A 8 GLU 2HG : -0.631: 0
: 2123:A 32 LYS HA :A 35 GLU 1HG : -0.617: 0
: 2123:A 27 CYS HA :A 30 TYR CZ : -0.587: 0
: 2123:A 30 TYR HD2 :A 14 ALA 1HB : -0.419: 0
: 2123:A 77 THR HA :A 73 ARG 2HG : -0.538: 0
: 2123:A 73 ARG 1HH1 :A 73 ARG 2HD : -0.408: 0
: 2123:A 60 TRP N :A 59 ARG 2HD : -0.535: 0
: 2123:A 105 LYS 1HE :A 101 GLU 1HG : -0.535: 0
: 2123:A 105 LYS 1HG :A 101 GLU O : -0.514: 0
: 2123:A 7 ALA 1HB :A 115 VAL 2HG2 : -0.517: 0
: 2123:A 115 VAL O :A 119 VAL 3HG2 : -0.495: 0
: 2123:A 111 VAL O :A 115 VAL 3HG2 : -0.401: 0
: 2123:A 43 ILE 1HG1 :A 39 LYS O : -0.500: 0
: 2123:A 39 LYS 2HB :A 36 GLU O : -0.435: 0
: 2123:A 104 VAL 3HG2 :A 100 ASN O : -0.485: 0
: 2123:A 100 ASN 1HB :A 103 GLU 2HG : -0.400: 0
: 2123:A 56 ASN C :A 56 ASN 2HD2 : -0.461: 0
: 2123:A 15 GLU 1HG :A 11 TYR O : -0.441: 0
: 2123:A 112 ARG O :A 116 ILE 2HG1 : -0.421: 0
#sum2 ::20.73 clashscore : 20.73 clashscore B<40
#summary::2123 atoms:2123 atoms B<40:237256 potential dots:14830.0 A^2:44 bumps:44 bumps B<40:603.5 score
Output from PDB validation software
Summary from PDB validation
May. 11, 00:42:13 2013
[ Text modified to reflect that this was run under PSVS - Aneerban Bhattacharya: Dec 2005 ]
The following checks were made on :
-----------------------------------------
CLOSE CONTACTS
==> Distances smaller than 2.2 Angstroms are considered as close contacts
for heavy atoms, 1.6 Angstroms for hydrogens.
Chain Atom Res Seq Chain Atom Res Seq Mol_ID Distance
-------------------------------------------------------------------------
A 2HG GLU 44 - A 2HB SER 121 10 Dist = 1.31
A HE2 PHE 17 - A 2HE LYS 29 20 Dist = 1.34
A HD1 PHE 66 - A HE1 TRP 83 19 Dist = 1.37
A 2HD2 LEU 89 - A HG LEU 107 12 Dist = 1.38
A 2HE LYS 57 - A 1HD LYS 61 11 Dist = 1.42
A HA LYS 32 - A 1HG GLU 35 10 Dist = 1.42
A HA ASP 110 - A 2HD LYS 113 13 Dist = 1.44
A HE2 TYR 11 - A 1HH1 ARG 112 11 Dist = 1.45
A HA LYS 32 - A 1HG GLU 35 4 Dist = 1.45
A 1HD LYS 61 - A 1HG GLU 63 15 Dist = 1.45
A 1HD2 LEU 23 - A 2HG2 VAL 104 17 Dist = 1.45
A HA LYS 32 - A 1HG GLU 35 8 Dist = 1.46
A 3HD2 LEU 89 - A 3HD1 LEU 99 18 Dist = 1.46
A 2HB HIS 126 - A HD2 HIS 128 5 Dist = 1.47
A HA LYS 105 - A 2HE LYS 108 17 Dist = 1.47
A HE2 PHE 17 - A 1HZ LYS 29 10 Dist = 1.48
A 2HD1 LEU 89 - A HG LEU 107 18 Dist = 1.50
A 3HD1 LEU 89 - A HG LEU 107 11 Dist = 1.50
A 2HB LEU 82 - A 1HD1 LEU 114 19 Dist = 1.51
A HE2 TYR 30 - A 1HD1 LEU 89 10 Dist = 1.52
A 2HD1 LEU 89 - A HG LEU 107 20 Dist = 1.52
A HA ASP 110 - A 2HD LYS 113 10 Dist = 1.53
A 1HG GLU 96 - A 2HD2 LEU 99 12 Dist = 1.53
A 1HD LYS 57 - A 2HB LYS 61 20 Dist = 1.53
A 2HD2 LEU 23 - A 1HG2 VAL 104 4 Dist = 1.53
A 2HG1 ILE 3 - A 2HD1 LEU 122 12 Dist = 1.53
A HD2 TYR 10 - A 2HG LYS 29 1 Dist = 1.54
A 3HG2 VAL 42 - A 2HD1 LEU 47 4 Dist = 1.54
A 1HD1 LEU 89 - A HG LEU 107 19 Dist = 1.54
A 2HB ASN 76 - A 2HG1 ILE 79 2 Dist = 1.56
A OE1 GLU 15 - A 1HZ LYS 108 5 Dist = 1.56
A 2HG GLU 109 - A 1HH2 ARG 112 3 Dist = 1.56
A 2HD1 LEU 89 - A HG LEU 107 15 Dist = 1.56
A 1HE LYS 61 - A 2HB GLU 63 5 Dist = 1.56
A HA LYS 32 - A 1HG GLU 35 13 Dist = 1.56
A 1HB ARG 73 - A 2HG PRO 80 11 Dist = 1.57
A 2HD1 LEU 89 - A HG LEU 107 7 Dist = 1.57
A HA GLU 36 - A 2HE LYS 39 10 Dist = 1.57
A 1HD2 ASN 75 - A HZ PHE 117 19 Dist = 1.58
A 1HD1 LEU 89 - A HG LEU 107 2 Dist = 1.58
A 1HA GLY 93 - A 1HB LEU 97 2 Dist = 1.58
A OE2 GLU 49 - A 2HZ LYS 67 2 Dist = 1.58
A OE2 GLU 103 - A 2HZ LYS 106 19 Dist = 1.58
A HE2 PHE 17 - A 2HE LYS 29 8 Dist = 1.58
A OE1 GLU 36 - A 2HZ LYS 39 6 Dist = 1.59
A 2HD1 LEU 89 - A HG LEU 107 1 Dist = 1.59
A 2HZ LYS 106 - A OD1 ASP 110 15 Dist = 1.59
A OE1 GLU 13 - A 1HZ LYS 29 7 Dist = 1.59
A OE1 GLU 15 - A 3HZ LYS 108 16 Dist = 1.60
A OE2 GLU 36 - A 3HZ LYS 39 9 Dist = 1.60
A 3HZ LYS 106 - A OD2 ASP 110 6 Dist = 1.60
A OE1 GLU 13 - A 3HZ LYS 29 10 Dist = 1.60
A OE1 GLU 13 - A 3HZ LYS 29 17 Dist = 1.60
A OE1 GLU 44 - A HD1 HIS 124 9 Dist = 1.60
A OE2 GLU 28 - A 1HZ LYS 32 19 Dist = 1.60
DISTANCES AND ANGLES
We have checked your intra and intermolecular distances and angles with the
procedures currently in place at PDB:
==> Bond and angle checks are performed by first computing the average rms
error for all bonds and angles relative to standard values for nucleotide
units [L. Clowney et al., Geometric Parameters in Nucleic Acids: Nitrogenous
Bases, J.Am.Chem.Soc. 1996, 118, 509-518; A. Gelbin et al., Geometric
Parameters in Nucleic Acids: Sugar and Phosphate Constituents, J.Am.Chem.Soc.
1996, 118, 519-529] and amino acid units [R.A. Engh and R. Huber, Accurate
Bond and Angle Parameters for X-ray protein structure refinement, Acta
Crystallogr. 1991, A47, 392-400]. Any bond or angle which deviates from the
dictionary values by more than six times this computed rms error is
identified as an outlier.
*** Covalent Bond Lengths:
The RMS deviation for covalent bonds relative to the standard
dictionary is 0.004 Angstroms
The following table contains a list of the covalent bonds
greater than 6.0*RMSD.
Deviation Residue Chain Sequence Model AT1 - AT2 Bond Dictionary
Name ID Number Distance Value
------------------------------------------------------------------------
-0.023 GLU A 16 6 N - CA 1.435 1.458
-0.025 GLU A 16 6 CB - CG 1.495 1.520
-0.025 LYS A 57 14 N - CA 1.433 1.458
*** Covalent Angle Values:
The RMS deviation for covalent angles relative to the standard
dictionary is 0.6 degrees.
The following table contains a list of the covalent bond angles
greater than 6.0*RMSD.
Deviation Residue Chain Sequence Model AT1 - AT2 - AT3 Bond Dictionary
Name ID Number Angle Value
--------------------------------------------------------------------------------
-3.7 HIS A 126 1 N - CA - C 107.5 111.2
3.6 THR A 5 2 C - N - CA 125.3 121.7
-3.7 ASN A 45 4 N - CA - CB 106.8 110.5
-3.8 LYS A 102 6 N - CA - CB 106.7 110.5
-3.5 LYS A 102 11 N - CA - CB 107.0 110.5
3.6 GLU A 92 12 N - CA - C 114.8 111.2
-4.0 GLY A 93 17 N - CA - C 108.5 112.5
-3.8 GLU A 96 19 N - CA - C 107.4 111.2
TORSION ANGLES
The torsion angle distributions have been checked. The postscript file of the
conformation rings showing the torsion angle distributions will be sent in a
separate E-mail message.
CHIRALITY
The chirality has been checked. O1P, O2P, and hydrogen atoms which do not
follow the convention defined in the IUBMB (Liebecq, C. Compendium of
Biochemical Nomenclature and Related Documents, 2nd ed.; Portland Press:
London and Chapel Hill, 1992) and IUPAC nomenclature (J.L. Markley, A. Bax,
Y. Arata, C.W. Hilbers, R. Kaptein, B.D. Sykes, P.E. Wright and K. W�thrich,
Recommendations for the Presentation of NMR Structures of Proteins and
Nucleic Acids, Pure & Appl. Chem., Vol. 70, pp. 117-142, 1998) have been
standardized. Any other stereochemical violations are listed below.
E/Z NOMENCLATURE
E/Z nomenclature of hydrogens and/or nitrogens on Arg, Asn or Gln residues needs
to be corrected to conform with the standard for E/Z orientation presented in
[J.L. Markley, et al., Recommendations for the Presentation of NMR Structures of
Proteins and Nucleic Acids, Pure & Appl. Chem., 1998, 70, 117-142].
Model Chain Residue Residue Atom Name Original
Name Number Atom Name
----- ----- ------- ------- -------- ---------
1 A GLN 25 1HE2
1 A GLN 25 2HE2
1 A ASN 45 1HD2
1 A ASN 45 2HD2
1 A ASN 46 1HD2
1 A ASN 46 2HD2
1 A ASN 52 1HD2
1 A ASN 52 2HD2
1 A ASN 53 1HD2
1 A ASN 53 2HD2
1 A ASN 56 1HD2
1 A ASN 56 2HD2
1 A ASN 64 1HD2
1 A ASN 64 2HD2
1 A ASN 75 1HD2
1 A ASN 75 2HD2
1 A ASN 76 1HD2
1 A ASN 76 2HD2
1 A ASN 100 1HD2
1 A ASN 100 2HD2
1 A ASN 120 1HD2
1 A ASN 120 2HD2
2 A GLN 25 1HE2
2 A GLN 25 2HE2
2 A ASN 45 1HD2
2 A ASN 45 2HD2
2 A ASN 46 1HD2
2 A ASN 46 2HD2
2 A ASN 52 1HD2
2 A ASN 52 2HD2
2 A ASN 53 1HD2
2 A ASN 53 2HD2
2 A ASN 56 1HD2
2 A ASN 56 2HD2
2 A ASN 64 1HD2
2 A ASN 64 2HD2
2 A ASN 75 1HD2
2 A ASN 75 2HD2
2 A ASN 76 1HD2
2 A ASN 76 2HD2
2 A ASN 100 1HD2
2 A ASN 100 2HD2
2 A ASN 120 1HD2
2 A ASN 120 2HD2
3 A GLN 25 1HE2
3 A GLN 25 2HE2
3 A ASN 45 1HD2
3 A ASN 45 2HD2
3 A ASN 46 1HD2
3 A ASN 46 2HD2
