Detailed results of SR478_NMR_em_bcr3 by PSVS
Output from PDBStat
Constraints analysis
table of NOE constraints
# -------------- SUMMARY OF RESTRAINTS ---------------
# TOTAL NUMBER OF NOE RESTRAINTS : 870
# INTRA-RESIDUE RESTRAINTS (I=J) : 188
# SEQUENTIAL RESTRAINTS (I-J)=1 : 232
# BACKBONE-BACKBONE : 62
# BACKBONE-SIDE CHAIN : 43
# SIDE CHAIN-SIDE CHAIN : 127
# MEDIUM RANGE RESTRAINTS 1<(I-J)<5 : 271
# BACKBONE-BACKBONE : 75
# BACKBONE-SIDE CHAIN : 74
# SIDE CHAIN-SIDE CHAIN : 122
# LONG RANGE RESTRAINTS (I-J)>=5 : 179
# TOTAL HYDROGEN BOND RESTRAINTS : 0
# LONG RANGE H-BOND RESTR. (I-J)>=5 : 0
# DISULFIDE RESTRAINTS : 0
# INTRA-CHAIN RESTRAINTS : 870
# INTER-CHAIN RESTRAINTS : 0
# AMBIGUOUS RESTRAINTS : 0
# -----------------------------------------------------
# -----------------------------------------------------
# -----------------------------------------------------
# RES # INTRA INTER seq med lng InterChain
MET 1 0 0.0 0.0 0.0 0.0 0.0
SER 2 0 1.5 1.5 0.0 0.0 0.0
GLU 3 0 4.5 4.5 0.0 0.0 0.0
LEU 4 9 11.5 3.0 1.0 7.5 0.0
PHE 5 0 0.0 0.0 0.0 0.0 0.0
SER 6 0 1.5 0.5 1.0 0.0 0.0
VAL 7 6 6.5 1.0 2.0 3.5 0.0
PRO 8 0 5.0 1.0 4.0 0.0 0.0
TYR 9 1 9.5 3.5 1.0 5.0 0.0
PHE 10 1 15.5 3.5 2.0 10.0 0.0
ILE 11 5 19.0 2.5 8.0 8.5 0.0
GLU 12 3 5.5 3.5 2.0 0.0 0.0
ASN 13 0 6.0 2.5 2.5 1.0 0.0
LEU 14 5 22.0 2.0 7.0 13.0 0.0
LYS 15 1 6.5 2.0 4.5 0.0 0.0
GLN 16 1 7.5 2.5 2.5 2.5 0.0
HIS 17 0 7.0 2.0 3.5 1.5 0.0
ILE 18 5 16.0 2.5 6.0 7.5 0.0
GLU 19 2 4.5 2.5 1.5 0.5 0.0
MET 20 1 5.5 1.5 4.0 0.0 0.0
ASN 21 1 6.5 3.0 2.0 1.5 0.0
GLN 22 3 6.5 4.5 1.0 1.0 0.0
SER 23 0 4.0 3.5 0.5 0.0 0.0
GLU 24 3 9.0 3.0 2.0 4.0 0.0
ASP 25 0 6.5 3.0 2.0 1.5 0.0
LYS 26 7 9.5 4.0 2.0 3.5 0.0
ILE 27 6 15.0 4.0 3.0 8.0 0.0
HIS 28 0 3.0 1.5 1.5 0.0 0.0
ALA 29 0 14.5 1.5 7.0 6.0 0.0
MET 30 0 11.5 2.0 4.0 5.5 0.0
ASN 31 0 4.0 1.5 2.5 0.0 0.0
SER 32 0 3.0 1.5 1.0 0.5 0.0
TYR 33 4 19.5 3.0 7.0 9.5 0.0
TYR 34 4 12.5 2.5 4.5 5.5 0.0
ARG 35 1 1.5 1.0 0.5 0.0 0.0
SER 36 0 2.5 1.5 1.0 0.0 0.0
VAL 37 5 21.5 5.0 8.0 8.5 0.0
VAL 38 1 15.0 5.5 7.0 2.5 0.0
SER 39 1 6.5 3.5 3.0 0.0 0.0
THR 40 2 14.0 6.0 8.0 0.0 0.0
LEU 41 4 14.5 5.0 7.0 2.5 0.0
VAL 42 4 14.0 4.5 6.5 3.0 0.0
GLN 43 1 7.5 4.5 3.0 0.0 0.0
ASP 44 1 5.5 1.5 4.0 0.0 0.0
GLN 45 2 5.5 3.5 1.0 1.0 0.0
LEU 46 10 8.0 5.5 2.0 0.5 0.0
THR 47 3 12.0 5.5 5.5 1.0 0.0
LYS 48 2 17.5 6.0 11.5 0.0 0.0
ASN 49 3 9.0 5.0 4.0 0.0 0.0
ALA 50 1 10.5 4.5 6.0 0.0 0.0
VAL 51 6 18.5 5.0 13.0 0.5 0.0
VAL 52 7 17.0 5.5 7.5 4.0 0.0
LEU 53 8 13.5 4.5 9.0 0.0 0.0
LYS 54 6 10.5 2.5 8.0 0.0 0.0
ARG 55 0 5.5 1.5 3.0 1.0 0.0
ILE 56 2 16.0 3.5 10.0 2.5 0.0
GLN 57 0 7.5 3.0 4.5 0.0 0.0
HIS 58 0 2.5 1.0 1.5 0.0 0.0
LEU 59 4 17.5 3.5 4.5 9.5 0.0
ASP 60 0 7.5 4.0 2.5 1.0 0.0
GLU 61 3 11.5 2.5 2.5 6.5 0.0
ALA 62 0 14.0 2.5 2.5 9.0 0.0
TYR 63 3 13.0 4.0 4.5 4.5 0.0
ASN 64 0 5.0 2.5 2.0 0.5 0.0
LYS 65 0 4.0 1.0 3.0 0.0 0.0
VAL 66 6 11.0 3.0 2.5 5.5 0.0
LYS 67 3 14.0 4.5 5.5 4.0 0.0
ARG 68 5 8.5 4.5 3.5 0.5 0.0
GLY 69 1 7.0 4.5 1.5 1.0 0.0
GLU 70 3 10.0 3.5 4.5 2.0 0.0
SER 71 1 4.5 3.0 0.5 1.0 0.0
LYS 72 6 4.5 3.5 1.0 0.0 0.0
LEU 73 10 6.5 5.0 1.5 0.0 0.0
GLU 74 5 5.5 4.5 1.0 0.0 0.0
HIS 75 0 2.0 1.0 1.0 0.0 0.0
HIS 76 0 0.0 0.0 0.0 0.0 0.0
HIS 77 0 0.0 0.0 0.0 0.0 0.0
HIS 78 0 0.0 0.0 0.0 0.0 0.0
HIS 79 0 0.0 0.0 0.0 0.0 0.0
HIS 80 0 0.0 0.0 0.0 0.0 0.0
MET 1 0 0.0 0.0 0.0 0.0 0.0
SER 2 0 0.0 0.0 0.0 0.0 0.0
GLU 3 0 0.0 0.0 0.0 0.0 0.0
LEU 4 0 0.0 0.0 0.0 0.0 0.0
PHE 5 0 0.0 0.0 0.0 0.0 0.0
SER 6 0 0.0 0.0 0.0 0.0 0.0
VAL 7 0 0.0 0.0 0.0 0.0 0.0
PRO 8 0 0.0 0.0 0.0 0.0 0.0
TYR 9 0 0.0 0.0 0.0 0.0 0.0
PHE 10 0 0.0 0.0 0.0 0.0 0.0
ILE 11 0 0.0 0.0 0.0 0.0 0.0
GLU 12 0 0.0 0.0 0.0 0.0 0.0
ASN 13 0 0.0 0.0 0.0 0.0 0.0
LEU 14 0 0.0 0.0 0.0 0.0 0.0
LYS 15 0 0.0 0.0 0.0 0.0 0.0
GLN 16 0 0.0 0.0 0.0 0.0 0.0
HIS 17 0 0.0 0.0 0.0 0.0 0.0
ILE 18 0 0.0 0.0 0.0 0.0 0.0
GLU 19 0 0.0 0.0 0.0 0.0 0.0
MET 20 0 0.0 0.0 0.0 0.0 0.0
ASN 21 0 0.0 0.0 0.0 0.0 0.0
GLN 22 0 0.0 0.0 0.0 0.0 0.0
SER 23 0 0.0 0.0 0.0 0.0 0.0
GLU 24 0 0.0 0.0 0.0 0.0 0.0
ASP 25 0 0.0 0.0 0.0 0.0 0.0
LYS 26 0 0.0 0.0 0.0 0.0 0.0
ILE 27 0 0.0 0.0 0.0 0.0 0.0
HIS 28 0 0.0 0.0 0.0 0.0 0.0
ALA 29 0 0.0 0.0 0.0 0.0 0.0
MET 30 0 0.0 0.0 0.0 0.0 0.0
ASN 31 0 0.0 0.0 0.0 0.0 0.0
SER 32 0 0.0 0.0 0.0 0.0 0.0
TYR 33 0 0.0 0.0 0.0 0.0 0.0
TYR 34 0 0.0 0.0 0.0 0.0 0.0
ARG 35 0 0.0 0.0 0.0 0.0 0.0
SER 36 0 0.0 0.0 0.0 0.0 0.0
VAL 37 0 0.0 0.0 0.0 0.0 0.0
VAL 38 0 0.0 0.0 0.0 0.0 0.0
SER 39 0 0.0 0.0 0.0 0.0 0.0
THR 40 0 0.0 0.0 0.0 0.0 0.0
LEU 41 0 0.0 0.0 0.0 0.0 0.0
VAL 42 0 0.0 0.0 0.0 0.0 0.0
GLN 43 0 0.0 0.0 0.0 0.0 0.0
ASP 44 0 0.0 0.0 0.0 0.0 0.0
GLN 45 0 0.0 0.0 0.0 0.0 0.0
LEU 46 0 0.0 0.0 0.0 0.0 0.0
THR 47 0 0.0 0.0 0.0 0.0 0.0
LYS 48 0 0.0 0.0 0.0 0.0 0.0
ASN 49 0 0.0 0.0 0.0 0.0 0.0
ALA 50 0 0.0 0.0 0.0 0.0 0.0
VAL 51 0 0.0 0.0 0.0 0.0 0.0
VAL 52 0 0.0 0.0 0.0 0.0 0.0
LEU 53 0 0.0 0.0 0.0 0.0 0.0
LYS 54 0 0.0 0.0 0.0 0.0 0.0
ARG 55 0 0.0 0.0 0.0 0.0 0.0
ILE 56 0 0.0 0.0 0.0 0.0 0.0
GLN 57 0 0.0 0.0 0.0 0.0 0.0
HIS 58 0 0.0 0.0 0.0 0.0 0.0
LEU 59 0 0.0 0.0 0.0 0.0 0.0
ASP 60 0 0.0 0.0 0.0 0.0 0.0
GLU 61 0 0.0 0.0 0.0 0.0 0.0
ALA 62 0 0.0 0.0 0.0 0.0 0.0
TYR 63 0 0.0 0.0 0.0 0.0 0.0
ASN 64 0 0.0 0.0 0.0 0.0 0.0
LYS 65 0 0.0 0.0 0.0 0.0 0.0
VAL 66 0 0.0 0.0 0.0 0.0 0.0
LYS 67 0 0.0 0.0 0.0 0.0 0.0
ARG 68 0 0.0 0.0 0.0 0.0 0.0
GLY 69 0 0.0 0.0 0.0 0.0 0.0
GLU 70 0 0.0 0.0 0.0 0.0 0.0
SER 71 0 0.0 0.0 0.0 0.0 0.0
LYS 72 0 0.0 0.0 0.0 0.0 0.0
LEU 73 0 0.0 0.0 0.0 0.0 0.0
GLU 74 0 0.0 0.0 0.0 0.0 0.0
HIS 75 0 0.0 0.0 0.0 0.0 0.0
HIS 76 0 0.0 0.0 0.0 0.0 0.0
HIS 77 0 0.0 0.0 0.0 0.0 0.0
HIS 78 0 0.0 0.0 0.0 0.0 0.0
HIS 79 0 0.0 0.0 0.0 0.0 0.0
HIS 80 0 0.0 0.0 0.0 0.0 0.0
# TOTAL 188 682.0 232.0 271.0 179.0 0.0
# TOTAL NUMBER OF RESTRAINTS (CHECKING): 870.0
List of conformationally-resticting NOE constraints
assign ((segid A and resid 7 and name HA )) ((segid A and resid 7 and name HG1# )) 4.09 2.29 0.00
assign ((segid A and resid 37 and name HG1# )) ((segid A and resid 40 and name HB )) 6.00 4.20 0.00
assign ((segid A and resid 37 and name HG2# )) ((segid A and resid 40 and name HB )) 6.00 4.20 0.00
assign ((segid A and resid 37 and name HA )) ((segid A and resid 40 and name HB )) 6.00 4.20 0.00
assign ((segid A and resid 40 and name HB )) ((segid A and resid 41 and name HN )) 6.00 4.20 0.00
assign ((segid A and resid 52 and name HA )) ((segid A and resid 52 and name HG1# )) 3.45 1.65 0.00
assign ((segid A and resid 38 and name HA )) ((segid A and resid 41 and name HG )) 4.15 2.35 0.00
assign ((segid A and resid 11 and name HA )) ((segid A and resid 14 and name HD2# )) 4.20 2.40 0.00
assign ((segid A and resid 11 and name HA )) ((segid A and resid 62 and name HB# )) 6.00 4.20 0.00
assign ((segid A and resid 11 and name HA )) ((segid A and resid 14 and name HD1# )) 4.20 2.40 0.00
assign ((segid A and resid 7 and name HB )) ((segid A and resid 8 and name HA )) 4.84 3.04 0.00
assign ((segid A and resid 8 and name HA )) ((segid A and resid 11 and name HB )) 3.59 1.79 0.00
assign ((segid A and resid 8 and name HA )) ((segid A and resid 11 and name HG2# )) 4.48 2.68 0.00
assign ((segid A and resid 8 and name HA )) ((segid A and resid 11 and name HD1# )) 4.62 2.82 0.00
assign ((segid A and resid 66 and name HA )) ((segid A and resid 66 and name HG2# )) 3.43 1.63 0.00
assign ((segid A and resid 37 and name HA )) ((segid A and resid 37 and name HG1# )) 3.53 1.73 0.00
assign ((segid A and resid 37 and name HA )) ((segid A and resid 41 and name HG )) 6.00 4.20 0.00
assign ((segid A and resid 37 and name HA )) ((segid A and resid 37 and name HG2# )) 3.53 1.73 0.00
assign ((segid A and resid 51 and name HA )) ((segid A and resid 51 and name HG2# )) 3.27 1.47 0.00
assign ((segid A and resid 51 and name HA )) ((segid A and resid 51 and name HG1# )) 3.27 1.47 0.00
assign ((segid A and resid 47 and name HA )) ((segid A and resid 47 and name HG2# )) 3.59 1.79 0.00
assign ((segid A and resid 27 and name HA )) ((segid A and resid 27 and name HD1# )) 3.32 1.52 0.00
assign ((segid A and resid 17 and name HA )) ((segid A and resid 19 and name HN )) 6.00 4.20 0.00
assign ((segid A and resid 17 and name HA )) ((segid A and resid 20 and name HN )) 6.00 4.20 0.00
assign ((segid A and resid 17 and name HA )) ((segid A and resid 25 and name HN )) 6.00 4.20 0.00
assign ((segid A and resid 15 and name HA )) ((segid A and resid 18 and name HG1# )) 3.48 1.68 0.00
assign ((segid A and resid 65 and name HA )) ((segid A and resid 67 and name HN )) 6.00 4.20 0.00
assign ((segid A and resid 18 and name HG2# )) ((segid A and resid 26 and name HA )) 3.55 1.75 0.00
assign ((segid A and resid 18 and name HD1# )) ((segid A and resid 26 and name HA )) 4.87 3.07 0.00
assign ((segid A and resid 26 and name HA )) ((segid A and resid 29 and name HB# )) 3.65 1.85 0.00
assign ((segid A and resid 29 and name HB# )) ((segid A and resid 30 and name HA )) 6.00 4.20 0.00
assign ((segid A and resid 30 and name HA )) ((segid A and resid 66 and name HG2# )) 5.20 3.40 0.00
assign ((segid A and resid 14 and name HA )) ((segid A and resid 18 and name HG1# )) 6.00 4.20 0.00
assign ((segid A and resid 66 and name HB )) ((segid A and resid 67 and name HA )) 6.00 4.20 0.00
assign ((segid A and resid 27 and name HD1# )) ((segid A and resid 67 and name HA )) 4.10 2.30 0.00
assign ((segid A and resid 67 and name HA )) ((segid A and resid 67 and name HG1 )) 3.44 1.64 0.00
assign ((segid A and resid 11 and name HG2# )) ((segid A and resid 14 and name HA )) 6.00 4.20 0.00
assign ((segid A and resid 14 and name HA )) ((segid A and resid 18 and name HD1# )) 6.00 4.20 0.00
assign ((segid A and resid 18 and name HB )) ((segid A and resid 19 and name HA )) 4.76 2.96 0.00
assign ((segid A and resid 53 and name HA )) ((segid A and resid 56 and name HD1# )) 4.44 2.64 0.00
assign ((segid A and resid 72 and name HA )) ((segid A and resid 73 and name HN )) 3.57 1.77 0.00
assign ((segid A and resid 58 and name HA )) ((segid A and resid 61 and name HN )) 4.14 2.34 0.00
assign ((segid A and resid 64 and name HA )) ((segid A and resid 67 and name HN )) 3.84 2.04 0.00
assign ((segid A and resid 49 and name HA )) ((segid A and resid 50 and name HB# )) 6.00 4.20 0.00
assign ((segid A and resid 46 and name HA )) ((segid A and resid 48 and name HN )) 3.66 1.86 0.00
assign ((segid A and resid 46 and name HN )) ((segid A and resid 46 and name HA )) 2.93 1.13 0.00
assign ((segid A and resid 46 and name HA )) ((segid A and resid 52 and name HN )) 5.69 3.89 0.00
assign ((segid A and resid 74 and name HA )) ((segid A and resid 74 and name HG1 )) 4.11 2.31 0.00
assign ((segid A and resid 74 and name HA )) ((segid A and resid 74 and name HG2 )) 4.11 2.31 0.00
assign ((segid A and resid 3 and name HA )) ((segid A and resid 4 and name HA )) 4.61 2.81 0.00
assign ((segid A and resid 73 and name HA )) ((segid A and resid 73 and name HG )) 3.51 1.71 0.00
assign ((segid A and resid 24 and name HA )) ((segid A and resid 25 and name HB1 )) 6.00 4.20 0.00
assign ((segid A and resid 25 and name HB2 )) ((segid A and resid 30 and name HN )) 5.83 4.03 0.00
assign ((segid A and resid 46 and name HB2 )) ((segid A and resid 46 and name HD2# )) 3.28 1.48 0.00
assign ((segid A and resid 54 and name HD1 )) ((segid A and resid 54 and name HE# )) 2.36 0.56 0.00
assign ((segid A and resid 54 and name HD2 )) ((segid A and resid 54 and name HE# )) 2.36 0.56 0.00
assign ((segid A and resid 73 and name HB1 )) ((segid A and resid 73 and name HD1# )) 3.73 1.93 0.00
assign ((segid A and resid 73 and name HB1 )) ((segid A and resid 73 and name HD2# )) 3.73 1.93 0.00
assign ((segid A and resid 73 and name HB2 )) ((segid A and resid 73 and name HD1# )) 3.73 1.93 0.00
assign ((segid A and resid 73 and name HB2 )) ((segid A and resid 73 and name HD2# )) 3.73 1.93 0.00
assign ((segid A and resid 53 and name HD1# )) ((segid A and resid 56 and name HB )) 5.17 3.37 0.00
assign ((segid A and resid 7 and name HG2# )) ((segid A and resid 61 and name HG2 )) 6.00 4.20 0.00
assign ((segid A and resid 17 and name HN )) ((segid A and resid 24 and name HG2 )) 6.00 4.20 0.00
assign ((segid A and resid 16 and name HG1 )) ((segid A and resid 17 and name HN )) 6.00 4.20 0.00
assign ((segid A and resid 16 and name HG1 )) ((segid A and resid 21 and name HN )) 5.17 3.37 0.00
assign ((segid A and resid 16 and name HG2 )) ((segid A and resid 17 and name HN )) 6.00 4.20 0.00
assign ((segid A and resid 26 and name HN )) ((segid A and resid 26 and name HB2 )) 4.06 2.26 0.00
assign ((segid A and resid 72 and name HN )) ((segid A and resid 72 and name HB1 )) 4.02 2.22 0.00
assign ((segid A and resid 66 and name HN )) ((segid A and resid 66 and name HB )) 4.17 2.37 0.00
assign ((segid A and resid 64 and name HN )) ((segid A and resid 66 and name HB )) 6.00 4.20 0.00
assign ((segid A and resid 27 and name HD1# )) ((segid A and resid 67 and name HB1 )) 6.00 4.20 0.00
assign ((segid A and resid 24 and name HB1 )) ((segid A and resid 25 and name HN )) 6.00 4.20 0.00
assign ((segid A and resid 48 and name HN )) ((segid A and resid 51 and name HB )) 6.00 4.20 0.00
assign ((segid A and resid 17 and name HN )) ((segid A and resid 24 and name HB1 )) 6.00 4.20 0.00
assign ((segid A and resid 52 and name HB )) ((segid A and resid 53 and name HN )) 4.56 2.76 0.00
assign ((segid A and resid 70 and name HB2 )) ((segid A and resid 71 and name HN )) 4.13 2.33 0.00
assign ((segid A and resid 67 and name HN )) ((segid A and resid 68 and name HB1 )) 6.00 4.20 0.00
assign ((segid A and resid 22 and name HB2 )) ((segid A and resid 23 and name HN )) 4.16 2.36 0.00
assign ((segid A and resid 46 and name HG )) ((segid A and resid 48 and name HN )) 6.00 4.20 0.00