3 A ASN 52 1HD2
3 A ASN 52 2HD2
3 A ASN 53 1HD2
3 A ASN 53 2HD2
3 A ASN 56 1HD2
3 A ASN 56 2HD2
3 A ASN 64 1HD2
3 A ASN 64 2HD2
3 A ASN 75 1HD2
3 A ASN 75 2HD2
3 A ASN 76 1HD2
3 A ASN 76 2HD2
3 A ASN 100 1HD2
3 A ASN 100 2HD2
3 A ASN 120 1HD2
3 A ASN 120 2HD2
4 A GLN 25 1HE2
4 A GLN 25 2HE2
4 A ASN 45 1HD2
4 A ASN 45 2HD2
4 A ASN 46 1HD2
4 A ASN 46 2HD2
4 A ASN 52 1HD2
4 A ASN 52 2HD2
4 A ASN 53 1HD2
4 A ASN 53 2HD2
4 A ASN 56 1HD2
4 A ASN 56 2HD2
4 A ASN 64 1HD2
4 A ASN 64 2HD2
4 A ASN 75 1HD2
4 A ASN 75 2HD2
4 A ASN 76 1HD2
4 A ASN 76 2HD2
4 A ASN 100 1HD2
4 A ASN 100 2HD2
4 A ASN 120 1HD2
4 A ASN 120 2HD2
5 A GLN 25 1HE2
5 A GLN 25 2HE2
5 A ASN 45 1HD2
5 A ASN 45 2HD2
5 A ASN 46 1HD2
5 A ASN 46 2HD2
5 A ASN 52 1HD2
5 A ASN 52 2HD2
5 A ASN 53 1HD2
5 A ASN 53 2HD2
5 A ASN 56 1HD2
5 A ASN 56 2HD2
5 A ASN 64 1HD2
5 A ASN 64 2HD2
5 A ASN 75 1HD2
5 A ASN 75 2HD2
5 A ASN 76 1HD2
5 A ASN 76 2HD2
5 A ASN 100 1HD2
5 A ASN 100 2HD2
5 A ASN 120 1HD2
5 A ASN 120 2HD2
6 A GLN 25 1HE2
6 A GLN 25 2HE2
6 A ASN 45 1HD2
6 A ASN 45 2HD2
6 A ASN 46 1HD2
6 A ASN 46 2HD2
6 A ASN 52 1HD2
6 A ASN 52 2HD2
6 A ASN 53 1HD2
6 A ASN 53 2HD2
6 A ASN 56 1HD2
6 A ASN 56 2HD2
6 A ASN 64 1HD2
6 A ASN 64 2HD2
6 A ASN 75 1HD2
6 A ASN 75 2HD2
6 A ASN 76 1HD2
6 A ASN 76 2HD2
6 A ASN 100 1HD2
6 A ASN 100 2HD2
6 A ASN 120 1HD2
6 A ASN 120 2HD2
7 A GLN 25 1HE2
7 A GLN 25 2HE2
7 A ASN 45 1HD2
7 A ASN 45 2HD2
7 A ASN 46 1HD2
7 A ASN 46 2HD2
7 A ASN 52 1HD2
7 A ASN 52 2HD2
7 A ASN 53 1HD2
7 A ASN 53 2HD2
7 A ASN 56 1HD2
7 A ASN 56 2HD2
7 A ASN 64 1HD2
7 A ASN 64 2HD2
7 A ASN 75 1HD2
7 A ASN 75 2HD2
7 A ASN 76 1HD2
7 A ASN 76 2HD2
7 A ASN 100 1HD2
7 A ASN 100 2HD2
7 A ASN 120 1HD2
7 A ASN 120 2HD2
8 A GLN 25 1HE2
8 A GLN 25 2HE2
8 A ASN 45 1HD2
8 A ASN 45 2HD2
8 A ASN 46 1HD2
8 A ASN 46 2HD2
8 A ASN 52 1HD2
8 A ASN 52 2HD2
8 A ASN 53 1HD2
8 A ASN 53 2HD2
8 A ASN 56 1HD2
8 A ASN 56 2HD2
8 A ASN 64 1HD2
8 A ASN 64 2HD2
8 A ASN 75 1HD2
8 A ASN 75 2HD2
8 A ASN 76 1HD2
8 A ASN 76 2HD2
8 A ASN 100 1HD2
8 A ASN 100 2HD2
8 A ASN 120 1HD2
8 A ASN 120 2HD2
9 A GLN 25 1HE2
9 A GLN 25 2HE2
9 A ASN 45 1HD2
9 A ASN 45 2HD2
9 A ASN 46 1HD2
9 A ASN 46 2HD2
9 A ASN 52 1HD2
9 A ASN 52 2HD2
9 A ASN 53 1HD2
9 A ASN 53 2HD2
9 A ASN 56 1HD2
9 A ASN 56 2HD2
9 A ASN 64 1HD2
9 A ASN 64 2HD2
9 A ASN 75 1HD2
9 A ASN 75 2HD2
9 A ASN 76 1HD2
9 A ASN 76 2HD2
9 A ASN 100 1HD2
9 A ASN 100 2HD2
9 A ASN 120 1HD2
9 A ASN 120 2HD2
10 A GLN 25 1HE2
10 A GLN 25 2HE2
10 A ASN 45 1HD2
10 A ASN 45 2HD2
10 A ASN 46 1HD2
10 A ASN 46 2HD2
10 A ASN 52 1HD2
10 A ASN 52 2HD2
10 A ASN 53 1HD2
10 A ASN 53 2HD2
10 A ASN 56 1HD2
10 A ASN 56 2HD2
10 A ASN 64 1HD2
10 A ASN 64 2HD2
10 A ASN 75 1HD2
10 A ASN 75 2HD2
10 A ASN 76 1HD2
10 A ASN 76 2HD2
10 A ASN 100 1HD2
10 A ASN 100 2HD2
10 A ASN 120 1HD2
10 A ASN 120 2HD2
11 A GLN 25 1HE2
11 A GLN 25 2HE2
11 A ASN 45 1HD2
11 A ASN 45 2HD2
11 A ASN 46 1HD2
11 A ASN 46 2HD2
11 A ASN 52 1HD2
11 A ASN 52 2HD2
11 A ASN 53 1HD2
11 A ASN 53 2HD2
11 A ASN 56 1HD2
11 A ASN 56 2HD2
11 A ASN 64 1HD2
11 A ASN 64 2HD2
11 A ASN 75 1HD2
11 A ASN 75 2HD2
11 A ASN 76 1HD2
11 A ASN 76 2HD2
11 A ASN 100 1HD2
11 A ASN 100 2HD2
11 A ASN 120 1HD2
11 A ASN 120 2HD2
12 A GLN 25 1HE2
12 A GLN 25 2HE2
12 A ASN 45 1HD2
12 A ASN 45 2HD2
12 A ASN 46 1HD2
12 A ASN 46 2HD2
12 A ASN 52 1HD2
12 A ASN 52 2HD2
12 A ASN 53 1HD2
12 A ASN 53 2HD2
12 A ASN 56 1HD2
12 A ASN 56 2HD2
12 A ASN 64 1HD2
12 A ASN 64 2HD2
12 A ASN 75 1HD2
12 A ASN 75 2HD2
12 A ASN 76 1HD2
12 A ASN 76 2HD2
12 A ASN 100 1HD2
12 A ASN 100 2HD2
12 A ASN 120 1HD2
12 A ASN 120 2HD2
13 A GLN 25 1HE2
13 A GLN 25 2HE2
13 A ASN 45 1HD2
13 A ASN 45 2HD2
13 A ASN 46 1HD2
13 A ASN 46 2HD2
13 A ASN 52 1HD2
13 A ASN 52 2HD2
13 A ASN 53 1HD2
13 A ASN 53 2HD2
13 A ASN 56 1HD2
13 A ASN 56 2HD2
13 A ASN 64 1HD2
13 A ASN 64 2HD2
13 A ASN 75 1HD2
13 A ASN 75 2HD2
13 A ASN 76 1HD2
13 A ASN 76 2HD2
13 A ASN 100 1HD2
13 A ASN 100 2HD2
13 A ASN 120 1HD2
13 A ASN 120 2HD2
14 A GLN 25 1HE2
14 A GLN 25 2HE2
14 A ASN 45 1HD2
14 A ASN 45 2HD2
14 A ASN 46 1HD2
14 A ASN 46 2HD2
14 A ASN 52 1HD2
14 A ASN 52 2HD2
14 A ASN 53 1HD2
14 A ASN 53 2HD2
14 A ASN 56 1HD2
14 A ASN 56 2HD2
14 A ASN 64 1HD2
14 A ASN 64 2HD2
14 A ASN 75 1HD2
14 A ASN 75 2HD2
14 A ASN 76 1HD2
14 A ASN 76 2HD2
14 A ASN 100 1HD2
14 A ASN 100 2HD2
14 A ASN 120 1HD2
14 A ASN 120 2HD2
15 A GLN 25 1HE2
15 A GLN 25 2HE2
15 A ASN 45 1HD2
15 A ASN 45 2HD2
15 A ASN 46 1HD2
15 A ASN 46 2HD2
15 A ASN 52 1HD2
15 A ASN 52 2HD2
15 A ASN 53 1HD2
15 A ASN 53 2HD2
15 A ASN 56 1HD2
15 A ASN 56 2HD2
15 A ASN 64 1HD2
15 A ASN 64 2HD2
15 A ASN 75 1HD2
15 A ASN 75 2HD2
15 A ASN 76 1HD2
15 A ASN 76 2HD2
15 A ASN 100 1HD2
15 A ASN 100 2HD2
15 A ASN 120 1HD2
15 A ASN 120 2HD2
16 A GLN 25 1HE2
16 A GLN 25 2HE2
16 A ASN 45 1HD2
16 A ASN 45 2HD2
16 A ASN 46 1HD2
16 A ASN 46 2HD2
16 A ASN 52 1HD2
16 A ASN 52 2HD2
16 A ASN 53 1HD2
16 A ASN 53 2HD2
16 A ASN 56 1HD2
16 A ASN 56 2HD2
16 A ASN 64 1HD2
16 A ASN 64 2HD2
16 A ASN 75 1HD2
16 A ASN 75 2HD2
16 A ASN 76 1HD2
16 A ASN 76 2HD2
16 A ASN 100 1HD2
16 A ASN 100 2HD2
16 A ASN 120 1HD2
16 A ASN 120 2HD2
17 A GLN 25 1HE2
17 A GLN 25 2HE2
17 A ASN 45 1HD2
17 A ASN 45 2HD2
17 A ASN 46 1HD2
17 A ASN 46 2HD2
17 A ASN 52 1HD2
17 A ASN 52 2HD2
17 A ASN 53 1HD2
17 A ASN 53 2HD2
17 A ASN 56 1HD2
17 A ASN 56 2HD2
17 A ASN 64 1HD2
17 A ASN 64 2HD2
17 A ASN 75 1HD2
17 A ASN 75 2HD2
17 A ASN 76 1HD2
17 A ASN 76 2HD2
17 A ASN 100 1HD2
17 A ASN 100 2HD2
17 A ASN 120 1HD2
17 A ASN 120 2HD2
18 A GLN 25 1HE2
18 A GLN 25 2HE2
18 A ASN 45 1HD2
18 A ASN 45 2HD2
18 A ASN 46 1HD2
18 A ASN 46 2HD2
18 A ASN 52 1HD2
18 A ASN 52 2HD2
18 A ASN 53 1HD2
18 A ASN 53 2HD2
18 A ASN 56 1HD2
18 A ASN 56 2HD2
18 A ASN 64 1HD2
18 A ASN 64 2HD2
18 A ASN 75 1HD2
18 A ASN 75 2HD2
18 A ASN 76 1HD2
18 A ASN 76 2HD2
18 A ASN 100 1HD2
18 A ASN 100 2HD2
18 A ASN 120 1HD2
18 A ASN 120 2HD2
19 A GLN 25 1HE2
19 A GLN 25 2HE2
19 A ASN 45 1HD2
19 A ASN 45 2HD2
19 A ASN 46 1HD2
19 A ASN 46 2HD2
19 A ASN 52 1HD2
19 A ASN 52 2HD2
19 A ASN 53 1HD2
19 A ASN 53 2HD2
19 A ASN 56 1HD2
19 A ASN 56 2HD2
19 A ASN 64 1HD2
19 A ASN 64 2HD2
19 A ASN 75 1HD2
19 A ASN 75 2HD2
19 A ASN 76 1HD2
19 A ASN 76 2HD2
19 A ASN 100 1HD2
19 A ASN 100 2HD2
19 A ASN 120 1HD2
19 A ASN 120 2HD2
20 A GLN 25 1HE2
20 A GLN 25 2HE2
20 A ASN 45 1HD2
20 A ASN 45 2HD2
20 A ASN 46 1HD2
20 A ASN 46 2HD2
20 A ASN 52 1HD2
20 A ASN 52 2HD2
20 A ASN 53 1HD2
20 A ASN 53 2HD2
20 A ASN 56 1HD2
20 A ASN 56 2HD2
20 A ASN 64 1HD2
20 A ASN 64 2HD2
20 A ASN 75 1HD2
20 A ASN 75 2HD2
20 A ASN 76 1HD2
20 A ASN 76 2HD2
20 A ASN 100 1HD2
20 A ASN 100 2HD2
20 A ASN 120 1HD2
20 A ASN 120 2HD2
OTHER IMPORTANT ISSUES
==> The following residues are missing:
(Note: The SEQ number starts from 1 for each chain according to SEQRES
sequence record.)
RES MOD#C SEQ
MET( 1 A-128 )
SER( 1 A-127 )
ILE( 1 A-126 )
SER( 1 A-125 )
THR( 1 A-124 )
SER( 1 A-123 )
ALA( 1 A-122 )
GLU( 1 A-121 )
VAL( 1 A-120 )
TYR( 1 A-119 )
TYR( 1 A-118 )
GLU( 1 A-117 )
GLU( 1 A-116 )
ALA( 1 A-115 )
GLU( 1 A-114 )
GLU( 1 A-113 )
PHE( 1 A-112 )
LEU( 1 A-111 )
SER( 1 A-110 )
LYS( 1 A-109 )
GLY( 1 A-108 )
ASP( 1 A-107 )
LEU( 1 A-106 )
VAL( 1 A-105 )
GLN( 1 A-104 )
ALA( 1 A-103 )
CYS( 1 A-102 )
GLU( 1 A-101 )
LYS( 1 A-100 )
TYR( 1 A -99 )
TYR( 1 A -98 )
LYS( 1 A -97 )
ALA( 1 A -96 )
ALA( 1 A -95 )
GLU( 1 A -94 )
GLU( 1 A -93 )
ALA( 1 A -92 )
ILE( 1 A -91 )
LYS( 1 A -90 )
LEU( 1 A -89 )
LEU( 1 A -88 )
VAL( 1 A -87 )
ILE( 1 A -86 )
GLU( 1 A -85 )
ASN( 1 A -84 )
ASN( 1 A -83 )
LEU( 1 A -82 )
LYS( 1 A -81 )
GLU( 1 A -80 )
ILE( 1 A -79 )
THR( 1 A -78 )
ASN( 1 A -77 )
ASN( 1 A -76 )
VAL( 1 A -75 )
LYS( 1 A -74 )
ASN( 1 A -73 )
LYS( 1 A -72 )
GLY( 1 A -71 )
ARG( 1 A -70 )
TRP( 1 A -69 )
LYS( 1 A -68 )
SER( 1 A -67 )
GLU( 1 A -66 )
ASN( 1 A -65 )
LEU( 1 A -64 )
PHE( 1 A -63 )
LYS( 1 A -62 )
ALA( 1 A -61 )
SER( 1 A -60 )
LYS( 1 A -59 )
LEU( 1 A -58 )
LEU( 1 A -57 )
ARG( 1 A -56 )
SER( 1 A -55 )
ASN( 1 A -54 )
ASN( 1 A -53 )
THR( 1 A -52 )
GLU( 1 A -51 )
ILE( 1 A -50 )
PRO( 1 A -49 )
ILE( 1 A -48 )
LEU( 1 A -47 )
TRP( 1 A -46 )