assign ((segid A and resid 46 and name HN )) ((segid A and resid 46 and name HG )) 5.36 3.56 0.00
assign ((segid A and resid 73 and name HG )) ((segid A and resid 74 and name HN )) 4.46 2.66 0.00
assign ((segid A and resid 73 and name HN )) ((segid A and resid 73 and name HG )) 4.49 2.69 0.00
assign ((segid A and resid 14 and name HD1# )) ((segid A and resid 62 and name HN )) 6.00 4.20 0.00
assign ((segid A and resid 59 and name HG )) ((segid A and resid 60 and name HN )) 6.00 4.20 0.00
assign ((segid A and resid 30 and name HN )) ((segid A and resid 66 and name HG2# )) 3.96 2.16 0.00
assign ((segid A and resid 46 and name HN )) ((segid A and resid 46 and name HD1# )) 4.91 3.11 0.00
assign ((segid A and resid 27 and name HD1# )) ((segid A and resid 67 and name HG1 )) 4.93 3.13 0.00
assign ((segid A and resid 46 and name HB1 )) ((segid A and resid 46 and name HD1# )) 3.28 1.48 0.00
assign ((segid A and resid 46 and name HB2 )) ((segid A and resid 46 and name HD1# )) 3.28 1.48 0.00
assign ((segid A and resid 53 and name HN )) ((segid A and resid 53 and name HD1# )) 4.53 2.73 0.00
assign ((segid A and resid 53 and name HA )) ((segid A and resid 53 and name HD1# )) 4.21 2.41 0.00
assign ((segid A and resid 53 and name HN )) ((segid A and resid 53 and name HD2# )) 4.53 2.73 0.00
assign ((segid A and resid 53 and name HD2# )) ((segid A and resid 56 and name HB )) 5.17 3.37 0.00
assign ((segid A and resid 53 and name HA )) ((segid A and resid 53 and name HD2# )) 4.21 2.41 0.00
assign ((segid A and resid 4 and name HD2# )) ((segid A and resid 10 and name HE# )) 4.98 3.18 0.00
assign ((segid A and resid 4 and name HD2# )) ((segid A and resid 10 and name HD# )) 6.00 4.20 0.00
assign ((segid A and resid 4 and name HD2# )) ((segid A and resid 9 and name HE# )) 4.75 2.95 0.00
assign ((segid A and resid 4 and name HA )) ((segid A and resid 4 and name HD2# )) 4.34 2.54 0.00
assign ((segid A and resid 4 and name HN )) ((segid A and resid 4 and name HD2# )) 6.00 4.20 0.00
assign ((segid A and resid 4 and name HD1# )) ((segid A and resid 10 and name HE# )) 4.98 3.18 0.00
assign ((segid A and resid 4 and name HD1# )) ((segid A and resid 10 and name HD# )) 6.00 4.20 0.00
assign ((segid A and resid 4 and name HD1# )) ((segid A and resid 9 and name HE# )) 4.75 2.95 0.00
assign ((segid A and resid 4 and name HD1# )) ((segid A and resid 9 and name HD# )) 5.59 3.79 0.00
assign ((segid A and resid 4 and name HA )) ((segid A and resid 4 and name HD1# )) 4.34 2.54 0.00
assign ((segid A and resid 7 and name HN )) ((segid A and resid 7 and name HG2# )) 4.85 3.05 0.00
assign ((segid A and resid 46 and name HN )) ((segid A and resid 47 and name HG2# )) 6.00 4.20 0.00
assign ((segid A and resid 47 and name HG2# )) ((segid A and resid 52 and name HN )) 4.99 3.19 0.00
assign ((segid A and resid 46 and name HN )) ((segid A and resid 46 and name HD2# )) 4.91 3.11 0.00
assign ((segid A and resid 47 and name HG2# )) ((segid A and resid 48 and name HN )) 3.95 2.15 0.00
assign ((segid A and resid 47 and name HG2# )) ((segid A and resid 51 and name HN )) 6.00 4.20 0.00
assign ((segid A and resid 7 and name HA )) ((segid A and resid 7 and name HG2# )) 4.09 2.29 0.00
assign ((segid A and resid 52 and name HN )) ((segid A and resid 52 and name HG2# )) 5.13 3.33 0.00
assign ((segid A and resid 52 and name HA )) ((segid A and resid 52 and name HG2# )) 3.45 1.65 0.00
assign ((segid A and resid 46 and name HB1 )) ((segid A and resid 46 and name HD2# )) 3.28 1.48 0.00
assign ((segid A and resid 10 and name HZ )) ((segid A and resid 37 and name HG1# )) 4.31 2.51 0.00
assign ((segid A and resid 10 and name HE# )) ((segid A and resid 37 and name HG1# )) 4.11 2.31 0.00
assign ((segid A and resid 10 and name HE# )) ((segid A and resid 37 and name HG2# )) 4.11 2.31 0.00
assign ((segid A and resid 48 and name HN )) ((segid A and resid 51 and name HG2# )) 5.55 3.75 0.00
assign ((segid A and resid 47 and name HN )) ((segid A and resid 51 and name HG2# )) 4.92 3.12 0.00
assign ((segid A and resid 30 and name HA )) ((segid A and resid 66 and name HG1# )) 5.20 3.40 0.00
assign ((segid A and resid 30 and name HN )) ((segid A and resid 66 and name HG1# )) 3.88 2.08 0.00
assign ((segid A and resid 52 and name HN )) ((segid A and resid 52 and name HG1# )) 5.13 3.33 0.00
assign ((segid A and resid 59 and name HA )) ((segid A and resid 62 and name HB# )) 6.00 4.20 0.00
assign ((segid A and resid 14 and name HD1# )) ((segid A and resid 62 and name HB# )) 3.91 2.11 0.00
assign ((segid A and resid 40 and name HA )) ((segid A and resid 40 and name HG2# )) 3.55 1.75 0.00
assign ((segid A and resid 37 and name HA )) ((segid A and resid 40 and name HG2# )) 5.52 3.72 0.00
assign ((segid A and resid 7 and name HG1# )) ((segid A and resid 61 and name HG1 )) 6.00 4.20 0.00
assign ((segid A and resid 7 and name HG1# )) ((segid A and resid 61 and name HG2 )) 6.00 4.20 0.00
assign ((segid A and resid 37 and name HN )) ((segid A and resid 59 and name HD1# )) 5.39 3.59 0.00
assign ((segid A and resid 42 and name HG2# )) ((segid A and resid 43 and name HN )) 5.19 3.39 0.00
assign ((segid A and resid 42 and name HN )) ((segid A and resid 42 and name HG2# )) 4.09 2.29 0.00
assign ((segid A and resid 50 and name HB# )) ((segid A and resid 51 and name HN )) 4.61 2.81 0.00
assign ((segid A and resid 48 and name HE2 )) ((segid A and resid 50 and name HB# )) 5.27 3.47 0.00
assign ((segid A and resid 48 and name HE1 )) ((segid A and resid 50 and name HB# )) 5.27 3.47 0.00
assign ((segid A and resid 50 and name HB# )) ((segid A and resid 51 and name HB )) 6.00 4.20 0.00
assign ((segid A and resid 50 and name HN )) ((segid A and resid 50 and name HB# )) 3.44 1.64 0.00
assign ((segid A and resid 50 and name HB# )) ((segid A and resid 53 and name HN )) 5.44 3.64 0.00
assign ((segid A and resid 56 and name HG2# )) ((segid A and resid 57 and name HN )) 6.00 4.20 0.00
assign ((segid A and resid 56 and name HG2# )) ((segid A and resid 60 and name HN )) 6.00 4.20 0.00
assign ((segid A and resid 53 and name HA )) ((segid A and resid 56 and name HG2# )) 6.00 4.20 0.00
assign ((segid A and resid 56 and name HG2# )) ((segid A and resid 57 and name HA )) 6.00 4.20 0.00
assign ((segid A and resid 38 and name HA )) ((segid A and resid 56 and name HG2# )) 6.00 4.20 0.00
assign ((segid A and resid 25 and name HN )) ((segid A and resid 29 and name HB# )) 3.80 2.00 0.00
assign ((segid A and resid 24 and name HB1 )) ((segid A and resid 29 and name HB# )) 5.10 3.30 0.00
assign ((segid A and resid 14 and name HA )) ((segid A and resid 29 and name HB# )) 4.23 2.43 0.00
assign ((segid A and resid 27 and name HG2# )) ((segid A and resid 29 and name HB# )) 5.40 3.60 0.00
assign ((segid A and resid 27 and name HG2# )) ((segid A and resid 28 and name HN )) 6.00 4.20 0.00