LYS( 1 A -45 )
SER( 1 A -44 )
ALA( 1 A -43 )
TRP( 1 A -42 )
THR( 1 A -41 )
LEU( 1 A -40 )
HIS( 1 A -39 )
VAL( 1 A -38 )
GLU( 1 A -37 )
GLY( 1 A -36 )
PHE( 1 A -35 )
HIS( 1 A -34 )
GLU( 1 A -33 )
LEU( 1 A -32 )
SER( 1 A -31 )
LEU( 1 A -30 )
ASN( 1 A -29 )
GLU( 1 A -28 )
LYS( 1 A -27 )
GLU( 1 A -26 )
VAL( 1 A -25 )
LYS( 1 A -24 )
LYS( 1 A -23 )
LEU( 1 A -22 )
LYS( 1 A -21 )
GLU( 1 A -20 )
ASP( 1 A -19 )
VAL( 1 A -18 )
ARG( 1 A -17 )
LYS( 1 A -16 )
LEU( 1 A -15 )
VAL( 1 A -14 )
ILE( 1 A -13 )
PHE( 1 A -12 )
ALA( 1 A -11 )
VAL( 1 A -10 )
ASN( 1 A -9 )
SER( 1 A -8 )
LEU( 1 A -7 )
GLU( 1 A -6 )
HIS( 1 A -5 )
HIS( 1 A -4 )
HIS( 1 A -3 )
HIS( 1 A -2 )
HIS( 1 A -1 )
HIS( 1 A 0 )
MET( 2 A-128 )
SER( 2 A-127 )
ILE( 2 A-126 )
SER( 2 A-125 )
THR( 2 A-124 )
SER( 2 A-123 )
ALA( 2 A-122 )
GLU( 2 A-121 )
VAL( 2 A-120 )
TYR( 2 A-119 )
TYR( 2 A-118 )
GLU( 2 A-117 )
GLU( 2 A-116 )
ALA( 2 A-115 )
GLU( 2 A-114 )
GLU( 2 A-113 )
PHE( 2 A-112 )
LEU( 2 A-111 )
SER( 2 A-110 )
LYS( 2 A-109 )
GLY( 2 A-108 )
ASP( 2 A-107 )
LEU( 2 A-106 )
VAL( 2 A-105 )
GLN( 2 A-104 )
ALA( 2 A-103 )
CYS( 2 A-102 )
GLU( 2 A-101 )
LYS( 2 A-100 )
TYR( 2 A -99 )
TYR( 2 A -98 )
LYS( 2 A -97 )
ALA( 2 A -96 )
ALA( 2 A -95 )
GLU( 2 A -94 )
GLU( 2 A -93 )
ALA( 2 A -92 )
ILE( 2 A -91 )
LYS( 2 A -90 )
LEU( 2 A -89 )
LEU( 2 A -88 )
VAL( 2 A -87 )
ILE( 2 A -86 )
GLU( 2 A -85 )
ASN( 2 A -84 )
ASN( 2 A -83 )
LEU( 2 A -82 )
LYS( 2 A -81 )
GLU( 2 A -80 )
ILE( 2 A -79 )
THR( 2 A -78 )
ASN( 2 A -77 )
ASN( 2 A -76 )
VAL( 2 A -75 )
LYS( 2 A -74 )
ASN( 2 A -73 )
LYS( 2 A -72 )
GLY( 2 A -71 )
ARG( 2 A -70 )
TRP( 2 A -69 )
LYS( 2 A -68 )
SER( 2 A -67 )
GLU( 2 A -66 )
ASN( 2 A -65 )
LEU( 2 A -64 )
PHE( 2 A -63 )
LYS( 2 A -62 )
ALA( 2 A -61 )
SER( 2 A -60 )
LYS( 2 A -59 )
LEU( 2 A -58 )
LEU( 2 A -57 )
ARG( 2 A -56 )
SER( 2 A -55 )
ASN( 2 A -54 )
ASN( 2 A -53 )
THR( 2 A -52 )
GLU( 2 A -51 )
ILE( 2 A -50 )
PRO( 2 A -49 )
ILE( 2 A -48 )
LEU( 2 A -47 )
TRP( 2 A -46 )
LYS( 2 A -45 )
SER( 2 A -44 )
ALA( 2 A -43 )
TRP( 2 A -42 )
THR( 2 A -41 )
LEU( 2 A -40 )
HIS( 2 A -39 )
VAL( 2 A -38 )
GLU( 2 A -37 )
GLY( 2 A -36 )
PHE( 2 A -35 )
HIS( 2 A -34 )
GLU( 2 A -33 )
LEU( 2 A -32 )
SER( 2 A -31 )
LEU( 2 A -30 )
ASN( 2 A -29 )
GLU( 2 A -28 )
LYS( 2 A -27 )
GLU( 2 A -26 )
VAL( 2 A -25 )
LYS( 2 A -24 )
LYS( 2 A -23 )
LEU( 2 A -22 )
LYS( 2 A -21 )
GLU( 2 A -20 )
ASP( 2 A -19 )
VAL( 2 A -18 )
ARG( 2 A -17 )
LYS( 2 A -16 )
LEU( 2 A -15 )
VAL( 2 A -14 )
ILE( 2 A -13 )
PHE( 2 A -12 )
ALA( 2 A -11 )
VAL( 2 A -10 )
ASN( 2 A -9 )
SER( 2 A -8 )
LEU( 2 A -7 )
GLU( 2 A -6 )
HIS( 2 A -5 )
HIS( 2 A -4 )
HIS( 2 A -3 )
HIS( 2 A -2 )
HIS( 2 A -1 )
HIS( 2 A 0 )
MET( 3 A-128 )
SER( 3 A-127 )
ILE( 3 A-126 )
SER( 3 A-125 )
THR( 3 A-124 )
SER( 3 A-123 )
ALA( 3 A-122 )
GLU( 3 A-121 )
VAL( 3 A-120 )
TYR( 3 A-119 )
TYR( 3 A-118 )
GLU( 3 A-117 )
GLU( 3 A-116 )
ALA( 3 A-115 )
GLU( 3 A-114 )
GLU( 3 A-113 )
PHE( 3 A-112 )
LEU( 3 A-111 )
SER( 3 A-110 )
LYS( 3 A-109 )
GLY( 3 A-108 )
ASP( 3 A-107 )
LEU( 3 A-106 )
VAL( 3 A-105 )
GLN( 3 A-104 )
ALA( 3 A-103 )
CYS( 3 A-102 )
GLU( 3 A-101 )
LYS( 3 A-100 )
TYR( 3 A -99 )
TYR( 3 A -98 )
LYS( 3 A -97 )
ALA( 3 A -96 )
ALA( 3 A -95 )
GLU( 3 A -94 )
GLU( 3 A -93 )
ALA( 3 A -92 )
ILE( 3 A -91 )
LYS( 3 A -90 )
LEU( 3 A -89 )
LEU( 3 A -88 )
VAL( 3 A -87 )
ILE( 3 A -86 )
GLU( 3 A -85 )
ASN( 3 A -84 )
ASN( 3 A -83 )
LEU( 3 A -82 )
LYS( 3 A -81 )
GLU( 3 A -80 )
ILE( 3 A -79 )
THR( 3 A -78 )
ASN( 3 A -77 )
ASN( 3 A -76 )
VAL( 3 A -75 )
LYS( 3 A -74 )
ASN( 3 A -73 )
LYS( 3 A -72 )
GLY( 3 A -71 )
ARG( 3 A -70 )
TRP( 3 A -69 )
LYS( 3 A -68 )
SER( 3 A -67 )
GLU( 3 A -66 )
ASN( 3 A -65 )
LEU( 3 A -64 )
PHE( 3 A -63 )
LYS( 3 A -62 )
ALA( 3 A -61 )
SER( 3 A -60 )
LYS( 3 A -59 )
LEU( 3 A -58 )
LEU( 3 A -57 )
ARG( 3 A -56 )
SER( 3 A -55 )
ASN( 3 A -54 )
ASN( 3 A -53 )
THR( 3 A -52 )
GLU( 3 A -51 )
ILE( 3 A -50 )
PRO( 3 A -49 )
ILE( 3 A -48 )
LEU( 3 A -47 )
TRP( 3 A -46 )
LYS( 3 A -45 )
SER( 3 A -44 )
ALA( 3 A -43 )
TRP( 3 A -42 )
THR( 3 A -41 )
LEU( 3 A -40 )
HIS( 3 A -39 )
VAL( 3 A -38 )
GLU( 3 A -37 )
GLY( 3 A -36 )
PHE( 3 A -35 )
HIS( 3 A -34 )
GLU( 3 A -33 )
LEU( 3 A -32 )
SER( 3 A -31 )
LEU( 3 A -30 )
ASN( 3 A -29 )
GLU( 3 A -28 )
LYS( 3 A -27 )
GLU( 3 A -26 )
VAL( 3 A -25 )
LYS( 3 A -24 )
LYS( 3 A -23 )
LEU( 3 A -22 )
LYS( 3 A -21 )
GLU( 3 A -20 )
ASP( 3 A -19 )
VAL( 3 A -18 )
ARG( 3 A -17 )
LYS( 3 A -16 )
LEU( 3 A -15 )
VAL( 3 A -14 )
ILE( 3 A -13 )
PHE( 3 A -12 )
ALA( 3 A -11 )
VAL( 3 A -10 )
ASN( 3 A -9 )
SER( 3 A -8 )
LEU( 3 A -7 )
GLU( 3 A -6 )
HIS( 3 A -5 )
HIS( 3 A -4 )
HIS( 3 A -3 )
HIS( 3 A -2 )
HIS( 3 A -1 )
HIS( 3 A 0 )
MET( 4 A-128 )
SER( 4 A-127 )
ILE( 4 A-126 )
SER( 4 A-125 )
THR( 4 A-124 )
SER( 4 A-123 )
ALA( 4 A-122 )
GLU( 4 A-121 )
VAL( 4 A-120 )
TYR( 4 A-119 )
TYR( 4 A-118 )
GLU( 4 A-117 )
GLU( 4 A-116 )
ALA( 4 A-115 )
GLU( 4 A-114 )
GLU( 4 A-113 )
PHE( 4 A-112 )
LEU( 4 A-111 )
SER( 4 A-110 )
LYS( 4 A-109 )
GLY( 4 A-108 )
ASP( 4 A-107 )
LEU( 4 A-106 )
VAL( 4 A-105 )
GLN( 4 A-104 )
ALA( 4 A-103 )
CYS( 4 A-102 )
GLU( 4 A-101 )
LYS( 4 A-100 )
TYR( 4 A -99 )
TYR( 4 A -98 )
LYS( 4 A -97 )
ALA( 4 A -96 )
ALA( 4 A -95 )
GLU( 4 A -94 )
GLU( 4 A -93 )
ALA( 4 A -92 )
ILE( 4 A -91 )
LYS( 4 A -90 )
LEU( 4 A -89 )
LEU( 4 A -88 )
VAL( 4 A -87 )
ILE( 4 A -86 )
GLU( 4 A -85 )
ASN( 4 A -84 )
ASN( 4 A -83 )
LEU( 4 A -82 )
LYS( 4 A -81 )
GLU( 4 A -80 )
ILE( 4 A -79 )
THR( 4 A -78 )
ASN( 4 A -77 )
ASN( 4 A -76 )
VAL( 4 A -75 )
LYS( 4 A -74 )
ASN( 4 A -73 )
LYS( 4 A -72 )
GLY( 4 A -71 )
ARG( 4 A -70 )
TRP( 4 A -69 )
LYS( 4 A -68 )
SER( 4 A -67 )
GLU( 4 A -66 )
ASN( 4 A -65 )
LEU( 4 A -64 )
PHE( 4 A -63 )
LYS( 4 A -62 )
ALA( 4 A -61 )
SER( 4 A -60 )
LYS( 4 A -59 )
LEU( 4 A -58 )
LEU( 4 A -57 )
ARG( 4 A -56 )
SER( 4 A -55 )
ASN( 4 A -54 )
ASN( 4 A -53 )
THR( 4 A -52 )
GLU( 4 A -51 )
ILE( 4 A -50 )
PRO( 4 A -49 )
ILE( 4 A -48 )
LEU( 4 A -47 )
TRP( 4 A -46 )
LYS( 4 A -45 )
SER( 4 A -44 )
ALA( 4 A -43 )
TRP( 4 A -42 )
THR( 4 A -41 )
LEU( 4 A -40 )
HIS( 4 A -39 )
VAL( 4 A -38 )
GLU( 4 A -37 )
GLY( 4 A -36 )
PHE( 4 A -35 )
HIS( 4 A -34 )
GLU( 4 A -33 )
LEU( 4 A -32 )
SER( 4 A -31 )
LEU( 4 A -30 )
ASN( 4 A -29 )
GLU( 4 A -28 )
LYS( 4 A -27 )
GLU( 4 A -26 )
VAL( 4 A -25 )
LYS( 4 A -24 )
LYS( 4 A -23 )
LEU( 4 A -22 )
LYS( 4 A -21 )
GLU( 4 A -20 )
ASP( 4 A -19 )
VAL( 4 A -18 )
ARG( 4 A -17 )
LYS( 4 A -16 )
LEU( 4 A -15 )
VAL( 4 A -14 )
ILE( 4 A -13 )
PHE( 4 A -12 )
ALA( 4 A -11 )
VAL( 4 A -10 )
ASN( 4 A -9 )
SER( 4 A -8 )
LEU( 4 A -7 )
GLU( 4 A -6 )
HIS( 4 A -5 )
HIS( 4 A -4 )
HIS( 4 A -3 )
HIS( 4 A -2 )
HIS( 4 A -1 )
HIS( 4 A 0 )
MET( 5 A-128 )
SER( 5 A-127 )
ILE( 5 A-126 )
SER( 5 A-125 )
THR( 5 A-124 )
SER( 5 A-123 )
ALA( 5 A-122 )
GLU( 5 A-121 )
VAL( 5 A-120 )
TYR( 5 A-119 )
TYR( 5 A-118 )
GLU( 5 A-117 )
GLU( 5 A-116 )
ALA( 5 A-115 )
GLU( 5 A-114 )
GLU( 5 A-113 )
PHE( 5 A-112 )
LEU( 5 A-111 )
SER( 5 A-110 )
LYS( 5 A-109 )
GLY( 5 A-108 )
ASP( 5 A-107 )
LEU( 5 A-106 )
VAL( 5 A-105 )
GLN( 5 A-104 )
ALA( 5 A-103 )
CYS( 5 A-102 )
GLU( 5 A-101 )
LYS( 5 A-100 )
TYR( 5 A -99 )
TYR( 5 A -98 )
LYS( 5 A -97 )
ALA( 5 A -96 )
ALA( 5 A -95 )
GLU( 5 A -94 )
GLU( 5 A -93 )
ALA( 5 A -92 )
ILE( 5 A -91 )
LYS( 5 A -90 )
LEU( 5 A -89 )
LEU( 5 A -88 )
VAL( 5 A -87 )
ILE( 5 A -86 )
GLU( 5 A -85 )
ASN( 5 A -84 )
ASN( 5 A -83 )
LEU( 5 A -82 )
LYS( 5 A -81 )
GLU( 5 A -80 )
ILE( 5 A -79 )
THR( 5 A -78 )
ASN( 5 A -77 )
ASN( 5 A -76 )
VAL( 5 A -75 )
LYS( 5 A -74 )
ASN( 5 A -73 )
LYS( 5 A -72 )
GLY( 5 A -71 )
ARG( 5 A -70 )
TRP( 5 A -69 )
LYS( 5 A -68 )
SER( 5 A -67 )
GLU( 5 A -66 )
ASN( 5 A -65 )
LEU( 5 A -64 )
PHE( 5 A -63 )
LYS( 5 A -62 )
ALA( 5 A -61 )
SER( 5 A -60 )
LYS( 5 A -59 )
LEU( 5 A -58 )
LEU( 5 A -57 )
ARG( 5 A -56 )
SER( 5 A -55 )
ASN( 5 A -54 )
ASN( 5 A -53 )
THR( 5 A -52 )
GLU( 5 A -51 )
ILE( 5 A -50 )
PRO( 5 A -49 )
ILE( 5 A -48 )
LEU( 5 A -47 )
TRP( 5 A -46 )
LYS( 5 A -45 )
SER( 5 A -44 )
ALA( 5 A -43 )
TRP( 5 A -42 )
THR( 5 A -41 )
LEU( 5 A -40 )
HIS( 5 A -39 )
VAL( 5 A -38 )
GLU( 5 A -37 )
GLY( 5 A -36 )
PHE( 5 A -35 )
HIS( 5 A -34 )
GLU( 5 A -33 )
LEU( 5 A -32 )
SER( 5 A -31 )
LEU( 5 A -30 )
ASN( 5 A -29 )
GLU( 5 A -28 )
LYS( 5 A -27 )
GLU( 5 A -26 )
VAL( 5 A -25 )
LYS( 5 A -24 )
LYS( 5 A -23 )
LEU( 5 A -22 )
LYS( 5 A -21 )
GLU( 5 A -20 )
ASP( 5 A -19 )
VAL( 5 A -18 )
ARG( 5 A -17 )
LYS( 5 A -16 )
LEU( 5 A -15 )
VAL( 5 A -14 )
ILE( 5 A -13 )
PHE( 5 A -12 )
ALA( 5 A -11 )
VAL( 5 A -10 )
ASN( 5 A -9 )
SER( 5 A -8 )
LEU( 5 A -7 )
GLU( 5 A -6 )
HIS( 5 A -5 )
HIS( 5 A -4 )
HIS( 5 A -3 )
HIS( 5 A -2 )
HIS( 5 A -1 )
HIS( 5 A 0 )
MET( 6 A-128 )
SER( 6 A-127 )
ILE( 6 A-126 )
SER( 6 A-125 )
THR( 6 A-124 )
SER( 6 A-123 )
ALA( 6 A-122 )
GLU( 6 A-121 )
VAL( 6 A-120 )
TYR( 6 A-119 )
TYR( 6 A-118 )
GLU( 6 A-117 )
GLU( 6 A-116 )
ALA( 6 A-115 )
GLU( 6 A-114 )
GLU( 6 A-113 )
PHE( 6 A-112 )
LEU( 6 A-111 )
SER( 6 A-110 )
LYS( 6 A-109 )
GLY( 6 A-108 )
ASP( 6 A-107 )
LEU( 6 A-106 )
VAL( 6 A-105 )
GLN( 6 A-104 )
ALA( 6 A-103 )
CYS( 6 A-102 )
GLU( 6 A-101 )
LYS( 6 A-100 )
TYR( 6 A -99 )
TYR( 6 A -98 )
LYS( 6 A -97 )
ALA( 6 A -96 )
ALA( 6 A -95 )
GLU( 6 A -94 )
GLU( 6 A -93 )
ALA( 6 A -92 )
ILE( 6 A -91 )
LYS( 6 A -90 )
LEU( 6 A -89 )
LEU( 6 A -88 )
VAL( 6 A -87 )
ILE( 6 A -86 )
GLU( 6 A -85 )
ASN( 6 A -84 )
ASN( 6 A -83 )
LEU( 6 A -82 )
LYS( 6 A -81 )
GLU( 6 A -80 )
ILE( 6 A -79 )
THR( 6 A -78 )
ASN( 6 A -77 )
ASN( 6 A -76 )
VAL( 6 A -75 )
LYS( 6 A -74 )
ASN( 6 A -73 )
LYS( 6 A -72 )
GLY( 6 A -71 )
ARG( 6 A -70 )
TRP( 6 A -69 )
LYS( 6 A -68 )
SER( 6 A -67 )
GLU( 6 A -66 )
ASN( 6 A -65 )
LEU( 6 A -64 )
PHE( 6 A -63 )
LYS( 6 A -62 )
ALA( 6 A -61 )
SER( 6 A -60 )
LYS( 6 A -59 )
LEU( 6 A -58 )
LEU( 6 A -57 )
ARG( 6 A -56 )
SER( 6 A -55 )
ASN( 6 A -54 )
ASN( 6 A -53 )
THR( 6 A -52 )
GLU( 6 A -51 )
ILE( 6 A -50 )
PRO( 6 A -49 )
ILE( 6 A -48 )
LEU( 6 A -47 )
TRP( 6 A -46 )
LYS( 6 A -45 )
SER( 6 A -44 )
ALA( 6 A -43 )
TRP( 6 A -42 )
THR( 6 A -41 )
LEU( 6 A -40 )
HIS( 6 A -39 )
VAL( 6 A -38 )
GLU( 6 A -37 )
GLY( 6 A -36 )
PHE( 6 A -35 )
HIS( 6 A -34 )
GLU( 6 A -33 )
LEU( 6 A -32 )
SER( 6 A -31 )
LEU( 6 A -30 )
ASN( 6 A -29 )
GLU( 6 A -28 )
LYS( 6 A -27 )
GLU( 6 A -26 )
VAL( 6 A -25 )
LYS( 6 A -24 )
LYS( 6 A -23 )
LEU( 6 A -22 )
LYS( 6 A -21 )
GLU( 6 A -20 )
ASP( 6 A -19 )
VAL( 6 A -18 )
ARG( 6 A -17 )
LYS( 6 A -16 )
LEU( 6 A -15 )
VAL( 6 A -14 )
ILE( 6 A -13 )
PHE( 6 A -12 )
ALA( 6 A -11 )
VAL( 6 A -10 )
ASN( 6 A -9 )
SER( 6 A -8 )
LEU( 6 A -7 )
GLU( 6 A -6 )
HIS( 6 A -5 )
HIS( 6 A -4 )
HIS( 6 A -3 )
HIS( 6 A -2 )
HIS( 6 A -1 )
HIS( 6 A 0 )
MET( 7 A-128 )
SER( 7 A-127 )
ILE( 7 A-126 )
SER( 7 A-125 )
THR( 7 A-124 )
SER( 7 A-123 )
ALA( 7 A-122 )
GLU( 7 A-121 )
VAL( 7 A-120 )
TYR( 7 A-119 )
TYR( 7 A-118 )
GLU( 7 A-117 )
GLU( 7 A-116 )
ALA( 7 A-115 )
GLU( 7 A-114 )
GLU( 7 A-113 )
PHE( 7 A-112 )
LEU( 7 A-111 )
SER( 7 A-110 )
LYS( 7 A-109 )
GLY( 7 A-108 )
ASP( 7 A-107 )
LEU( 7 A-106 )
VAL( 7 A-105 )
GLN( 7 A-104 )
ALA( 7 A-103 )
CYS( 7 A-102 )
GLU( 7 A-101 )
LYS( 7 A-100 )
TYR( 7 A -99 )
TYR( 7 A -98 )
LYS( 7 A -97 )
ALA( 7 A -96 )
ALA( 7 A -95 )
GLU( 7 A -94 )
GLU( 7 A -93 )
ALA( 7 A -92 )
ILE( 7 A -91 )
LYS( 7 A -90 )
LEU( 7 A -89 )
LEU( 7 A -88 )
VAL( 7 A -87 )
ILE( 7 A -86 )
GLU( 7 A -85 )
ASN( 7 A -84 )
ASN( 7 A -83 )
LEU( 7 A -82 )
LYS( 7 A -81 )
GLU( 7 A -80 )
ILE( 7 A -79 )
THR( 7 A -78 )
ASN( 7 A -77 )
ASN( 7 A -76 )
VAL( 7 A -75 )
LYS( 7 A -74 )
ASN( 7 A -73 )
LYS( 7 A -72 )
GLY( 7 A -71 )
ARG( 7 A -70 )
TRP( 7 A -69 )
LYS( 7 A -68 )
SER( 7 A -67 )
GLU( 7 A -66 )
ASN( 7 A -65 )
LEU( 7 A -64 )
PHE( 7 A -63 )
LYS( 7 A -62 )
ALA( 7 A -61 )
SER( 7 A -60 )
LYS( 7 A -59 )
LEU( 7 A -58 )
LEU( 7 A -57 )
ARG( 7 A -56 )
SER( 7 A -55 )
ASN( 7 A -54 )
ASN( 7 A -53 )
THR( 7 A -52 )
GLU( 7 A -51 )
ILE( 7 A -50 )
PRO( 7 A -49 )
ILE( 7 A -48 )
LEU( 7 A -47 )
TRP( 7 A -46 )
LYS( 7 A -45 )
SER( 7 A -44 )
ALA( 7 A -43 )
TRP( 7 A -42 )
THR( 7 A -41 )
LEU( 7 A -40 )
HIS( 7 A -39 )
VAL( 7 A -38 )
GLU( 7 A -37 )
GLY( 7 A -36 )
PHE( 7 A -35 )
HIS( 7 A -34 )
GLU( 7 A -33 )
LEU( 7 A -32 )
SER( 7 A -31 )
LEU( 7 A -30 )
ASN( 7 A -29 )
GLU( 7 A -28 )
LYS( 7 A -27 )
GLU( 7 A -26 )
VAL( 7 A -25 )
LYS( 7 A -24 )
LYS( 7 A -23 )
LEU( 7 A -22 )
LYS( 7 A -21 )
GLU( 7 A -20 )
ASP( 7 A -19 )
VAL( 7 A -18 )
ARG( 7 A -17 )
LYS( 7 A -16 )
LEU( 7 A -15 )
VAL( 7 A -14 )
ILE( 7 A -13 )
PHE( 7 A -12 )
ALA( 7 A -11 )
VAL( 7 A -10 )
ASN( 7 A -9 )
SER( 7 A -8 )
LEU( 7 A -7 )
GLU( 7 A -6 )
HIS( 7 A -5 )
HIS( 7 A -4 )
HIS( 7 A -3 )
HIS( 7 A -2 )
HIS( 7 A -1 )
HIS( 7 A 0 )
MET( 8 A-128 )
SER( 8 A-127 )
ILE( 8 A-126 )
SER( 8 A-125 )
THR( 8 A-124 )
SER( 8 A-123 )
ALA( 8 A-122 )
GLU( 8 A-121 )
VAL( 8 A-120 )
TYR( 8 A-119 )
TYR( 8 A-118 )
GLU( 8 A-117 )
GLU( 8 A-116 )
ALA( 8 A-115 )
GLU( 8 A-114 )
GLU( 8 A-113 )
PHE( 8 A-112 )
LEU( 8 A-111 )
SER( 8 A-110 )
LYS( 8 A-109 )
GLY( 8 A-108 )
ASP( 8 A-107 )
LEU( 8 A-106 )
VAL( 8 A-105 )
GLN( 8 A-104 )
ALA( 8 A-103 )
CYS( 8 A-102 )
GLU( 8 A-101 )
LYS( 8 A-100 )
TYR( 8 A -99 )
TYR( 8 A -98 )
LYS( 8 A -97 )
ALA( 8 A -96 )
ALA( 8 A -95 )
GLU( 8 A -94 )
GLU( 8 A -93 )
ALA( 8 A -92 )
ILE( 8 A -91 )
LYS( 8 A -90 )
LEU( 8 A -89 )
LEU( 8 A -88 )
VAL( 8 A -87 )
ILE( 8 A -86 )
GLU( 8 A -85 )
ASN( 8 A -84 )
ASN( 8 A -83 )
LEU( 8 A -82 )
LYS( 8 A -81 )
GLU( 8 A -80 )
ILE( 8 A -79 )
THR( 8 A -78 )
ASN( 8 A -77 )
ASN( 8 A -76 )
VAL( 8 A -75 )
LYS( 8 A -74 )
ASN( 8 A -73 )
LYS( 8 A -72 )
GLY( 8 A -71 )
ARG( 8 A -70 )
TRP( 8 A -69 )
LYS( 8 A -68 )
SER( 8 A -67 )
GLU( 8 A -66 )
ASN( 8 A -65 )
LEU( 8 A -64 )
PHE( 8 A -63 )
LYS( 8 A -62 )
ALA( 8 A -61 )
SER( 8 A -60 )
LYS( 8 A -59 )
LEU( 8 A -58 )
LEU( 8 A -57 )
ARG( 8 A -56 )
SER( 8 A -55 )
ASN( 8 A -54 )
ASN( 8 A -53 )
THR( 8 A -52 )
GLU( 8 A -51 )
ILE( 8 A -50 )
PRO( 8 A -49 )
ILE( 8 A -48 )
LEU( 8 A -47 )
TRP( 8 A -46 )
LYS( 8 A -45 )
SER( 8 A -44 )
ALA( 8 A -43 )
TRP( 8 A -42 )
THR( 8 A -41 )
LEU( 8 A -40 )
HIS( 8 A -39 )
VAL( 8 A -38 )
GLU( 8 A -37 )
GLY( 8 A -36 )
PHE( 8 A -35 )
HIS( 8 A -34 )
GLU( 8 A -33 )
LEU( 8 A -32 )
SER( 8 A -31 )
LEU( 8 A -30 )
ASN( 8 A -29 )
GLU( 8 A -28 )
LYS( 8 A -27 )
GLU( 8 A -26 )
VAL( 8 A -25 )
LYS( 8 A -24 )
LYS( 8 A -23 )
LEU( 8 A -22 )
LYS( 8 A -21 )
GLU( 8 A -20 )
ASP( 8 A -19 )
VAL( 8 A -18 )
ARG( 8 A -17 )
LYS( 8 A -16 )
LEU( 8 A -15 )
VAL( 8 A -14 )
ILE( 8 A -13 )
PHE( 8 A -12 )
ALA( 8 A -11 )
VAL( 8 A -10 )
ASN( 8 A -9 )
SER( 8 A -8 )
LEU( 8 A -7 )
GLU( 8 A -6 )
HIS( 8 A -5 )
HIS( 8 A -4 )
HIS( 8 A -3 )
HIS( 8 A -2 )
HIS( 8 A -1 )
HIS( 8 A 0 )
MET( 9 A-128 )
SER( 9 A-127 )
ILE( 9 A-126 )
SER( 9 A-125 )
THR( 9 A-124 )
SER( 9 A-123 )
ALA( 9 A-122 )
GLU( 9 A-121 )
VAL( 9 A-120 )
TYR( 9 A-119 )
TYR( 9 A-118 )
GLU( 9 A-117 )
GLU( 9 A-116 )
ALA( 9 A-115 )
GLU( 9 A-114 )
GLU( 9 A-113 )
PHE( 9 A-112 )
LEU( 9 A-111 )
SER( 9 A-110 )
LYS( 9 A-109 )
GLY( 9 A-108 )
ASP( 9 A-107 )
LEU( 9 A-106 )
VAL( 9 A-105 )
GLN( 9 A-104 )
ALA( 9 A-103 )
CYS( 9 A-102 )
GLU( 9 A-101 )
LYS( 9 A-100 )
TYR( 9 A -99 )
TYR( 9 A -98 )
LYS( 9 A -97 )
ALA( 9 A -96 )
ALA( 9 A -95 )
GLU( 9 A -94 )
GLU( 9 A -93 )
ALA( 9 A -92 )
ILE( 9 A -91 )
LYS( 9 A -90 )
LEU( 9 A -89 )
LEU( 9 A -88 )
VAL( 9 A -87 )
ILE( 9 A -86 )
GLU( 9 A -85 )
ASN( 9 A -84 )
ASN( 9 A -83 )
LEU( 9 A -82 )
LYS( 9 A -81 )
GLU( 9 A -80 )
ILE( 9 A -79 )
THR( 9 A -78 )
ASN( 9 A -77 )
ASN( 9 A -76 )
VAL( 9 A -75 )
LYS( 9 A -74 )
ASN( 9 A -73 )
LYS( 9 A -72 )
GLY( 9 A -71 )
ARG( 9 A -70 )
TRP( 9 A -69 )
LYS( 9 A -68 )
SER( 9 A -67 )
GLU( 9 A -66 )
ASN( 9 A -65 )
LEU( 9 A -64 )
PHE( 9 A -63 )
LYS( 9 A -62 )
ALA( 9 A -61 )
SER( 9 A -60 )
LYS( 9 A -59 )
LEU( 9 A -58 )
LEU( 9 A -57 )
ARG( 9 A -56 )
SER( 9 A -55 )
ASN( 9 A -54 )
ASN( 9 A -53 )
THR( 9 A -52 )
GLU( 9 A -51 )
ILE( 9 A -50 )
PRO( 9 A -49 )
ILE( 9 A -48 )
LEU( 9 A -47 )
TRP( 9 A -46 )
LYS( 9 A -45 )
SER( 9 A -44 )
ALA( 9 A -43 )
TRP( 9 A -42 )
THR( 9 A -41 )
LEU( 9 A -40 )
HIS( 9 A -39 )
VAL( 9 A -38 )
GLU( 9 A -37 )
GLY( 9 A -36 )
PHE( 9 A -35 )
HIS( 9 A -34 )
GLU( 9 A -33 )
LEU( 9 A -32 )
SER( 9 A -31 )
LEU( 9 A -30 )
ASN( 9 A -29 )
GLU( 9 A -28 )
LYS( 9 A -27 )
GLU( 9 A -26 )
VAL( 9 A -25 )
LYS( 9 A -24 )
LYS( 9 A -23 )
LEU( 9 A -22 )
LYS( 9 A -21 )
GLU( 9 A -20 )
ASP( 9 A -19 )
VAL( 9 A -18 )
ARG( 9 A -17 )
LYS( 9 A -16 )
LEU( 9 A -15 )
VAL( 9 A -14 )
ILE( 9 A -13 )
PHE( 9 A -12 )
ALA( 9 A -11 )
VAL( 9 A -10 )