assign ((segid A and resid 26 and name HN )) ((segid A and resid 27 and name HG2# )) 6.00 4.20 0.00
assign ((segid A and resid 18 and name HG2# )) ((segid A and resid 18 and name HG1# )) 3.19 1.39 0.00
assign ((segid A and resid 15 and name HN )) ((segid A and resid 18 and name HG2# )) 5.06 3.26 0.00
assign ((segid A and resid 18 and name HN )) ((segid A and resid 18 and name HG2# )) 4.43 2.63 0.00
assign ((segid A and resid 18 and name HG2# )) ((segid A and resid 19 and name HN )) 4.72 2.92 0.00
assign ((segid A and resid 15 and name HA )) ((segid A and resid 18 and name HG2# )) 5.28 3.48 0.00
assign ((segid A and resid 18 and name HG2# )) ((segid A and resid 29 and name HB# )) 3.62 1.82 0.00
assign ((segid A and resid 11 and name HG2# )) ((segid A and resid 14 and name HN )) 6.00 4.20 0.00
assign ((segid A and resid 11 and name HG2# )) ((segid A and resid 62 and name HA )) 4.50 2.70 0.00
assign ((segid A and resid 11 and name HG2# )) ((segid A and resid 62 and name HB# )) 3.41 1.61 0.00
assign ((segid A and resid 11 and name HD1# )) ((segid A and resid 62 and name HA )) 4.49 2.69 0.00
assign ((segid A and resid 11 and name HB )) ((segid A and resid 11 and name HD1# )) 2.98 1.18 0.00
assign ((segid A and resid 11 and name HD1# )) ((segid A and resid 62 and name HB# )) 3.56 1.76 0.00
assign ((segid A and resid 18 and name HN )) ((segid A and resid 18 and name HD1# )) 4.99 3.19 0.00
assign ((segid A and resid 18 and name HD1# )) ((segid A and resid 19 and name HA )) 6.00 4.20 0.00
assign ((segid A and resid 18 and name HD1# )) ((segid A and resid 29 and name HB# )) 4.55 2.75 0.00
assign ((segid A and resid 15 and name HA )) ((segid A and resid 18 and name HD1# )) 3.94 2.14 0.00
assign ((segid A and resid 18 and name HA )) ((segid A and resid 18 and name HD1# )) 4.49 2.69 0.00
assign ((segid A and resid 56 and name HN )) ((segid A and resid 56 and name HD1# )) 5.19 3.39 0.00
assign ((segid A and resid 42 and name HB )) ((segid A and resid 56 and name HD1# )) 5.86 4.06 0.00
assign ((segid A and resid 56 and name HB )) ((segid A and resid 56 and name HD1# )) 3.48 1.68 0.00
assign ((segid A and resid 27 and name HD1# )) ((segid A and resid 67 and name HN )) 6.00 4.20 0.00
assign ((segid A and resid 27 and name HD1# )) ((segid A and resid 71 and name HN )) 4.98 3.18 0.00
assign ((segid A and resid 27 and name HN )) ((segid A and resid 27 and name HD1# )) 5.00 3.20 0.00
assign ((segid A and resid 27 and name HD1# )) ((segid A and resid 63 and name HE# )) 6.00 4.20 0.00
assign ((segid A and resid 27 and name HB )) ((segid A and resid 27 and name HD1# )) 3.28 1.48 0.00
assign ((segid A and resid 21 and name HA )) ((segid A and resid 24 and name HB2 )) 6.00 4.20 0.00
assign ((segid A and resid 7 and name HG2# )) ((segid A and resid 61 and name HG1 )) 6.00 4.20 0.00
assign ((segid A and resid 41 and name HN )) ((segid A and resid 41 and name HG )) 6.00 4.20 0.00
assign ((segid A and resid 14 and name HD2# )) ((segid A and resid 62 and name HB# )) 3.91 2.11 0.00
assign ((segid A and resid 34 and name HD# )) ((segid A and resid 59 and name HD2# )) 6.00 4.20 0.00
assign ((segid A and resid 47 and name HG2# )) ((segid A and resid 51 and name HG2# )) 2.79 0.99 0.00
assign ((segid A and resid 47 and name HG2# )) ((segid A and resid 51 and name HG1# )) 2.79 0.99 0.00
assign ((segid A and resid 18 and name HG1# )) ((segid A and resid 29 and name HB# )) 5.35 3.55 0.00
assign ((segid A and resid 29 and name HB# )) ((segid A and resid 66 and name HB )) 5.61 3.81 0.00
assign ((segid A and resid 16 and name HA )) ((segid A and resid 20 and name HE# )) 4.71 2.91 0.00
assign ((segid A and resid 19 and name HN )) ((segid A and resid 20 and name HE# )) 5.85 4.05 0.00
assign ((segid A and resid 17 and name HN )) ((segid A and resid 20 and name HE# )) 6.00 4.20 0.00
assign ((segid A and resid 3 and name HA )) ((segid A and resid 4 and name HN )) 3.44 1.64 0.00
assign ((segid A and resid 4 and name HN )) ((segid A and resid 4 and name HG )) 4.88 3.08 0.00
assign ((segid A and resid 4 and name HN )) ((segid A and resid 4 and name HD1# )) 6.00 4.20 0.00
assign ((segid A and resid 48 and name HN )) ((segid A and resid 51 and name HN )) 6.00 4.20 0.00
assign ((segid A and resid 47 and name HN )) ((segid A and resid 48 and name HN )) 5.37 3.57 0.00
assign ((segid A and resid 47 and name HA )) ((segid A and resid 48 and name HN )) 3.28 1.48 0.00
assign ((segid A and resid 47 and name HB )) ((segid A and resid 48 and name HN )) 6.00 4.20 0.00
assign ((segid A and resid 48 and name HN )) ((segid A and resid 51 and name HG1# )) 5.55 3.75 0.00
assign ((segid A and resid 25 and name HB1 )) ((segid A and resid 26 and name HN )) 4.39 2.59 0.00
assign ((segid A and resid 26 and name HN )) ((segid A and resid 26 and name HB1 )) 4.06 2.26 0.00
assign ((segid A and resid 26 and name HN )) ((segid A and resid 26 and name HG1 )) 5.14 3.34 0.00
assign ((segid A and resid 26 and name HN )) ((segid A and resid 26 and name HG2 )) 5.14 3.34 0.00
assign ((segid A and resid 45 and name HN )) ((segid A and resid 46 and name HG )) 6.00 4.20 0.00
assign ((segid A and resid 12 and name HN )) ((segid A and resid 14 and name HN )) 5.65 3.85 0.00
assign ((segid A and resid 14 and name HN )) ((segid A and resid 16 and name HN )) 5.85 4.05 0.00
assign ((segid A and resid 14 and name HN )) ((segid A and resid 14 and name HG )) 3.83 2.03 0.00
assign ((segid A and resid 14 and name HN )) ((segid A and resid 14 and name HD2# )) 3.96 2.16 0.00
assign ((segid A and resid 14 and name HN )) ((segid A and resid 14 and name HD1# )) 3.96 2.16 0.00
assign ((segid A and resid 14 and name HN )) ((segid A and resid 33 and name HE# )) 6.00 4.20 0.00
assign ((segid A and resid 26 and name HA )) ((segid A and resid 29 and name HN )) 4.98 3.18 0.00
assign ((segid A and resid 28 and name HN )) ((segid A and resid 29 and name HN )) 4.15 2.35 0.00
assign ((segid A and resid 25 and name HB2 )) ((segid A and resid 29 and name HN )) 4.46 2.66 0.00
assign ((segid A and resid 29 and name HN )) ((segid A and resid 31 and name HN )) 6.00 4.20 0.00
assign ((segid A and resid 27 and name HN )) ((segid A and resid 29 and name HN )) 5.65 3.85 0.00
assign ((segid A and resid 27 and name HB )) ((segid A and resid 29 and name HN )) 6.00 4.20 0.00
assign ((segid A and resid 27 and name HG2# )) ((segid A and resid 29 and name HN )) 6.00 4.20 0.00
assign ((segid A and resid 59 and name HN )) ((segid A and resid 59 and name HD1# )) 4.98 3.18 0.00
assign ((segid A and resid 59 and name HN )) ((segid A and resid 59 and name HD2# )) 4.98 3.18 0.00
assign ((segid A and resid 56 and name HD1# )) ((segid A and resid 59 and name HN )) 6.00 4.20 0.00
assign ((segid A and resid 56 and name HB )) ((segid A and resid 59 and name HN )) 6.00 4.20 0.00