ASN( 9 A -9 )
SER( 9 A -8 )
LEU( 9 A -7 )
GLU( 9 A -6 )
HIS( 9 A -5 )
HIS( 9 A -4 )
HIS( 9 A -3 )
HIS( 9 A -2 )
HIS( 9 A -1 )
HIS( 9 A 0 )
MET( 10 A-128 )
SER( 10 A-127 )
ILE( 10 A-126 )
SER( 10 A-125 )
THR( 10 A-124 )
SER( 10 A-123 )
ALA( 10 A-122 )
GLU( 10 A-121 )
VAL( 10 A-120 )
TYR( 10 A-119 )
TYR( 10 A-118 )
GLU( 10 A-117 )
GLU( 10 A-116 )
ALA( 10 A-115 )
GLU( 10 A-114 )
GLU( 10 A-113 )
PHE( 10 A-112 )
LEU( 10 A-111 )
SER( 10 A-110 )
LYS( 10 A-109 )
GLY( 10 A-108 )
ASP( 10 A-107 )
LEU( 10 A-106 )
VAL( 10 A-105 )
GLN( 10 A-104 )
ALA( 10 A-103 )
CYS( 10 A-102 )
GLU( 10 A-101 )
LYS( 10 A-100 )
TYR( 10 A -99 )
TYR( 10 A -98 )
LYS( 10 A -97 )
ALA( 10 A -96 )
ALA( 10 A -95 )
GLU( 10 A -94 )
GLU( 10 A -93 )
ALA( 10 A -92 )
ILE( 10 A -91 )
LYS( 10 A -90 )
LEU( 10 A -89 )
LEU( 10 A -88 )
VAL( 10 A -87 )
ILE( 10 A -86 )
GLU( 10 A -85 )
ASN( 10 A -84 )
ASN( 10 A -83 )
LEU( 10 A -82 )
LYS( 10 A -81 )
GLU( 10 A -80 )
ILE( 10 A -79 )
THR( 10 A -78 )
ASN( 10 A -77 )
ASN( 10 A -76 )
VAL( 10 A -75 )
LYS( 10 A -74 )
ASN( 10 A -73 )
LYS( 10 A -72 )
GLY( 10 A -71 )
ARG( 10 A -70 )
TRP( 10 A -69 )
LYS( 10 A -68 )
SER( 10 A -67 )
GLU( 10 A -66 )
ASN( 10 A -65 )
LEU( 10 A -64 )
PHE( 10 A -63 )
LYS( 10 A -62 )
ALA( 10 A -61 )
SER( 10 A -60 )
LYS( 10 A -59 )
LEU( 10 A -58 )
LEU( 10 A -57 )
ARG( 10 A -56 )
SER( 10 A -55 )
ASN( 10 A -54 )
ASN( 10 A -53 )
THR( 10 A -52 )
GLU( 10 A -51 )
ILE( 10 A -50 )
PRO( 10 A -49 )
ILE( 10 A -48 )
LEU( 10 A -47 )
TRP( 10 A -46 )
LYS( 10 A -45 )
SER( 10 A -44 )
ALA( 10 A -43 )
TRP( 10 A -42 )
THR( 10 A -41 )
LEU( 10 A -40 )
HIS( 10 A -39 )
VAL( 10 A -38 )
GLU( 10 A -37 )
GLY( 10 A -36 )
PHE( 10 A -35 )
HIS( 10 A -34 )
GLU( 10 A -33 )
LEU( 10 A -32 )
SER( 10 A -31 )
LEU( 10 A -30 )
ASN( 10 A -29 )
GLU( 10 A -28 )
LYS( 10 A -27 )
GLU( 10 A -26 )
VAL( 10 A -25 )
LYS( 10 A -24 )
LYS( 10 A -23 )
LEU( 10 A -22 )
LYS( 10 A -21 )
GLU( 10 A -20 )
ASP( 10 A -19 )
VAL( 10 A -18 )
ARG( 10 A -17 )
LYS( 10 A -16 )
LEU( 10 A -15 )
VAL( 10 A -14 )
ILE( 10 A -13 )
PHE( 10 A -12 )
ALA( 10 A -11 )
VAL( 10 A -10 )
ASN( 10 A -9 )
SER( 10 A -8 )
LEU( 10 A -7 )
GLU( 10 A -6 )
HIS( 10 A -5 )
HIS( 10 A -4 )
HIS( 10 A -3 )
HIS( 10 A -2 )
HIS( 10 A -1 )
HIS( 10 A 0 )
MET( 11 A-128 )
SER( 11 A-127 )
ILE( 11 A-126 )
SER( 11 A-125 )
THR( 11 A-124 )
SER( 11 A-123 )
ALA( 11 A-122 )
GLU( 11 A-121 )
VAL( 11 A-120 )
TYR( 11 A-119 )
TYR( 11 A-118 )
GLU( 11 A-117 )
GLU( 11 A-116 )
ALA( 11 A-115 )
GLU( 11 A-114 )
GLU( 11 A-113 )
PHE( 11 A-112 )
LEU( 11 A-111 )
SER( 11 A-110 )
LYS( 11 A-109 )
GLY( 11 A-108 )
ASP( 11 A-107 )
LEU( 11 A-106 )
VAL( 11 A-105 )
GLN( 11 A-104 )
ALA( 11 A-103 )
CYS( 11 A-102 )
GLU( 11 A-101 )
LYS( 11 A-100 )
TYR( 11 A -99 )
TYR( 11 A -98 )
LYS( 11 A -97 )
ALA( 11 A -96 )
ALA( 11 A -95 )
GLU( 11 A -94 )
GLU( 11 A -93 )
ALA( 11 A -92 )
ILE( 11 A -91 )
LYS( 11 A -90 )
LEU( 11 A -89 )
LEU( 11 A -88 )
VAL( 11 A -87 )
ILE( 11 A -86 )
GLU( 11 A -85 )
ASN( 11 A -84 )
ASN( 11 A -83 )
LEU( 11 A -82 )
LYS( 11 A -81 )
GLU( 11 A -80 )
ILE( 11 A -79 )
THR( 11 A -78 )
ASN( 11 A -77 )
ASN( 11 A -76 )
VAL( 11 A -75 )
LYS( 11 A -74 )
ASN( 11 A -73 )
LYS( 11 A -72 )
GLY( 11 A -71 )
ARG( 11 A -70 )
TRP( 11 A -69 )
LYS( 11 A -68 )
SER( 11 A -67 )
GLU( 11 A -66 )
ASN( 11 A -65 )
LEU( 11 A -64 )
PHE( 11 A -63 )
LYS( 11 A -62 )
ALA( 11 A -61 )
SER( 11 A -60 )
LYS( 11 A -59 )
LEU( 11 A -58 )
LEU( 11 A -57 )
ARG( 11 A -56 )
SER( 11 A -55 )
ASN( 11 A -54 )
ASN( 11 A -53 )
THR( 11 A -52 )
GLU( 11 A -51 )
ILE( 11 A -50 )
PRO( 11 A -49 )
ILE( 11 A -48 )
LEU( 11 A -47 )
TRP( 11 A -46 )
LYS( 11 A -45 )
SER( 11 A -44 )
ALA( 11 A -43 )
TRP( 11 A -42 )
THR( 11 A -41 )
LEU( 11 A -40 )
HIS( 11 A -39 )
VAL( 11 A -38 )
GLU( 11 A -37 )
GLY( 11 A -36 )
PHE( 11 A -35 )
HIS( 11 A -34 )
GLU( 11 A -33 )
LEU( 11 A -32 )
SER( 11 A -31 )
LEU( 11 A -30 )
ASN( 11 A -29 )
GLU( 11 A -28 )
LYS( 11 A -27 )
GLU( 11 A -26 )
VAL( 11 A -25 )
LYS( 11 A -24 )
LYS( 11 A -23 )
LEU( 11 A -22 )
LYS( 11 A -21 )
GLU( 11 A -20 )
ASP( 11 A -19 )
VAL( 11 A -18 )
ARG( 11 A -17 )
LYS( 11 A -16 )
LEU( 11 A -15 )
VAL( 11 A -14 )
ILE( 11 A -13 )
PHE( 11 A -12 )
ALA( 11 A -11 )
VAL( 11 A -10 )
ASN( 11 A -9 )
SER( 11 A -8 )
LEU( 11 A -7 )
GLU( 11 A -6 )
HIS( 11 A -5 )
HIS( 11 A -4 )
HIS( 11 A -3 )
HIS( 11 A -2 )
HIS( 11 A -1 )
HIS( 11 A 0 )
MET( 12 A-128 )
SER( 12 A-127 )
ILE( 12 A-126 )
SER( 12 A-125 )
THR( 12 A-124 )
SER( 12 A-123 )
ALA( 12 A-122 )
GLU( 12 A-121 )
VAL( 12 A-120 )
TYR( 12 A-119 )
TYR( 12 A-118 )
GLU( 12 A-117 )
GLU( 12 A-116 )
ALA( 12 A-115 )
GLU( 12 A-114 )
GLU( 12 A-113 )
PHE( 12 A-112 )
LEU( 12 A-111 )
SER( 12 A-110 )
LYS( 12 A-109 )
GLY( 12 A-108 )
ASP( 12 A-107 )
LEU( 12 A-106 )
VAL( 12 A-105 )
GLN( 12 A-104 )
ALA( 12 A-103 )
CYS( 12 A-102 )
GLU( 12 A-101 )
LYS( 12 A-100 )
TYR( 12 A -99 )
TYR( 12 A -98 )
LYS( 12 A -97 )
ALA( 12 A -96 )
ALA( 12 A -95 )
GLU( 12 A -94 )
GLU( 12 A -93 )
ALA( 12 A -92 )
ILE( 12 A -91 )
LYS( 12 A -90 )
LEU( 12 A -89 )
LEU( 12 A -88 )
VAL( 12 A -87 )
ILE( 12 A -86 )
GLU( 12 A -85 )
ASN( 12 A -84 )
ASN( 12 A -83 )
LEU( 12 A -82 )
LYS( 12 A -81 )
GLU( 12 A -80 )
ILE( 12 A -79 )
THR( 12 A -78 )
ASN( 12 A -77 )
ASN( 12 A -76 )
VAL( 12 A -75 )
LYS( 12 A -74 )
ASN( 12 A -73 )
LYS( 12 A -72 )
GLY( 12 A -71 )
ARG( 12 A -70 )
TRP( 12 A -69 )
LYS( 12 A -68 )
SER( 12 A -67 )
GLU( 12 A -66 )
ASN( 12 A -65 )
LEU( 12 A -64 )
PHE( 12 A -63 )
LYS( 12 A -62 )
ALA( 12 A -61 )
SER( 12 A -60 )
LYS( 12 A -59 )
LEU( 12 A -58 )
LEU( 12 A -57 )
ARG( 12 A -56 )
SER( 12 A -55 )
ASN( 12 A -54 )
ASN( 12 A -53 )
THR( 12 A -52 )
GLU( 12 A -51 )
ILE( 12 A -50 )
PRO( 12 A -49 )
ILE( 12 A -48 )
LEU( 12 A -47 )
TRP( 12 A -46 )
LYS( 12 A -45 )
SER( 12 A -44 )
ALA( 12 A -43 )
TRP( 12 A -42 )
THR( 12 A -41 )
LEU( 12 A -40 )
HIS( 12 A -39 )
VAL( 12 A -38 )
GLU( 12 A -37 )
GLY( 12 A -36 )
PHE( 12 A -35 )
HIS( 12 A -34 )
GLU( 12 A -33 )
LEU( 12 A -32 )
SER( 12 A -31 )
LEU( 12 A -30 )
ASN( 12 A -29 )
GLU( 12 A -28 )
LYS( 12 A -27 )
GLU( 12 A -26 )
VAL( 12 A -25 )
LYS( 12 A -24 )
LYS( 12 A -23 )
LEU( 12 A -22 )
LYS( 12 A -21 )
GLU( 12 A -20 )
ASP( 12 A -19 )
VAL( 12 A -18 )
ARG( 12 A -17 )
LYS( 12 A -16 )
LEU( 12 A -15 )
VAL( 12 A -14 )
ILE( 12 A -13 )
PHE( 12 A -12 )
ALA( 12 A -11 )
VAL( 12 A -10 )
ASN( 12 A -9 )
SER( 12 A -8 )
LEU( 12 A -7 )
GLU( 12 A -6 )
HIS( 12 A -5 )
HIS( 12 A -4 )
HIS( 12 A -3 )
HIS( 12 A -2 )
HIS( 12 A -1 )
HIS( 12 A 0 )
MET( 13 A-128 )
SER( 13 A-127 )
ILE( 13 A-126 )
SER( 13 A-125 )
THR( 13 A-124 )
SER( 13 A-123 )
ALA( 13 A-122 )
GLU( 13 A-121 )
VAL( 13 A-120 )
TYR( 13 A-119 )
TYR( 13 A-118 )
GLU( 13 A-117 )
GLU( 13 A-116 )
ALA( 13 A-115 )
GLU( 13 A-114 )
GLU( 13 A-113 )
PHE( 13 A-112 )
LEU( 13 A-111 )
SER( 13 A-110 )
LYS( 13 A-109 )
GLY( 13 A-108 )
ASP( 13 A-107 )
LEU( 13 A-106 )
VAL( 13 A-105 )
GLN( 13 A-104 )
ALA( 13 A-103 )
CYS( 13 A-102 )
GLU( 13 A-101 )
LYS( 13 A-100 )
TYR( 13 A -99 )
TYR( 13 A -98 )
LYS( 13 A -97 )
ALA( 13 A -96 )
ALA( 13 A -95 )
GLU( 13 A -94 )
GLU( 13 A -93 )
ALA( 13 A -92 )
ILE( 13 A -91 )
LYS( 13 A -90 )
LEU( 13 A -89 )
LEU( 13 A -88 )
VAL( 13 A -87 )
ILE( 13 A -86 )
GLU( 13 A -85 )
ASN( 13 A -84 )
ASN( 13 A -83 )
LEU( 13 A -82 )
LYS( 13 A -81 )
GLU( 13 A -80 )
ILE( 13 A -79 )
THR( 13 A -78 )
ASN( 13 A -77 )
ASN( 13 A -76 )
VAL( 13 A -75 )
LYS( 13 A -74 )
ASN( 13 A -73 )
LYS( 13 A -72 )
GLY( 13 A -71 )
ARG( 13 A -70 )
TRP( 13 A -69 )
LYS( 13 A -68 )
SER( 13 A -67 )
GLU( 13 A -66 )
ASN( 13 A -65 )
LEU( 13 A -64 )
PHE( 13 A -63 )
LYS( 13 A -62 )
ALA( 13 A -61 )
SER( 13 A -60 )
LYS( 13 A -59 )
LEU( 13 A -58 )
LEU( 13 A -57 )
ARG( 13 A -56 )
SER( 13 A -55 )
ASN( 13 A -54 )
ASN( 13 A -53 )
THR( 13 A -52 )
GLU( 13 A -51 )
ILE( 13 A -50 )
PRO( 13 A -49 )
ILE( 13 A -48 )
LEU( 13 A -47 )
TRP( 13 A -46 )
LYS( 13 A -45 )
SER( 13 A -44 )
ALA( 13 A -43 )
TRP( 13 A -42 )
THR( 13 A -41 )
LEU( 13 A -40 )
HIS( 13 A -39 )
VAL( 13 A -38 )
GLU( 13 A -37 )
GLY( 13 A -36 )
PHE( 13 A -35 )
HIS( 13 A -34 )
GLU( 13 A -33 )
LEU( 13 A -32 )
SER( 13 A -31 )
LEU( 13 A -30 )
ASN( 13 A -29 )
GLU( 13 A -28 )
LYS( 13 A -27 )
GLU( 13 A -26 )