assign ((segid A and resid 71 and name HA )) ((segid A and resid 72 and name HN )) 3.07 1.27 0.00
assign ((segid A and resid 72 and name HN )) ((segid A and resid 74 and name HG1 )) 6.00 4.20 0.00
assign ((segid A and resid 72 and name HN )) ((segid A and resid 74 and name HG2 )) 6.00 4.20 0.00
assign ((segid A and resid 72 and name HN )) ((segid A and resid 72 and name HB2 )) 4.02 2.22 0.00
assign ((segid A and resid 11 and name HN )) ((segid A and resid 62 and name HA )) 5.83 4.03 0.00
assign ((segid A and resid 11 and name HN )) ((segid A and resid 62 and name HB# )) 4.38 2.58 0.00
assign ((segid A and resid 11 and name HN )) ((segid A and resid 11 and name HD1# )) 4.03 2.23 0.00
assign ((segid A and resid 11 and name HN )) ((segid A and resid 13 and name HN )) 6.00 4.20 0.00
assign ((segid A and resid 11 and name HN )) ((segid A and resid 11 and name HB )) 3.37 1.57 0.00
assign ((segid A and resid 25 and name HN )) ((segid A and resid 28 and name HN )) 6.00 4.20 0.00
assign ((segid A and resid 38 and name HA )) ((segid A and resid 41 and name HN )) 6.00 4.20 0.00
assign ((segid A and resid 24 and name HB2 )) ((segid A and resid 25 and name HN )) 3.75 1.95 0.00
assign ((segid A and resid 32 and name HN )) ((segid A and resid 33 and name HN )) 3.91 2.11 0.00
assign ((segid A and resid 33 and name HN )) ((segid A and resid 33 and name HD# )) 4.91 3.11 0.00
assign ((segid A and resid 33 and name HN )) ((segid A and resid 33 and name HB1 )) 3.88 2.08 0.00
assign ((segid A and resid 30 and name HA )) ((segid A and resid 33 and name HN )) 4.44 2.64 0.00
assign ((segid A and resid 33 and name HN )) ((segid A and resid 33 and name HB2 )) 3.88 2.08 0.00
assign ((segid A and resid 48 and name HN )) ((segid A and resid 49 and name HN )) 4.56 2.76 0.00
assign ((segid A and resid 48 and name HA )) ((segid A and resid 49 and name HN )) 3.11 1.31 0.00
assign ((segid A and resid 49 and name HN )) ((segid A and resid 49 and name HB1 )) 4.14 2.34 0.00
assign ((segid A and resid 49 and name HN )) ((segid A and resid 49 and name HB2 )) 4.14 2.34 0.00
assign ((segid A and resid 11 and name HN )) ((segid A and resid 11 and name HG2# )) 4.12 2.32 0.00
assign ((segid A and resid 2 and name HB1 )) ((segid A and resid 3 and name HN )) 5.08 3.28 0.00
assign ((segid A and resid 3 and name HN )) ((segid A and resid 4 and name HN )) 6.00 4.20 0.00
assign ((segid A and resid 2 and name HB2 )) ((segid A and resid 3 and name HN )) 5.08 3.28 0.00
assign ((segid A and resid 40 and name HN )) ((segid A and resid 41 and name HN )) 3.76 1.96 0.00
assign ((segid A and resid 40 and name HG2# )) ((segid A and resid 41 and name HN )) 4.55 2.75 0.00
assign ((segid A and resid 41 and name HN )) ((segid A and resid 42 and name HN )) 3.55 1.75 0.00
assign ((segid A and resid 33 and name HE# )) ((segid A and resid 37 and name HN )) 6.00 4.20 0.00
assign ((segid A and resid 33 and name HD# )) ((segid A and resid 37 and name HN )) 6.00 4.20 0.00
assign ((segid A and resid 37 and name HN )) ((segid A and resid 37 and name HB )) 3.77 1.97 0.00
assign ((segid A and resid 37 and name HN )) ((segid A and resid 59 and name HD2# )) 5.39 3.59 0.00
assign ((segid A and resid 33 and name HN )) ((segid A and resid 37 and name HN )) 6.00 4.20 0.00
assign ((segid A and resid 29 and name HB# )) ((segid A and resid 33 and name HN )) 6.00 4.20 0.00
assign ((segid A and resid 24 and name HN )) ((segid A and resid 25 and name HN )) 4.41 2.61 0.00
assign ((segid A and resid 23 and name HN )) ((segid A and resid 24 and name HN )) 3.39 1.59 0.00
assign ((segid A and resid 24 and name HN )) ((segid A and resid 24 and name HB2 )) 3.59 1.79 0.00
assign ((segid A and resid 21 and name HA )) ((segid A and resid 24 and name HN )) 6.00 4.20 0.00
assign ((segid A and resid 73 and name HA )) ((segid A and resid 74 and name HN )) 3.22 1.42 0.00
assign ((segid A and resid 10 and name HN )) ((segid A and resid 11 and name HN )) 4.31 2.51 0.00
assign ((segid A and resid 9 and name HD# )) ((segid A and resid 10 and name HN )) 4.68 2.88 0.00
assign ((segid A and resid 9 and name HN )) ((segid A and resid 10 and name HN )) 4.24 2.44 0.00
assign ((segid A and resid 65 and name HN )) ((segid A and resid 67 and name HN )) 4.71 2.91 0.00
assign ((segid A and resid 67 and name HN )) ((segid A and resid 68 and name HN )) 3.61 1.81 0.00
assign ((segid A and resid 67 and name HN )) ((segid A and resid 67 and name HB2 )) 3.66 1.86 0.00
assign ((segid A and resid 8 and name HA )) ((segid A and resid 10 and name HN )) 4.85 3.05 0.00
assign ((segid A and resid 59 and name HN )) ((segid A and resid 60 and name HN )) 3.82 2.02 0.00
assign ((segid A and resid 59 and name HD1# )) ((segid A and resid 60 and name HN )) 6.00 4.20 0.00
assign ((segid A and resid 66 and name HN )) ((segid A and resid 67 and name HG1 )) 4.66 2.86 0.00
assign ((segid A and resid 66 and name HN )) ((segid A and resid 66 and name HG2# )) 3.70 1.90 0.00
assign ((segid A and resid 66 and name HN )) ((segid A and resid 66 and name HG1# )) 3.70 1.90 0.00
assign ((segid A and resid 59 and name HD2# )) ((segid A and resid 60 and name HN )) 6.00 4.20 0.00
assign ((segid A and resid 46 and name HN )) ((segid A and resid 47 and name HB )) 5.29 3.49 0.00
assign ((segid A and resid 65 and name HN )) ((segid A and resid 66 and name HN )) 3.41 1.61 0.00
assign ((segid A and resid 18 and name HN )) ((segid A and resid 25 and name HN )) 6.00 4.20 0.00
assign ((segid A and resid 38 and name HN )) ((segid A and resid 41 and name HN )) 6.00 4.20 0.00
assign ((segid A and resid 37 and name HN )) ((segid A and resid 38 and name HN )) 3.55 1.75 0.00
assign ((segid A and resid 38 and name HN )) ((segid A and resid 39 and name HN )) 3.87 2.07 0.00
assign ((segid A and resid 38 and name HN )) ((segid A and resid 40 and name HN )) 5.93 4.13 0.00
assign ((segid A and resid 15 and name HA )) ((segid A and resid 18 and name HN )) 4.44 2.64 0.00
assign ((segid A and resid 37 and name HG2# )) ((segid A and resid 38 and name HN )) 4.63 2.83 0.00
assign ((segid A and resid 33 and name HE# )) ((segid A and resid 38 and name HN )) 6.00 4.20 0.00
assign ((segid A and resid 33 and name HD# )) ((segid A and resid 38 and name HN )) 6.00 4.20 0.00
assign ((segid A and resid 34 and name HE# )) ((segid A and resid 38 and name HN )) 6.00 4.20 0.00
assign ((segid A and resid 37 and name HB )) ((segid A and resid 38 and name HN )) 4.58 2.78 0.00
assign ((segid A and resid 49 and name HA )) ((segid A and resid 52 and name HN )) 4.98 3.18 0.00
assign ((segid A and resid 52 and name HN )) ((segid A and resid 52 and name HB )) 3.70 1.90 0.00
assign ((segid A and resid 69 and name HN )) ((segid A and resid 70 and name HN )) 3.33 1.53 0.00
assign ((segid A and resid 68 and name HN )) ((segid A and resid 70 and name HN )) 4.88 3.08 0.00
assign ((segid A and resid 67 and name HA )) ((segid A and resid 70 and name HN )) 4.29 2.49 0.00
assign ((segid A and resid 27 and name HD1# )) ((segid A and resid 70 and name HN )) 5.25 3.45 0.00
assign ((segid A and resid 38 and name HN )) ((segid A and resid 40 and name HB )) 6.00 4.20 0.00
assign ((segid A and resid 18 and name HN )) ((segid A and resid 29 and name HB# )) 4.93 3.13 0.00
assign ((segid A and resid 18 and name HN )) ((segid A and resid 18 and name HG1# )) 3.55 1.75 0.00
assign ((segid A and resid 37 and name HG1# )) ((segid A and resid 38 and name HN )) 4.63 2.83 0.00
assign ((segid A and resid 7 and name HN )) ((segid A and resid 7 and name HB )) 4.07 2.27 0.00
assign ((segid A and resid 7 and name HN )) ((segid A and resid 7 and name HG1# )) 4.85 3.05 0.00
assign ((segid A and resid 7 and name HN )) ((segid A and resid 10 and name HN )) 6.00 4.20 0.00
assign ((segid A and resid 34 and name HN )) ((segid A and resid 36 and name HN )) 6.00 4.20 0.00
assign ((segid A and resid 33 and name HN )) ((segid A and resid 34 and name HN )) 3.85 2.05 0.00
assign ((segid A and resid 34 and name HN )) ((segid A and resid 35 and name HN )) 4.15 2.35 0.00
assign ((segid A and resid 32 and name HN )) ((segid A and resid 34 and name HN )) 6.00 4.20 0.00
assign ((segid A and resid 56 and name HN )) ((segid A and resid 57 and name HN )) 4.26 2.46 0.00
assign ((segid A and resid 54 and name HA )) ((segid A and resid 57 and name HN )) 4.49 2.69 0.00
assign ((segid A and resid 55 and name HA )) ((segid A and resid 57 and name HN )) 4.84 3.04 0.00
assign ((segid A and resid 56 and name HD1# )) ((segid A and resid 57 and name HN )) 4.52 2.72 0.00
assign ((segid A and resid 30 and name HA )) ((segid A and resid 34 and name HN )) 3.89 2.09 0.00
assign ((segid A and resid 46 and name HN )) ((segid A and resid 47 and name HN )) 3.23 1.43 0.00
assign ((segid A and resid 46 and name HA )) ((segid A and resid 47 and name HN )) 3.55 1.75 0.00
assign ((segid A and resid 47 and name HN )) ((segid A and resid 47 and name HB )) 3.74 1.94 0.00
assign ((segid A and resid 47 and name HN )) ((segid A and resid 47 and name HG2# )) 4.21 2.41 0.00
assign ((segid A and resid 47 and name HN )) ((segid A and resid 51 and name HG1# )) 4.92 3.12 0.00
assign ((segid A and resid 49 and name HN )) ((segid A and resid 50 and name HN )) 4.18 2.38 0.00
assign ((segid A and resid 50 and name HN )) ((segid A and resid 52 and name HN )) 4.98 3.18 0.00
assign ((segid A and resid 50 and name HN )) ((segid A and resid 51 and name HN )) 3.87 2.07 0.00
assign ((segid A and resid 47 and name HG2# )) ((segid A and resid 50 and name HN )) 6.00 4.20 0.00
assign ((segid A and resid 74 and name HA )) ((segid A and resid 75 and name HN )) 3.37 1.57 0.00
assign ((segid A and resid 44 and name HN )) ((segid A and resid 45 and name HN )) 4.68 2.88 0.00
assign ((segid A and resid 40 and name HG2# )) ((segid A and resid 44 and name HN )) 6.00 4.20 0.00
assign ((segid A and resid 42 and name HN )) ((segid A and resid 44 and name HN )) 5.16 3.36 0.00
assign ((segid A and resid 41 and name HA )) ((segid A and resid 44 and name HN )) 5.42 3.62 0.00
assign ((segid A and resid 42 and name HB )) ((segid A and resid 44 and name HN )) 5.64 3.84 0.00
assign ((segid A and resid 25 and name HB2 )) ((segid A and resid 28 and name HN )) 3.71 1.91 0.00
assign ((segid A and resid 21 and name HN )) ((segid A and resid 22 and name HN )) 3.90 2.10 0.00
assign ((segid A and resid 16 and name HG2 )) ((segid A and resid 21 and name HN )) 5.17 3.37 0.00
assign ((segid A and resid 53 and name HN )) ((segid A and resid 54 and name HN )) 3.79 1.99 0.00
assign ((segid A and resid 54 and name HN )) ((segid A and resid 55 and name HN )) 4.11 2.31 0.00
assign ((segid A and resid 52 and name HN )) ((segid A and resid 54 and name HN )) 4.42 2.62 0.00
assign ((segid A and resid 54 and name HN )) ((segid A and resid 56 and name HN )) 5.09 3.29 0.00
assign ((segid A and resid 51 and name HA )) ((segid A and resid 54 and name HN )) 4.33 2.53 0.00
assign ((segid A and resid 51 and name HN )) ((segid A and resid 54 and name HN )) 6.00 4.20 0.00
assign ((segid A and resid 34 and name HE# )) ((segid A and resid 63 and name HN )) 6.00 4.20 0.00
assign ((segid A and resid 62 and name HN )) ((segid A and resid 63 and name HN )) 3.73 1.93 0.00
assign ((segid A and resid 62 and name HB# )) ((segid A and resid 63 and name HN )) 4.21 2.41 0.00
assign ((segid A and resid 11 and name HD1# )) ((segid A and resid 63 and name HN )) 5.65 3.85 0.00
assign ((segid A and resid 59 and name HG )) ((segid A and resid 63 and name HN )) 6.00 4.20 0.00
assign ((segid A and resid 63 and name HN )) ((segid A and resid 66 and name HB )) 6.00 4.20 0.00
assign ((segid A and resid 34 and name HD# )) ((segid A and resid 63 and name HN )) 5.78 3.98 0.00
assign ((segid A and resid 61 and name HA )) ((segid A and resid 63 and name HN )) 6.00 4.20 0.00
assign ((segid A and resid 22 and name HN )) ((segid A and resid 24 and name HN )) 4.71 2.91 0.00
assign ((segid A and resid 21 and name HA )) ((segid A and resid 22 and name HN )) 3.42 1.62 0.00
assign ((segid A and resid 16 and name HA )) ((segid A and resid 22 and name HN )) 6.00 4.20 0.00
assign ((segid A and resid 54 and name HN )) ((segid A and resid 57 and name HN )) 5.82 4.02 0.00
assign ((segid A and resid 59 and name HN )) ((segid A and resid 61 and name HN )) 6.00 4.20 0.00
assign ((segid A and resid 11 and name HD1# )) ((segid A and resid 61 and name HN )) 5.89 4.09 0.00
assign ((segid A and resid 9 and name HN )) ((segid A and resid 9 and name HD# )) 5.30 3.50 0.00
assign ((segid A and resid 7 and name HA )) ((segid A and resid 9 and name HN )) 3.73 1.93 0.00
assign ((segid A and resid 7 and name HB )) ((segid A and resid 9 and name HN )) 4.79 2.99 0.00
assign ((segid A and resid 30 and name HN )) ((segid A and resid 33 and name HD# )) 6.00 4.20 0.00
assign ((segid A and resid 26 and name HN )) ((segid A and resid 27 and name HN )) 3.81 2.01 0.00
assign ((segid A and resid 27 and name HN )) ((segid A and resid 28 and name HN )) 3.96 2.16 0.00
assign ((segid A and resid 26 and name HG1 )) ((segid A and resid 27 and name HN )) 5.13 3.33 0.00
assign ((segid A and resid 26 and name HG2 )) ((segid A and resid 27 and name HN )) 5.13 3.33 0.00
assign ((segid A and resid 27 and name HN )) ((segid A and resid 27 and name HG2# )) 3.91 2.11 0.00
assign ((segid A and resid 15 and name HN )) ((segid A and resid 16 and name HN )) 3.39 1.59 0.00
assign ((segid A and resid 29 and name HN )) ((segid A and resid 30 and name HN )) 3.64 1.84 0.00
assign ((segid A and resid 18 and name HG2# )) ((segid A and resid 30 and name HN )) 5.68 3.88 0.00
assign ((segid A and resid 28 and name HN )) ((segid A and resid 30 and name HN )) 5.61 3.81 0.00