VAL( 13 A -25 )
LYS( 13 A -24 )
LYS( 13 A -23 )
LEU( 13 A -22 )
LYS( 13 A -21 )
GLU( 13 A -20 )
ASP( 13 A -19 )
VAL( 13 A -18 )
ARG( 13 A -17 )
LYS( 13 A -16 )
LEU( 13 A -15 )
VAL( 13 A -14 )
ILE( 13 A -13 )
PHE( 13 A -12 )
ALA( 13 A -11 )
VAL( 13 A -10 )
ASN( 13 A -9 )
SER( 13 A -8 )
LEU( 13 A -7 )
GLU( 13 A -6 )
HIS( 13 A -5 )
HIS( 13 A -4 )
HIS( 13 A -3 )
HIS( 13 A -2 )
HIS( 13 A -1 )
HIS( 13 A 0 )
MET( 14 A-128 )
SER( 14 A-127 )
ILE( 14 A-126 )
SER( 14 A-125 )
THR( 14 A-124 )
SER( 14 A-123 )
ALA( 14 A-122 )
GLU( 14 A-121 )
VAL( 14 A-120 )
TYR( 14 A-119 )
TYR( 14 A-118 )
GLU( 14 A-117 )
GLU( 14 A-116 )
ALA( 14 A-115 )
GLU( 14 A-114 )
GLU( 14 A-113 )
PHE( 14 A-112 )
LEU( 14 A-111 )
SER( 14 A-110 )
LYS( 14 A-109 )
GLY( 14 A-108 )
ASP( 14 A-107 )
LEU( 14 A-106 )
VAL( 14 A-105 )
GLN( 14 A-104 )
ALA( 14 A-103 )
CYS( 14 A-102 )
GLU( 14 A-101 )
LYS( 14 A-100 )
TYR( 14 A -99 )
TYR( 14 A -98 )
LYS( 14 A -97 )
ALA( 14 A -96 )
ALA( 14 A -95 )
GLU( 14 A -94 )
GLU( 14 A -93 )
ALA( 14 A -92 )
ILE( 14 A -91 )
LYS( 14 A -90 )
LEU( 14 A -89 )
LEU( 14 A -88 )
VAL( 14 A -87 )
ILE( 14 A -86 )
GLU( 14 A -85 )
ASN( 14 A -84 )
ASN( 14 A -83 )
LEU( 14 A -82 )
LYS( 14 A -81 )
GLU( 14 A -80 )
ILE( 14 A -79 )
THR( 14 A -78 )
ASN( 14 A -77 )
ASN( 14 A -76 )
VAL( 14 A -75 )
LYS( 14 A -74 )
ASN( 14 A -73 )
LYS( 14 A -72 )
GLY( 14 A -71 )
ARG( 14 A -70 )
TRP( 14 A -69 )
LYS( 14 A -68 )
SER( 14 A -67 )
GLU( 14 A -66 )
ASN( 14 A -65 )
LEU( 14 A -64 )
PHE( 14 A -63 )
LYS( 14 A -62 )
ALA( 14 A -61 )
SER( 14 A -60 )
LYS( 14 A -59 )
LEU( 14 A -58 )
LEU( 14 A -57 )
ARG( 14 A -56 )
SER( 14 A -55 )
ASN( 14 A -54 )
ASN( 14 A -53 )
THR( 14 A -52 )
GLU( 14 A -51 )
ILE( 14 A -50 )
PRO( 14 A -49 )
ILE( 14 A -48 )
LEU( 14 A -47 )
TRP( 14 A -46 )
LYS( 14 A -45 )
SER( 14 A -44 )
ALA( 14 A -43 )
TRP( 14 A -42 )
THR( 14 A -41 )
LEU( 14 A -40 )
HIS( 14 A -39 )
VAL( 14 A -38 )
GLU( 14 A -37 )
GLY( 14 A -36 )
PHE( 14 A -35 )
HIS( 14 A -34 )
GLU( 14 A -33 )
LEU( 14 A -32 )
SER( 14 A -31 )
LEU( 14 A -30 )
ASN( 14 A -29 )
GLU( 14 A -28 )
LYS( 14 A -27 )
GLU( 14 A -26 )
VAL( 14 A -25 )
LYS( 14 A -24 )
LYS( 14 A -23 )
LEU( 14 A -22 )
LYS( 14 A -21 )
GLU( 14 A -20 )
ASP( 14 A -19 )
VAL( 14 A -18 )
ARG( 14 A -17 )
LYS( 14 A -16 )
LEU( 14 A -15 )
VAL( 14 A -14 )
ILE( 14 A -13 )
PHE( 14 A -12 )
ALA( 14 A -11 )
VAL( 14 A -10 )
ASN( 14 A -9 )
SER( 14 A -8 )
LEU( 14 A -7 )
GLU( 14 A -6 )
HIS( 14 A -5 )
HIS( 14 A -4 )
HIS( 14 A -3 )
HIS( 14 A -2 )
HIS( 14 A -1 )
HIS( 14 A 0 )
MET( 15 A-128 )
SER( 15 A-127 )
ILE( 15 A-126 )
SER( 15 A-125 )
THR( 15 A-124 )
SER( 15 A-123 )
ALA( 15 A-122 )
GLU( 15 A-121 )
VAL( 15 A-120 )
TYR( 15 A-119 )
TYR( 15 A-118 )
GLU( 15 A-117 )
GLU( 15 A-116 )
ALA( 15 A-115 )
GLU( 15 A-114 )
GLU( 15 A-113 )
PHE( 15 A-112 )
LEU( 15 A-111 )
SER( 15 A-110 )
LYS( 15 A-109 )
GLY( 15 A-108 )
ASP( 15 A-107 )
LEU( 15 A-106 )
VAL( 15 A-105 )
GLN( 15 A-104 )
ALA( 15 A-103 )
CYS( 15 A-102 )
GLU( 15 A-101 )
LYS( 15 A-100 )
TYR( 15 A -99 )
TYR( 15 A -98 )
LYS( 15 A -97 )
ALA( 15 A -96 )
ALA( 15 A -95 )
GLU( 15 A -94 )
GLU( 15 A -93 )
ALA( 15 A -92 )
ILE( 15 A -91 )
LYS( 15 A -90 )
LEU( 15 A -89 )
LEU( 15 A -88 )
VAL( 15 A -87 )
ILE( 15 A -86 )
GLU( 15 A -85 )
ASN( 15 A -84 )
ASN( 15 A -83 )
LEU( 15 A -82 )
LYS( 15 A -81 )
GLU( 15 A -80 )
ILE( 15 A -79 )
THR( 15 A -78 )
ASN( 15 A -77 )
ASN( 15 A -76 )
VAL( 15 A -75 )
LYS( 15 A -74 )
ASN( 15 A -73 )
LYS( 15 A -72 )
GLY( 15 A -71 )
ARG( 15 A -70 )
TRP( 15 A -69 )
LYS( 15 A -68 )
SER( 15 A -67 )
GLU( 15 A -66 )
ASN( 15 A -65 )
LEU( 15 A -64 )
PHE( 15 A -63 )
LYS( 15 A -62 )
ALA( 15 A -61 )
SER( 15 A -60 )
LYS( 15 A -59 )
LEU( 15 A -58 )
LEU( 15 A -57 )
ARG( 15 A -56 )
SER( 15 A -55 )
ASN( 15 A -54 )
ASN( 15 A -53 )
THR( 15 A -52 )
GLU( 15 A -51 )
ILE( 15 A -50 )
PRO( 15 A -49 )
ILE( 15 A -48 )
LEU( 15 A -47 )
TRP( 15 A -46 )
LYS( 15 A -45 )
SER( 15 A -44 )
ALA( 15 A -43 )
TRP( 15 A -42 )
THR( 15 A -41 )
LEU( 15 A -40 )
HIS( 15 A -39 )
VAL( 15 A -38 )
GLU( 15 A -37 )
GLY( 15 A -36 )
PHE( 15 A -35 )
HIS( 15 A -34 )
GLU( 15 A -33 )
LEU( 15 A -32 )
SER( 15 A -31 )
LEU( 15 A -30 )
ASN( 15 A -29 )
GLU( 15 A -28 )
LYS( 15 A -27 )
GLU( 15 A -26 )
VAL( 15 A -25 )
LYS( 15 A -24 )
LYS( 15 A -23 )
LEU( 15 A -22 )
LYS( 15 A -21 )
GLU( 15 A -20 )
ASP( 15 A -19 )
VAL( 15 A -18 )
ARG( 15 A -17 )
LYS( 15 A -16 )
LEU( 15 A -15 )
VAL( 15 A -14 )
ILE( 15 A -13 )
PHE( 15 A -12 )
ALA( 15 A -11 )
VAL( 15 A -10 )
ASN( 15 A -9 )
SER( 15 A -8 )
LEU( 15 A -7 )
GLU( 15 A -6 )
HIS( 15 A -5 )
HIS( 15 A -4 )
HIS( 15 A -3 )
HIS( 15 A -2 )
HIS( 15 A -1 )
HIS( 15 A 0 )
MET( 16 A-128 )
SER( 16 A-127 )
ILE( 16 A-126 )
SER( 16 A-125 )
THR( 16 A-124 )
SER( 16 A-123 )
ALA( 16 A-122 )
GLU( 16 A-121 )
VAL( 16 A-120 )
TYR( 16 A-119 )
TYR( 16 A-118 )
GLU( 16 A-117 )
GLU( 16 A-116 )
ALA( 16 A-115 )
GLU( 16 A-114 )
GLU( 16 A-113 )
PHE( 16 A-112 )
LEU( 16 A-111 )
SER( 16 A-110 )
LYS( 16 A-109 )
GLY( 16 A-108 )
ASP( 16 A-107 )
LEU( 16 A-106 )
VAL( 16 A-105 )
GLN( 16 A-104 )
ALA( 16 A-103 )
CYS( 16 A-102 )
GLU( 16 A-101 )
LYS( 16 A-100 )
TYR( 16 A -99 )
TYR( 16 A -98 )
LYS( 16 A -97 )
ALA( 16 A -96 )
ALA( 16 A -95 )
GLU( 16 A -94 )
GLU( 16 A -93 )
ALA( 16 A -92 )
ILE( 16 A -91 )
LYS( 16 A -90 )
LEU( 16 A -89 )
LEU( 16 A -88 )
VAL( 16 A -87 )
ILE( 16 A -86 )
GLU( 16 A -85 )
ASN( 16 A -84 )
ASN( 16 A -83 )
LEU( 16 A -82 )
LYS( 16 A -81 )
GLU( 16 A -80 )
ILE( 16 A -79 )
THR( 16 A -78 )
ASN( 16 A -77 )
ASN( 16 A -76 )
VAL( 16 A -75 )
LYS( 16 A -74 )
ASN( 16 A -73 )
LYS( 16 A -72 )
GLY( 16 A -71 )
ARG( 16 A -70 )
TRP( 16 A -69 )
LYS( 16 A -68 )
SER( 16 A -67 )
GLU( 16 A -66 )
ASN( 16 A -65 )
LEU( 16 A -64 )
PHE( 16 A -63 )
LYS( 16 A -62 )
ALA( 16 A -61 )
SER( 16 A -60 )
LYS( 16 A -59 )
LEU( 16 A -58 )
LEU( 16 A -57 )
ARG( 16 A -56 )
SER( 16 A -55 )
ASN( 16 A -54 )
ASN( 16 A -53 )
THR( 16 A -52 )
GLU( 16 A -51 )
ILE( 16 A -50 )
PRO( 16 A -49 )
ILE( 16 A -48 )
LEU( 16 A -47 )
TRP( 16 A -46 )
LYS( 16 A -45 )
SER( 16 A -44 )
ALA( 16 A -43 )
TRP( 16 A -42 )
THR( 16 A -41 )
LEU( 16 A -40 )
HIS( 16 A -39 )
VAL( 16 A -38 )
GLU( 16 A -37 )
GLY( 16 A -36 )
PHE( 16 A -35 )
HIS( 16 A -34 )
GLU( 16 A -33 )
LEU( 16 A -32 )
SER( 16 A -31 )
LEU( 16 A -30 )
ASN( 16 A -29 )
GLU( 16 A -28 )
LYS( 16 A -27 )
GLU( 16 A -26 )
VAL( 16 A -25 )
LYS( 16 A -24 )
LYS( 16 A -23 )
LEU( 16 A -22 )
LYS( 16 A -21 )
GLU( 16 A -20 )
ASP( 16 A -19 )
VAL( 16 A -18 )
ARG( 16 A -17 )
LYS( 16 A -16 )
LEU( 16 A -15 )
VAL( 16 A -14 )
ILE( 16 A -13 )
PHE( 16 A -12 )
ALA( 16 A -11 )
VAL( 16 A -10 )
ASN( 16 A -9 )
SER( 16 A -8 )
LEU( 16 A -7 )
GLU( 16 A -6 )
HIS( 16 A -5 )
HIS( 16 A -4 )
HIS( 16 A -3 )
HIS( 16 A -2 )
HIS( 16 A -1 )
HIS( 16 A 0 )
MET( 17 A-128 )
SER( 17 A-127 )
ILE( 17 A-126 )
SER( 17 A-125 )
THR( 17 A-124 )
SER( 17 A-123 )
ALA( 17 A-122 )
GLU( 17 A-121 )
VAL( 17 A-120 )
TYR( 17 A-119 )
TYR( 17 A-118 )
GLU( 17 A-117 )
GLU( 17 A-116 )
ALA( 17 A-115 )
GLU( 17 A-114 )
GLU( 17 A-113 )
PHE( 17 A-112 )
LEU( 17 A-111 )
SER( 17 A-110 )
LYS( 17 A-109 )
GLY( 17 A-108 )
ASP( 17 A-107 )
LEU( 17 A-106 )
VAL( 17 A-105 )
GLN( 17 A-104 )
ALA( 17 A-103 )
CYS( 17 A-102 )
GLU( 17 A-101 )
LYS( 17 A-100 )
TYR( 17 A -99 )
TYR( 17 A -98 )
LYS( 17 A -97 )
ALA( 17 A -96 )
ALA( 17 A -95 )
GLU( 17 A -94 )
GLU( 17 A -93 )
ALA( 17 A -92 )
ILE( 17 A -91 )
LYS( 17 A -90 )
LEU( 17 A -89 )
LEU( 17 A -88 )
VAL( 17 A -87 )
ILE( 17 A -86 )
GLU( 17 A -85 )
ASN( 17 A -84 )
ASN( 17 A -83 )
LEU( 17 A -82 )
LYS( 17 A -81 )
GLU( 17 A -80 )
ILE( 17 A -79 )
THR( 17 A -78 )
ASN( 17 A -77 )
ASN( 17 A -76 )
VAL( 17 A -75 )
LYS( 17 A -74 )
ASN( 17 A -73 )
LYS( 17 A -72 )
GLY( 17 A -71 )
ARG( 17 A -70 )
TRP( 17 A -69 )
LYS( 17 A -68 )
SER( 17 A -67 )
GLU( 17 A -66 )
ASN( 17 A -65 )
LEU( 17 A -64 )
PHE( 17 A -63 )
LYS( 17 A -62 )
ALA( 17 A -61 )
SER( 17 A -60 )
LYS( 17 A -59 )
LEU( 17 A -58 )
LEU( 17 A -57 )
ARG( 17 A -56 )
SER( 17 A -55 )
ASN( 17 A -54 )
ASN( 17 A -53 )
THR( 17 A -52 )
GLU( 17 A -51 )
ILE( 17 A -50 )
PRO( 17 A -49 )
ILE( 17 A -48 )
LEU( 17 A -47 )
TRP( 17 A -46 )
LYS( 17 A -45 )
SER( 17 A -44 )
ALA( 17 A -43 )
TRP( 17 A -42 )
THR( 17 A -41 )
LEU( 17 A -40 )
HIS( 17 A -39 )
VAL( 17 A -38 )
GLU( 17 A -37 )
GLY( 17 A -36 )
PHE( 17 A -35 )
HIS( 17 A -34 )
GLU( 17 A -33 )
LEU( 17 A -32 )
SER( 17 A -31 )
LEU( 17 A -30 )
ASN( 17 A -29 )
GLU( 17 A -28 )
LYS( 17 A -27 )
GLU( 17 A -26 )
VAL( 17 A -25 )
LYS( 17 A -24 )
LYS( 17 A -23 )
LEU( 17 A -22 )
LYS( 17 A -21 )
GLU( 17 A -20 )
ASP( 17 A -19 )
VAL( 17 A -18 )
ARG( 17 A -17 )
LYS( 17 A -16 )
LEU( 17 A -15 )
VAL( 17 A -14 )
ILE( 17 A -13 )
PHE( 17 A -12 )
ALA( 17 A -11 )
VAL( 17 A -10 )
ASN( 17 A -9 )
SER( 17 A -8 )
LEU( 17 A -7 )
GLU( 17 A -6 )
HIS( 17 A -5 )
HIS( 17 A -4 )
HIS( 17 A -3 )
HIS( 17 A -2 )
HIS( 17 A -1 )
HIS( 17 A 0 )
MET( 18 A-128 )
SER( 18 A-127 )
ILE( 18 A-126 )
SER( 18 A-125 )
THR( 18 A-124 )
SER( 18 A-123 )
ALA( 18 A-122 )
GLU( 18 A-121 )
VAL( 18 A-120 )
TYR( 18 A-119 )
TYR( 18 A-118 )
GLU( 18 A-117 )
GLU( 18 A-116 )
ALA( 18 A-115 )
GLU( 18 A-114 )
GLU( 18 A-113 )
PHE( 18 A-112 )
LEU( 18 A-111 )
SER( 18 A-110 )
LYS( 18 A-109 )
GLY( 18 A-108 )
ASP( 18 A-107 )
LEU( 18 A-106 )
VAL( 18 A-105 )
GLN( 18 A-104 )
ALA( 18 A-103 )
CYS( 18 A-102 )
GLU( 18 A-101 )
LYS( 18 A-100 )
TYR( 18 A -99 )
TYR( 18 A -98 )
LYS( 18 A -97 )
ALA( 18 A -96 )
ALA( 18 A -95 )
GLU( 18 A -94 )
GLU( 18 A -93 )
ALA( 18 A -92 )
ILE( 18 A -91 )
LYS( 18 A -90 )
LEU( 18 A -89 )
LEU( 18 A -88 )
VAL( 18 A -87 )
ILE( 18 A -86 )
GLU( 18 A -85 )
ASN( 18 A -84 )
ASN( 18 A -83 )
LEU( 18 A -82 )
LYS( 18 A -81 )
GLU( 18 A -80 )
ILE( 18 A -79 )
THR( 18 A -78 )
ASN( 18 A -77 )
ASN( 18 A -76 )
VAL( 18 A -75 )
LYS( 18 A -74 )
ASN( 18 A -73 )
LYS( 18 A -72 )
GLY( 18 A -71 )
ARG( 18 A -70 )
TRP( 18 A -69 )
LYS( 18 A -68 )
SER( 18 A -67 )
GLU( 18 A -66 )
ASN( 18 A -65 )
LEU( 18 A -64 )
PHE( 18 A -63 )
LYS( 18 A -62 )
ALA( 18 A -61 )
SER( 18 A -60 )
LYS( 18 A -59 )
LEU( 18 A -58 )
LEU( 18 A -57 )
ARG( 18 A -56 )
SER( 18 A -55 )
ASN( 18 A -54 )
ASN( 18 A -53 )
THR( 18 A -52 )
GLU( 18 A -51 )
ILE( 18 A -50 )
PRO( 18 A -49 )
ILE( 18 A -48 )
LEU( 18 A -47 )
TRP( 18 A -46 )
LYS( 18 A -45 )
SER( 18 A -44 )
ALA( 18 A -43 )
TRP( 18 A -42 )
THR( 18 A -41 )
LEU( 18 A -40 )
HIS( 18 A -39 )
VAL( 18 A -38 )
GLU( 18 A -37 )
GLY( 18 A -36 )
PHE( 18 A -35 )
HIS( 18 A -34 )
GLU( 18 A -33 )
LEU( 18 A -32 )
SER( 18 A -31 )
LEU( 18 A -30 )
ASN( 18 A -29 )
GLU( 18 A -28 )
LYS( 18 A -27 )
GLU( 18 A -26 )
VAL( 18 A -25 )
LYS( 18 A -24 )
LYS( 18 A -23 )
LEU( 18 A -22 )
LYS( 18 A -21 )
GLU( 18 A -20 )
ASP( 18 A -19 )
VAL( 18 A -18 )
ARG( 18 A -17 )
LYS( 18 A -16 )
LEU( 18 A -15 )
VAL( 18 A -14 )
ILE( 18 A -13 )
PHE( 18 A -12 )
ALA( 18 A -11 )
VAL( 18 A -10 )
ASN( 18 A -9 )
SER( 18 A -8 )
LEU( 18 A -7 )
GLU( 18 A -6 )
HIS( 18 A -5 )
HIS( 18 A -4 )
HIS( 18 A -3 )
HIS( 18 A -2 )
HIS( 18 A -1 )
HIS( 18 A 0 )
MET( 19 A-128 )
SER( 19 A-127 )
ILE( 19 A-126 )
SER( 19 A-125 )
THR( 19 A-124 )
SER( 19 A-123 )
ALA( 19 A-122 )
GLU( 19 A-121 )
VAL( 19 A-120 )
TYR( 19 A-119 )
TYR( 19 A-118 )
GLU( 19 A-117 )
GLU( 19 A-116 )
ALA( 19 A-115 )
GLU( 19 A-114 )
GLU( 19 A-113 )
PHE( 19 A-112 )
LEU( 19 A-111 )
SER( 19 A-110 )
LYS( 19 A-109 )
GLY( 19 A-108 )
ASP( 19 A-107 )
LEU( 19 A-106 )
VAL( 19 A-105 )
GLN( 19 A-104 )
ALA( 19 A-103 )
CYS( 19 A-102 )
GLU( 19 A-101 )
LYS( 19 A-100 )
TYR( 19 A -99 )
TYR( 19 A -98 )
LYS( 19 A -97 )
ALA( 19 A -96 )
ALA( 19 A -95 )
GLU( 19 A -94 )
GLU( 19 A -93 )
ALA( 19 A -92 )
ILE( 19 A -91 )
LYS( 19 A -90 )
LEU( 19 A -89 )
LEU( 19 A -88 )
VAL( 19 A -87 )
ILE( 19 A -86 )
GLU( 19 A -85 )
ASN( 19 A -84 )
ASN( 19 A -83 )
LEU( 19 A -82 )
LYS( 19 A -81 )
GLU( 19 A -80 )
ILE( 19 A -79 )
THR( 19 A -78 )
ASN( 19 A -77 )
ASN( 19 A -76 )
VAL( 19 A -75 )
LYS( 19 A -74 )
ASN( 19 A -73 )
LYS( 19 A -72 )
GLY( 19 A -71 )
ARG( 19 A -70 )
TRP( 19 A -69 )
LYS( 19 A -68 )
SER( 19 A -67 )
GLU( 19 A -66 )
ASN( 19 A -65 )
LEU( 19 A -64 )
PHE( 19 A -63 )
LYS( 19 A -62 )
ALA( 19 A -61 )
SER( 19 A -60 )
LYS( 19 A -59 )
LEU( 19 A -58 )
LEU( 19 A -57 )
ARG( 19 A -56 )
SER( 19 A -55 )
ASN( 19 A -54 )
ASN( 19 A -53 )
THR( 19 A -52 )
GLU( 19 A -51 )
ILE( 19 A -50 )
PRO( 19 A -49 )
ILE( 19 A -48 )
LEU( 19 A -47 )
TRP( 19 A -46 )
LYS( 19 A -45 )
SER( 19 A -44 )
ALA( 19 A -43 )
TRP( 19 A -42 )
THR( 19 A -41 )
LEU( 19 A -40 )
HIS( 19 A -39 )
VAL( 19 A -38 )
GLU( 19 A -37 )
GLY( 19 A -36 )
PHE( 19 A -35 )
HIS( 19 A -34 )
GLU( 19 A -33 )
LEU( 19 A -32 )
SER( 19 A -31 )
LEU( 19 A -30 )
ASN( 19 A -29 )
GLU( 19 A -28 )
LYS( 19 A -27 )
GLU( 19 A -26 )
VAL( 19 A -25 )
LYS( 19 A -24 )
LYS( 19 A -23 )
LEU( 19 A -22 )
LYS( 19 A -21 )
GLU( 19 A -20 )
ASP( 19 A -19 )
VAL( 19 A -18 )
ARG( 19 A -17 )
LYS( 19 A -16 )
LEU( 19 A -15 )
VAL( 19 A -14 )
ILE( 19 A -13 )
PHE( 19 A -12 )
ALA( 19 A -11 )
VAL( 19 A -10 )
ASN( 19 A -9 )
SER( 19 A -8 )
LEU( 19 A -7 )
GLU( 19 A -6 )
HIS( 19 A -5 )
HIS( 19 A -4 )
HIS( 19 A -3 )
HIS( 19 A -2 )
HIS( 19 A -1 )
HIS( 19 A 0 )
MET( 20 A-128 )
SER( 20 A-127 )
ILE( 20 A-126 )
SER( 20 A-125 )
THR( 20 A-124 )
SER( 20 A-123 )
ALA( 20 A-122 )
GLU( 20 A-121 )
VAL( 20 A-120 )
TYR( 20 A-119 )
TYR( 20 A-118 )
GLU( 20 A-117 )
GLU( 20 A-116 )
ALA( 20 A-115 )
GLU( 20 A-114 )
GLU( 20 A-113 )
PHE( 20 A-112 )
LEU( 20 A-111 )
SER( 20 A-110 )
LYS( 20 A-109 )
GLY( 20 A-108 )
ASP( 20 A-107 )
LEU( 20 A-106 )
VAL( 20 A-105 )
GLN( 20 A-104 )
ALA( 20 A-103 )
CYS( 20 A-102 )
GLU( 20 A-101 )
LYS( 20 A-100 )
TYR( 20 A -99 )
TYR( 20 A -98 )
LYS( 20 A -97 )
ALA( 20 A -96 )
ALA( 20 A -95 )
GLU( 20 A -94 )
GLU( 20 A -93 )
ALA( 20 A -92 )
ILE( 20 A -91 )
LYS( 20 A -90 )
LEU( 20 A -89 )
LEU( 20 A -88 )
VAL( 20 A -87 )
ILE( 20 A -86 )
GLU( 20 A -85 )
ASN( 20 A -84 )
ASN( 20 A -83 )
LEU( 20 A -82 )
LYS( 20 A -81 )
GLU( 20 A -80 )
ILE( 20 A -79 )
THR( 20 A -78 )
ASN( 20 A -77 )
ASN( 20 A -76 )
VAL( 20 A -75 )
LYS( 20 A -74 )
ASN( 20 A -73 )
LYS( 20 A -72 )
GLY( 20 A -71 )
ARG( 20 A -70 )
TRP( 20 A -69 )
LYS( 20 A -68 )
SER( 20 A -67 )
GLU( 20 A -66 )
ASN( 20 A -65 )
LEU( 20 A -64 )
PHE( 20 A -63 )
LYS( 20 A -62 )
ALA( 20 A -61 )
SER( 20 A -60 )
LYS( 20 A -59 )
LEU( 20 A -58 )
LEU( 20 A -57 )
ARG( 20 A -56 )
SER( 20 A -55 )
ASN( 20 A -54 )
ASN( 20 A -53 )
THR( 20 A -52 )
GLU( 20 A -51 )
ILE( 20 A -50 )
PRO( 20 A -49 )
ILE( 20 A -48 )
LEU( 20 A -47 )
TRP( 20 A -46 )
LYS( 20 A -45 )
SER( 20 A -44 )
ALA( 20 A -43 )
TRP( 20 A -42 )
THR( 20 A -41 )
LEU( 20 A -40 )
HIS( 20 A -39 )
VAL( 20 A -38 )
GLU( 20 A -37 )
GLY( 20 A -36 )
PHE( 20 A -35 )
HIS( 20 A -34 )
GLU( 20 A -33 )
LEU( 20 A -32 )
SER( 20 A -31 )
LEU( 20 A -30 )
ASN( 20 A -29 )
GLU( 20 A -28 )
LYS( 20 A -27 )
GLU( 20 A -26 )
VAL( 20 A -25 )
LYS( 20 A -24 )
LYS( 20 A -23 )
LEU( 20 A -22 )
LYS( 20 A -21 )
GLU( 20 A -20 )
ASP( 20 A -19 )
VAL( 20 A -18 )
ARG( 20 A -17 )
LYS( 20 A -16 )
LEU( 20 A -15 )
VAL( 20 A -14 )
ILE( 20 A -13 )
PHE( 20 A -12 )
ALA( 20 A -11 )
VAL( 20 A -10 )
ASN( 20 A -9 )
SER( 20 A -8 )
LEU( 20 A -7 )
GLU( 20 A -6 )
HIS( 20 A -5 )
HIS( 20 A -4 )
HIS( 20 A -3 )
HIS( 20 A -2 )
HIS( 20 A -1 )
HIS( 20 A 0 )
PDB Chain_ID: A
1 15
SEQRES: MET SER ILE SER THR SER ALA GLU VAL TYR TYR GLU GLU ALA GLU
COORDS: ... ... ... ... ... ... ... ... ... ... ... ... ... ... ...
16 30
SEQRES: GLU PHE LEU SER LYS GLY ASP LEU VAL GLN ALA CYS GLU LYS TYR
COORDS: ... ... ... ... ... ... ... ... ... ... ... ... ... ... ...
31 45
SEQRES: TYR LYS ALA ALA GLU GLU ALA ILE LYS LEU LEU VAL ILE GLU ASN
COORDS: ... ... ... ... ... ... ... ... ... ... ... ... ... ... ...
46 60
SEQRES: ASN LEU LYS GLU ILE THR ASN ASN VAL LYS ASN LYS GLY ARG TRP
COORDS: ... ... ... ... ... ... ... ... ... ... ... ... ... ... ...
61 75
SEQRES: LYS SER GLU ASN LEU PHE LYS ALA SER LYS LEU LEU ARG SER ASN
COORDS: ... ... ... ... ... ... ... ... ... ... ... ... ... ... ...
76 90
SEQRES: ASN THR GLU ILE PRO ILE LEU TRP LYS SER ALA TRP THR LEU HIS
COORDS: ... ... ... ... ... ... ... ... ... ... ... ... ... ... ...
91 105
SEQRES: VAL GLU GLY PHE HIS GLU LEU SER LEU ASN GLU LYS GLU VAL LYS
COORDS: ... ... ... ... ... ... ... ... ... ... ... ... ... ... ...
106 120
SEQRES: LYS LEU LYS GLU ASP VAL ARG LYS LEU VAL ILE PHE ALA VAL ASN
COORDS: ... ... ... ... ... ... ... ... ... ... ... ... ... ... ...