assign ((segid A and resid 55 and name HN )) ((segid A and resid 58 and name HN )) 5.64 3.84 0.00
assign ((segid A and resid 58 and name HN )) ((segid A and resid 59 and name HN )) 3.73 1.93 0.00
assign ((segid A and resid 30 and name HN )) ((segid A and resid 66 and name HB )) 4.48 2.68 0.00
assign ((segid A and resid 58 and name HN )) ((segid A and resid 59 and name HB2 )) 4.80 3.00 0.00
assign ((segid A and resid 27 and name HG2# )) ((segid A and resid 30 and name HN )) 6.00 4.20 0.00
assign ((segid A and resid 56 and name HD1# )) ((segid A and resid 58 and name HN )) 6.00 4.20 0.00
assign ((segid A and resid 8 and name HA )) ((segid A and resid 12 and name HN )) 4.13 2.33 0.00
assign ((segid A and resid 11 and name HB )) ((segid A and resid 12 and name HN )) 3.02 1.22 0.00
assign ((segid A and resid 41 and name HN )) ((segid A and resid 43 and name HN )) 5.54 3.74 0.00
assign ((segid A and resid 42 and name HN )) ((segid A and resid 43 and name HN )) 3.93 2.13 0.00
assign ((segid A and resid 43 and name HN )) ((segid A and resid 44 and name HN )) 3.37 1.57 0.00
assign ((segid A and resid 41 and name HA )) ((segid A and resid 43 and name HN )) 4.56 2.76 0.00
assign ((segid A and resid 42 and name HB )) ((segid A and resid 43 and name HN )) 4.21 2.41 0.00
assign ((segid A and resid 40 and name HG2# )) ((segid A and resid 43 and name HN )) 6.00 4.20 0.00
assign ((segid A and resid 42 and name HG1# )) ((segid A and resid 43 and name HN )) 5.19 3.39 0.00
assign ((segid A and resid 31 and name HN )) ((segid A and resid 33 and name HN )) 6.00 4.20 0.00
assign ((segid A and resid 30 and name HE# )) ((segid A and resid 31 and name HN )) 5.73 3.93 0.00
assign ((segid A and resid 29 and name HB# )) ((segid A and resid 31 and name HN )) 6.00 4.20 0.00
assign ((segid A and resid 30 and name HN )) ((segid A and resid 31 and name HN )) 4.01 2.21 0.00
assign ((segid A and resid 27 and name HG2# )) ((segid A and resid 31 and name HN )) 5.86 4.06 0.00
assign ((segid A and resid 10 and name HN )) ((segid A and resid 12 and name HN )) 5.63 3.83 0.00
assign ((segid A and resid 55 and name HN )) ((segid A and resid 57 and name HN )) 6.00 4.20 0.00
assign ((segid A and resid 55 and name HN )) ((segid A and resid 56 and name HN )) 3.65 1.85 0.00
assign ((segid A and resid 35 and name HN )) ((segid A and resid 36 and name HN )) 4.08 2.28 0.00
assign ((segid A and resid 18 and name HN )) ((segid A and resid 19 and name HN )) 3.54 1.74 0.00
assign ((segid A and resid 53 and name HA )) ((segid A and resid 56 and name HN )) 4.51 2.71 0.00
assign ((segid A and resid 52 and name HA )) ((segid A and resid 56 and name HN )) 5.38 3.58 0.00
assign ((segid A and resid 63 and name HN )) ((segid A and resid 64 and name HN )) 4.25 2.45 0.00
assign ((segid A and resid 64 and name HN )) ((segid A and resid 68 and name HN )) 6.00 4.20 0.00
assign ((segid A and resid 63 and name HE# )) ((segid A and resid 64 and name HN )) 4.38 2.58 0.00
assign ((segid A and resid 11 and name HD1# )) ((segid A and resid 64 and name HN )) 6.00 4.20 0.00
assign ((segid A and resid 11 and name HN )) ((segid A and resid 12 and name HN )) 4.10 2.30 0.00
assign ((segid A and resid 11 and name HG2# )) ((segid A and resid 12 and name HN )) 3.79 1.99 0.00
assign ((segid A and resid 52 and name HA )) ((segid A and resid 55 and name HN )) 4.36 2.56 0.00
assign ((segid A and resid 13 and name HN )) ((segid A and resid 33 and name HE# )) 5.63 3.83 0.00
assign ((segid A and resid 51 and name HN )) ((segid A and resid 52 and name HN )) 4.13 2.33 0.00
assign ((segid A and resid 48 and name HA )) ((segid A and resid 51 and name HN )) 5.24 3.44 0.00
assign ((segid A and resid 51 and name HN )) ((segid A and resid 51 and name HG2# )) 3.87 2.07 0.00
assign ((segid A and resid 51 and name HN )) ((segid A and resid 51 and name HG1# )) 3.87 2.07 0.00
assign ((segid A and resid 51 and name HN )) ((segid A and resid 53 and name HN )) 5.25 3.45 0.00
assign ((segid A and resid 15 and name HN )) ((segid A and resid 18 and name HD1# )) 4.40 2.60 0.00
assign ((segid A and resid 14 and name HN )) ((segid A and resid 15 and name HN )) 4.24 2.44 0.00
assign ((segid A and resid 11 and name HA )) ((segid A and resid 15 and name HN )) 5.52 3.72 0.00
assign ((segid A and resid 17 and name HN )) ((segid A and resid 19 and name HN )) 4.01 2.21 0.00
assign ((segid A and resid 11 and name HG2# )) ((segid A and resid 13 and name HN )) 6.00 4.20 0.00
assign ((segid A and resid 13 and name HN )) ((segid A and resid 14 and name HN )) 4.08 2.28 0.00
assign ((segid A and resid 12 and name HN )) ((segid A and resid 13 and name HN )) 3.37 1.57 0.00
assign ((segid A and resid 13 and name HN )) ((segid A and resid 15 and name HN )) 5.01 3.21 0.00
assign ((segid A and resid 70 and name HB1 )) ((segid A and resid 71 and name HN )) 4.13 2.33 0.00
assign ((segid A and resid 30 and name HN )) ((segid A and resid 32 and name HN )) 4.48 2.68 0.00
assign ((segid A and resid 31 and name HN )) ((segid A and resid 32 and name HN )) 4.39 2.59 0.00
assign ((segid A and resid 27 and name HD1# )) ((segid A and resid 68 and name HN )) 6.00 4.20 0.00
assign ((segid A and resid 52 and name HN )) ((segid A and resid 53 and name HN )) 4.08 2.28 0.00
assign ((segid A and resid 49 and name HA )) ((segid A and resid 53 and name HN )) 5.24 3.44 0.00
assign ((segid A and resid 53 and name HN )) ((segid A and resid 56 and name HB )) 6.00 4.20 0.00
assign ((segid A and resid 64 and name HA )) ((segid A and resid 68 and name HN )) 4.04 2.24 0.00
assign ((segid A and resid 67 and name HB2 )) ((segid A and resid 68 and name HN )) 4.15 2.35 0.00
assign ((segid A and resid 67 and name HG1 )) ((segid A and resid 68 and name HN )) 4.52 2.72 0.00
assign ((segid A and resid 66 and name HA )) ((segid A and resid 68 and name HN )) 6.00 4.20 0.00
assign ((segid A and resid 68 and name HN )) ((segid A and resid 70 and name HG1 )) 6.00 4.20 0.00
assign ((segid A and resid 68 and name HN )) ((segid A and resid 70 and name HG2 )) 6.00 4.20 0.00
assign ((segid A and resid 16 and name HN )) ((segid A and resid 20 and name HN )) 6.00 4.20 0.00
assign ((segid A and resid 18 and name HN )) ((segid A and resid 20 and name HN )) 5.93 4.13 0.00
assign ((segid A and resid 6 and name HN )) ((segid A and resid 7 and name HN )) 6.00 4.20 0.00
assign ((segid A and resid 21 and name HA )) ((segid A and resid 23 and name HN )) 4.54 2.74 0.00
assign ((segid A and resid 22 and name HA )) ((segid A and resid 23 and name HN )) 3.55 1.75 0.00
assign ((segid A and resid 22 and name HB1 )) ((segid A and resid 23 and name HN )) 4.16 2.36 0.00
assign ((segid A and resid 23 and name HN )) ((segid A and resid 24 and name HB2 )) 4.22 2.42 0.00
assign ((segid A and resid 4 and name HA )) ((segid A and resid 6 and name HN )) 6.00 4.20 0.00