121 135
SEQRES: SER LEU GLU HIS HIS HIS HIS HIS HIS MET SER ILE SER THR SER
COORDS: ... ... ... ... ... ... ... ... ... MET SER ILE SER THR SER
1 6
136 150
SEQRES: ALA GLU VAL TYR TYR GLU GLU ALA GLU GLU PHE LEU SER LYS GLY
COORDS: ALA GLU VAL TYR TYR GLU GLU ALA GLU GLU PHE LEU SER LYS GLY
7 21
151 165
SEQRES: ASP LEU VAL GLN ALA CYS GLU LYS TYR TYR LYS ALA ALA GLU GLU
COORDS: ASP LEU VAL GLN ALA CYS GLU LYS TYR TYR LYS ALA ALA GLU GLU
22 36
166 180
SEQRES: ALA ILE LYS LEU LEU VAL ILE GLU ASN ASN LEU LYS GLU ILE THR
COORDS: ALA ILE LYS LEU LEU VAL ILE GLU ASN ASN LEU LYS GLU ILE THR
37 51
181 195
SEQRES: ASN ASN VAL LYS ASN LYS GLY ARG TRP LYS SER GLU ASN LEU PHE
COORDS: ASN ASN VAL LYS ASN LYS GLY ARG TRP LYS SER GLU ASN LEU PHE
52 66
196 210
SEQRES: LYS ALA SER LYS LEU LEU ARG SER ASN ASN THR GLU ILE PRO ILE
COORDS: LYS ALA SER LYS LEU LEU ARG SER ASN ASN THR GLU ILE PRO ILE
67 81
211 225
SEQRES: LEU TRP LYS SER ALA TRP THR LEU HIS VAL GLU GLY PHE HIS GLU
COORDS: LEU TRP LYS SER ALA TRP THR LEU HIS VAL GLU GLY PHE HIS GLU
82 96
226 240
SEQRES: LEU SER LEU ASN GLU LYS GLU VAL LYS LYS LEU LYS GLU ASP VAL
COORDS: LEU SER LEU ASN GLU LYS GLU VAL LYS LYS LEU LYS GLU ASP VAL
97 111
241 255
SEQRES: ARG LYS LEU VAL ILE PHE ALA VAL ASN SER LEU GLU HIS HIS HIS
COORDS: ARG LYS LEU VAL ILE PHE ALA VAL ASN SER LEU GLU HIS HIS HIS
112 126
256 258
SEQRES: HIS HIS HIS
COORDS: HIS HIS HIS
127 129
==> The following residues have missing atoms:
RES MOD#C SEQ ATOMS
GLU( 1 A 8) HE2
GLU( 1 A 12) HE2
GLU( 1 A 13) HE2
GLU( 1 A 15) HE2
GLU( 1 A 16) HE2
ASP( 1 A 22) HD2
GLU( 1 A 28) HE2
GLU( 1 A 35) HE2
GLU( 1 A 36) HE2
GLU( 1 A 44) HE2
GLU( 1 A 49) HE2
GLU( 1 A 63) HE2
GLU( 1 A 78) HE2
GLU( 1 A 92) HE2
GLU( 1 A 96) HE2
GLU( 1 A 101) HE2
GLU( 1 A 103) HE2
GLU( 1 A 109) HE2
ASP( 1 A 110) HD2
GLU( 1 A 123) HE2
GLU( 2 A 8) HE2
GLU( 2 A 12) HE2
GLU( 2 A 13) HE2
GLU( 2 A 15) HE2
GLU( 2 A 16) HE2
ASP( 2 A 22) HD2
GLU( 2 A 28) HE2
GLU( 2 A 35) HE2
GLU( 2 A 36) HE2
GLU( 2 A 44) HE2
GLU( 2 A 49) HE2
GLU( 2 A 63) HE2
GLU( 2 A 78) HE2
GLU( 2 A 92) HE2
GLU( 2 A 96) HE2
GLU( 2 A 101) HE2
GLU( 2 A 103) HE2
GLU( 2 A 109) HE2
ASP( 2 A 110) HD2
GLU( 2 A 123) HE2
GLU( 3 A 8) HE2
GLU( 3 A 12) HE2
GLU( 3 A 13) HE2
GLU( 3 A 15) HE2
GLU( 3 A 16) HE2
ASP( 3 A 22) HD2
GLU( 3 A 28) HE2
GLU( 3 A 35) HE2
GLU( 3 A 36) HE2
GLU( 3 A 44) HE2
GLU( 3 A 49) HE2
GLU( 3 A 63) HE2
GLU( 3 A 78) HE2
GLU( 3 A 92) HE2
GLU( 3 A 96) HE2
GLU( 3 A 101) HE2
GLU( 3 A 103) HE2
GLU( 3 A 109) HE2
ASP( 3 A 110) HD2
GLU( 3 A 123) HE2
GLU( 4 A 8) HE2
GLU( 4 A 12) HE2
GLU( 4 A 13) HE2
GLU( 4 A 15) HE2
GLU( 4 A 16) HE2
ASP( 4 A 22) HD2
GLU( 4 A 28) HE2
GLU( 4 A 35) HE2
GLU( 4 A 36) HE2
GLU( 4 A 44) HE2
GLU( 4 A 49) HE2
GLU( 4 A 63) HE2
GLU( 4 A 78) HE2
GLU( 4 A 92) HE2
GLU( 4 A 96) HE2
GLU( 4 A 101) HE2
GLU( 4 A 103) HE2
GLU( 4 A 109) HE2
ASP( 4 A 110) HD2
GLU( 4 A 123) HE2
GLU( 5 A 8) HE2
GLU( 5 A 12) HE2
GLU( 5 A 13) HE2
GLU( 5 A 15) HE2
GLU( 5 A 16) HE2
ASP( 5 A 22) HD2
GLU( 5 A 28) HE2
GLU( 5 A 35) HE2
GLU( 5 A 36) HE2
GLU( 5 A 44) HE2
GLU( 5 A 49) HE2
GLU( 5 A 63) HE2
GLU( 5 A 78) HE2
GLU( 5 A 92) HE2
GLU( 5 A 96) HE2
GLU( 5 A 101) HE2
GLU( 5 A 103) HE2
GLU( 5 A 109) HE2
ASP( 5 A 110) HD2
GLU( 5 A 123) HE2
GLU( 6 A 8) HE2
GLU( 6 A 12) HE2
GLU( 6 A 13) HE2
GLU( 6 A 15) HE2
GLU( 6 A 16) HE2
ASP( 6 A 22) HD2
GLU( 6 A 28) HE2
GLU( 6 A 35) HE2
GLU( 6 A 36) HE2
GLU( 6 A 44) HE2
GLU( 6 A 49) HE2
GLU( 6 A 63) HE2
GLU( 6 A 78) HE2
GLU( 6 A 92) HE2
GLU( 6 A 96) HE2
GLU( 6 A 101) HE2
GLU( 6 A 103) HE2
GLU( 6 A 109) HE2
ASP( 6 A 110) HD2
GLU( 6 A 123) HE2
GLU( 7 A 8) HE2
GLU( 7 A 12) HE2
GLU( 7 A 13) HE2
GLU( 7 A 15) HE2
GLU( 7 A 16) HE2
ASP( 7 A 22) HD2
GLU( 7 A 28) HE2
GLU( 7 A 35) HE2
GLU( 7 A 36) HE2
GLU( 7 A 44) HE2
GLU( 7 A 49) HE2
GLU( 7 A 63) HE2
GLU( 7 A 78) HE2
GLU( 7 A 92) HE2
GLU( 7 A 96) HE2
GLU( 7 A 101) HE2
GLU( 7 A 103) HE2
GLU( 7 A 109) HE2
ASP( 7 A 110) HD2
GLU( 7 A 123) HE2
GLU( 8 A 8) HE2
GLU( 8 A 12) HE2
GLU( 8 A 13) HE2
GLU( 8 A 15) HE2
GLU( 8 A 16) HE2
ASP( 8 A 22) HD2
GLU( 8 A 28) HE2
GLU( 8 A 35) HE2
GLU( 8 A 36) HE2
GLU( 8 A 44) HE2
GLU( 8 A 49) HE2
GLU( 8 A 63) HE2
GLU( 8 A 78) HE2
GLU( 8 A 92) HE2
GLU( 8 A 96) HE2
GLU( 8 A 101) HE2
GLU( 8 A 103) HE2
GLU( 8 A 109) HE2
ASP( 8 A 110) HD2
GLU( 8 A 123) HE2
GLU( 9 A 8) HE2
GLU( 9 A 12) HE2
GLU( 9 A 13) HE2
GLU( 9 A 15) HE2
GLU( 9 A 16) HE2
ASP( 9 A 22) HD2
GLU( 9 A 28) HE2
GLU( 9 A 35) HE2
GLU( 9 A 36) HE2
GLU( 9 A 44) HE2
GLU( 9 A 49) HE2
GLU( 9 A 63) HE2
GLU( 9 A 78) HE2
GLU( 9 A 92) HE2
GLU( 9 A 96) HE2
GLU( 9 A 101) HE2
GLU( 9 A 103) HE2
GLU( 9 A 109) HE2
ASP( 9 A 110) HD2
GLU( 9 A 123) HE2
GLU( 10 A 8) HE2
GLU( 10 A 12) HE2
GLU( 10 A 13) HE2
GLU( 10 A 15) HE2
GLU( 10 A 16) HE2
ASP( 10 A 22) HD2
GLU( 10 A 28) HE2
GLU( 10 A 35) HE2
GLU( 10 A 36) HE2
GLU( 10 A 44) HE2
GLU( 10 A 49) HE2
GLU( 10 A 63) HE2
GLU( 10 A 78) HE2
GLU( 10 A 92) HE2
GLU( 10 A 96) HE2
GLU( 10 A 101) HE2
GLU( 10 A 103) HE2
GLU( 10 A 109) HE2
ASP( 10 A 110) HD2
GLU( 10 A 123) HE2
GLU( 11 A 8) HE2
GLU( 11 A 12) HE2
GLU( 11 A 13) HE2
GLU( 11 A 15) HE2
GLU( 11 A 16) HE2
ASP( 11 A 22) HD2
GLU( 11 A 28) HE2
GLU( 11 A 35) HE2
GLU( 11 A 36) HE2
GLU( 11 A 44) HE2
GLU( 11 A 49) HE2
GLU( 11 A 63) HE2
GLU( 11 A 78) HE2
GLU( 11 A 92) HE2
GLU( 11 A 96) HE2
GLU( 11 A 101) HE2
GLU( 11 A 103) HE2
GLU( 11 A 109) HE2
ASP( 11 A 110) HD2
GLU( 11 A 123) HE2
GLU( 12 A 8) HE2
GLU( 12 A 12) HE2
GLU( 12 A 13) HE2
GLU( 12 A 15) HE2
GLU( 12 A 16) HE2
ASP( 12 A 22) HD2
GLU( 12 A 28) HE2
GLU( 12 A 35) HE2
GLU( 12 A 36) HE2
GLU( 12 A 44) HE2
GLU( 12 A 49) HE2
GLU( 12 A 63) HE2
GLU( 12 A 78) HE2
GLU( 12 A 92) HE2
GLU( 12 A 96) HE2
GLU( 12 A 101) HE2
GLU( 12 A 103) HE2
GLU( 12 A 109) HE2
ASP( 12 A 110) HD2
GLU( 12 A 123) HE2
GLU( 13 A 8) HE2
GLU( 13 A 12) HE2
GLU( 13 A 13) HE2
GLU( 13 A 15) HE2
GLU( 13 A 16) HE2
ASP( 13 A 22) HD2
GLU( 13 A 28) HE2
GLU( 13 A 35) HE2
GLU( 13 A 36) HE2
GLU( 13 A 44) HE2
GLU( 13 A 49) HE2
GLU( 13 A 63) HE2
GLU( 13 A 78) HE2
GLU( 13 A 92) HE2
GLU( 13 A 96) HE2
GLU( 13 A 101) HE2
GLU( 13 A 103) HE2
GLU( 13 A 109) HE2
ASP( 13 A 110) HD2
GLU( 13 A 123) HE2
GLU( 14 A 8) HE2
GLU( 14 A 12) HE2
GLU( 14 A 13) HE2
GLU( 14 A 15) HE2
GLU( 14 A 16) HE2
ASP( 14 A 22) HD2
GLU( 14 A 28) HE2
GLU( 14 A 35) HE2
GLU( 14 A 36) HE2
GLU( 14 A 44) HE2
GLU( 14 A 49) HE2
GLU( 14 A 63) HE2
GLU( 14 A 78) HE2
GLU( 14 A 92) HE2
GLU( 14 A 96) HE2
GLU( 14 A 101) HE2
GLU( 14 A 103) HE2
GLU( 14 A 109) HE2
ASP( 14 A 110) HD2
GLU( 14 A 123) HE2
GLU( 15 A 8) HE2
GLU( 15 A 12) HE2
GLU( 15 A 13) HE2
GLU( 15 A 15) HE2
GLU( 15 A 16) HE2
ASP( 15 A 22) HD2
GLU( 15 A 28) HE2
GLU( 15 A 35) HE2
GLU( 15 A 36) HE2
GLU( 15 A 44) HE2
GLU( 15 A 49) HE2
GLU( 15 A 63) HE2
GLU( 15 A 78) HE2
GLU( 15 A 92) HE2
GLU( 15 A 96) HE2
GLU( 15 A 101) HE2
GLU( 15 A 103) HE2
GLU( 15 A 109) HE2
ASP( 15 A 110) HD2
GLU( 15 A 123) HE2
GLU( 16 A 8) HE2
GLU( 16 A 12) HE2
GLU( 16 A 13) HE2
GLU( 16 A 15) HE2
GLU( 16 A 16) HE2
ASP( 16 A 22) HD2
GLU( 16 A 28) HE2
GLU( 16 A 35) HE2
GLU( 16 A 36) HE2
GLU( 16 A 44) HE2
GLU( 16 A 49) HE2
GLU( 16 A 63) HE2
GLU( 16 A 78) HE2
GLU( 16 A 92) HE2
GLU( 16 A 96) HE2
GLU( 16 A 101) HE2
GLU( 16 A 103) HE2
GLU( 16 A 109) HE2
ASP( 16 A 110) HD2
GLU( 16 A 123) HE2
GLU( 17 A 8) HE2
GLU( 17 A 12) HE2
GLU( 17 A 13) HE2
GLU( 17 A 15) HE2
GLU( 17 A 16) HE2
ASP( 17 A 22) HD2
GLU( 17 A 28) HE2
GLU( 17 A 35) HE2
GLU( 17 A 36) HE2
GLU( 17 A 44) HE2
GLU( 17 A 49) HE2
GLU( 17 A 63) HE2
GLU( 17 A 78) HE2
GLU( 17 A 92) HE2
GLU( 17 A 96) HE2
GLU( 17 A 101) HE2
GLU( 17 A 103) HE2
GLU( 17 A 109) HE2
ASP( 17 A 110) HD2
GLU( 17 A 123) HE2
GLU( 18 A 8) HE2
GLU( 18 A 12) HE2
GLU( 18 A 13) HE2
GLU( 18 A 15) HE2
GLU( 18 A 16) HE2
ASP( 18 A 22) HD2
GLU( 18 A 28) HE2
GLU( 18 A 35) HE2
GLU( 18 A 36) HE2
GLU( 18 A 44) HE2
GLU( 18 A 49) HE2
GLU( 18 A 63) HE2
GLU( 18 A 78) HE2
GLU( 18 A 92) HE2
GLU( 18 A 96) HE2
GLU( 18 A 101) HE2
GLU( 18 A 103) HE2
GLU( 18 A 109) HE2
ASP( 18 A 110) HD2
GLU( 18 A 123) HE2
GLU( 19 A 8) HE2
GLU( 19 A 12) HE2
GLU( 19 A 13) HE2
GLU( 19 A 15) HE2
GLU( 19 A 16) HE2
ASP( 19 A 22) HD2
GLU( 19 A 28) HE2
GLU( 19 A 35) HE2
GLU( 19 A 36) HE2
GLU( 19 A 44) HE2
GLU( 19 A 49) HE2
GLU( 19 A 63) HE2
GLU( 19 A 78) HE2
GLU( 19 A 92) HE2
GLU( 19 A 96) HE2
GLU( 19 A 101) HE2
GLU( 19 A 103) HE2
GLU( 19 A 109) HE2
ASP( 19 A 110) HD2
GLU( 19 A 123) HE2
GLU( 20 A 8) HE2
GLU( 20 A 12) HE2
GLU( 20 A 13) HE2
GLU( 20 A 15) HE2
GLU( 20 A 16) HE2
ASP( 20 A 22) HD2
GLU( 20 A 28) HE2
GLU( 20 A 35) HE2
GLU( 20 A 36) HE2
GLU( 20 A 44) HE2
GLU( 20 A 49) HE2
GLU( 20 A 63) HE2
GLU( 20 A 78) HE2
GLU( 20 A 92) HE2
GLU( 20 A 96) HE2
GLU( 20 A 101) HE2
GLU( 20 A 103) HE2
GLU( 20 A 109) HE2
ASP( 20 A 110) HD2
GLU( 20 A 123) HE2
==> The following residues have extra atoms:
RES MOD#C SEQ ATOMS
HIS( 1 A 129) O2
HIS( 2 A 129) O2
HIS( 3 A 129) O2
HIS( 4 A 129) O2
HIS( 5 A 129) O2
HIS( 6 A 129) O2
HIS( 7 A 129) O2
HIS( 8 A 129) O2
HIS( 9 A 129) O2
HIS( 10 A 129) O2
HIS( 11 A 129) O2
HIS( 12 A 129) O2
HIS( 13 A 129) O2
HIS( 14 A 129) O2
HIS( 15 A 129) O2
HIS( 16 A 129) O2
HIS( 17 A 129) O2
HIS( 18 A 129) O2
HIS( 19 A 129) O2
HIS( 20 A 129) O2
CHECK TERMINAL ATOMS
--------------------
Terminal atom(s) showed in middle of sequence will be deleted:
1H MET 1 A
2H MET 1 A
3H MET 1 A
1H MET 1 A
2H MET 1 A
3H MET 1 A
1H MET 1 A
2H MET 1 A
3H MET 1 A
1H MET 1 A
2H MET 1 A
3H MET 1 A
1H MET 1 A
2H MET 1 A
3H MET 1 A
1H MET 1 A
2H MET 1 A
3H MET 1 A
1H MET 1 A
2H MET 1 A
3H MET 1 A
1H MET 1 A
2H MET 1 A
3H MET 1 A
1H MET 1 A
2H MET 1 A
3H MET 1 A
1H MET 1 A
2H MET 1 A
3H MET 1 A
1H MET 1 A
2H MET 1 A
3H MET 1 A
1H MET 1 A
2H MET 1 A
3H MET 1 A
1H MET 1 A
2H MET 1 A
3H MET 1 A
1H MET 1 A
2H MET 1 A
3H MET 1 A
1H MET 1 A
2H MET 1 A
3H MET 1 A
1H MET 1 A
2H MET 1 A
3H MET 1 A
1H MET 1 A
2H MET 1 A
3H MET 1 A
1H MET 1 A
2H MET 1 A
3H MET 1 A
1H MET 1 A
2H MET 1 A
3H MET 1 A
1H MET 1 A
2H MET 1 A
3H MET 1 A




