assign ((segid A and resid 4 and name HN )) ((segid A and resid 6 and name HN )) 6.00 4.20 0.00
assign ((segid A and resid 40 and name HN )) ((segid A and resid 42 and name HN )) 5.53 3.73 0.00
assign ((segid A and resid 40 and name HN )) ((segid A and resid 40 and name HG2# )) 3.99 2.19 0.00
assign ((segid A and resid 36 and name HN )) ((segid A and resid 38 and name HN )) 5.92 4.12 0.00
assign ((segid A and resid 36 and name HN )) ((segid A and resid 37 and name HN )) 3.70 1.90 0.00
assign ((segid A and resid 39 and name HN )) ((segid A and resid 40 and name HN )) 3.98 2.18 0.00
assign ((segid A and resid 39 and name HN )) ((segid A and resid 40 and name HB )) 6.00 4.20 0.00
assign ((segid A and resid 39 and name HN )) ((segid A and resid 41 and name HN )) 6.00 4.20 0.00
assign ((segid A and resid 39 and name HN )) ((segid A and resid 42 and name HB )) 5.60 3.80 0.00
assign ((segid A and resid 40 and name HG2# )) ((segid A and resid 42 and name HN )) 6.00 4.20 0.00
assign ((segid A and resid 42 and name HN )) ((segid A and resid 42 and name HG1# )) 4.09 2.29 0.00
assign ((segid A and resid 68 and name HN )) ((segid A and resid 69 and name HN )) 2.93 1.13 0.00
assign ((segid A and resid 68 and name HB1 )) ((segid A and resid 69 and name HN )) 3.52 1.72 0.00
assign ((segid A and resid 67 and name HG1 )) ((segid A and resid 69 and name HN )) 6.00 4.20 0.00
assign ((segid A and resid 18 and name HD1# )) ((segid A and resid 69 and name HN )) 6.00 4.20 0.00
assign ((segid A and resid 27 and name HD1# )) ((segid A and resid 69 and name HN )) 6.00 4.20 0.00
assign ((segid A and resid 67 and name HN )) ((segid A and resid 69 and name HN )) 5.15 3.35 0.00
assign ((segid A and resid 8 and name HA )) ((segid A and resid 11 and name HN )) 4.27 2.47 0.00
assign ((segid A and resid 63 and name HD# )) ((segid A and resid 64 and name HN )) 4.23 2.43 0.00
assign ((segid A and resid 63 and name HN )) ((segid A and resid 63 and name HD# )) 4.56 2.76 0.00
assign ((segid A and resid 10 and name HZ )) ((segid A and resid 37 and name HG2# )) 4.31 2.51 0.00
assign ((segid A and resid 34 and name HN )) ((segid A and resid 34 and name HD# )) 4.87 3.07 0.00
assign ((segid A and resid 34 and name HD# )) ((segid A and resid 63 and name HA )) 4.43 2.63 0.00
assign ((segid A and resid 34 and name HA )) ((segid A and resid 34 and name HD# )) 4.44 2.64 0.00
assign ((segid A and resid 34 and name HD# )) ((segid A and resid 59 and name HD1# )) 6.00 4.20 0.00
assign ((segid A and resid 9 and name HD# )) ((segid A and resid 10 and name HE# )) 4.69 2.89 0.00
assign ((segid A and resid 4 and name HA )) ((segid A and resid 9 and name HD# )) 5.06 3.26 0.00
assign ((segid A and resid 4 and name HD2# )) ((segid A and resid 9 and name HD# )) 5.59 3.79 0.00
assign ((segid A and resid 4 and name HN )) ((segid A and resid 9 and name HD# )) 5.00 3.20 0.00
assign ((segid A and resid 9 and name HD# )) ((segid A and resid 10 and name HZ )) 6.00 4.20 0.00
assign ((segid A and resid 33 and name HD# )) ((segid A and resid 34 and name HN )) 4.36 2.56 0.00
assign ((segid A and resid 14 and name HG )) ((segid A and resid 33 and name HD# )) 6.00 4.20 0.00
assign ((segid A and resid 33 and name HD# )) ((segid A and resid 62 and name HB# )) 6.00 4.20 0.00
assign ((segid A and resid 10 and name HE# )) ((segid A and resid 10 and name HD# )) 3.15 1.35 0.00
assign ((segid A and resid 9 and name HE# )) ((segid A and resid 10 and name HD# )) 5.15 3.35 0.00
assign ((segid A and resid 10 and name HD# )) ((segid A and resid 33 and name HE# )) 5.32 3.52 0.00
assign ((segid A and resid 10 and name HD# )) ((segid A and resid 37 and name HB )) 6.00 4.20 0.00
assign ((segid A and resid 10 and name HD# )) ((segid A and resid 59 and name HB2 )) 6.00 4.20 0.00
assign ((segid A and resid 4 and name HN )) ((segid A and resid 9 and name HE# )) 5.26 3.46 0.00
assign ((segid A and resid 9 and name HE# )) ((segid A and resid 10 and name HE# )) 4.70 2.90 0.00
assign ((segid A and resid 4 and name HA )) ((segid A and resid 9 and name HE# )) 4.82 3.02 0.00
assign ((segid A and resid 33 and name HE# )) ((segid A and resid 59 and name HG )) 4.78 2.98 0.00
assign ((segid A and resid 14 and name HG )) ((segid A and resid 33 and name HE# )) 4.77 2.97 0.00
assign ((segid A and resid 33 and name HE# )) ((segid A and resid 34 and name HN )) 5.84 4.04 0.00
assign ((segid A and resid 63 and name HN )) ((segid A and resid 63 and name HE# )) 6.00 4.20 0.00
assign ((segid A and resid 63 and name HE# )) ((segid A and resid 63 and name HD# )) 2.93 1.13 0.00
assign ((segid A and resid 63 and name HE# )) ((segid A and resid 67 and name HG1 )) 3.95 2.15 0.00
assign ((segid A and resid 34 and name HE# )) ((segid A and resid 60 and name HN )) 5.40 3.60 0.00
assign ((segid A and resid 34 and name HE# )) ((segid A and resid 34 and name HD# )) 3.07 1.27 0.00
assign ((segid A and resid 34 and name HA )) ((segid A and resid 34 and name HE# )) 5.41 3.61 0.00
assign ((segid A and resid 2 and name HB# )) ((segid A and resid 3 and name HN )) 4.45 2.65 0.00
assign ((segid A and resid 3 and name HA )) ((segid A and resid 4 and name HB# )) 4.47 2.67 0.00
assign ((segid A and resid 3 and name HG# )) ((segid A and resid 4 and name HN )) 5.84 3.98 0.00
assign ((segid A and resid 3 and name HG# )) ((segid A and resid 4 and name HD# )) 5.79 3.99 0.00
assign ((segid A and resid 4 and name HN )) ((segid A and resid 4 and name HB# )) 3.57 1.77 0.00
assign ((segid A and resid 4 and name HN )) ((segid A and resid 4 and name HD# )) 4.50 2.70 0.00
assign ((segid A and resid 4 and name HA )) ((segid A and resid 4 and name HD# )) 3.68 1.88 0.00
assign ((segid A and resid 4 and name HB# )) ((segid A and resid 4 and name HD# )) 2.80 1.00 0.00
assign ((segid A and resid 4 and name HD# )) ((segid A and resid 9 and name HE# )) 4.06 2.26 0.00
assign ((segid A and resid 4 and name HD# )) ((segid A and resid 9 and name HD# )) 4.37 2.57 0.00
assign ((segid A and resid 4 and name HD# )) ((segid A and resid 10 and name HE# )) 4.15 2.35 0.00
assign ((segid A and resid 7 and name HN )) ((segid A and resid 7 and name HG# )) 4.20 2.40 0.00
assign ((segid A and resid 7 and name HG# )) ((segid A and resid 10 and name HN )) 5.69 3.89 0.00
assign ((segid A and resid 7 and name HG# )) ((segid A and resid 61 and name HN )) 5.92 4.12 0.00
assign ((segid A and resid 7 and name HG# )) ((segid A and resid 61 and name HG# )) 3.63 1.83 0.00
assign ((segid A and resid 7 and name HG# )) ((segid A and resid 62 and name HN )) 5.92 4.12 0.00
assign ((segid A and resid 8 and name HB# )) ((segid A and resid 9 and name HN )) 4.11 2.31 0.00
assign ((segid A and resid 8 and name HB# )) ((segid A and resid 11 and name HN )) 5.50 3.70 0.00
assign ((segid A and resid 8 and name HB# )) ((segid A and resid 11 and name HG2# )) 5.14 3.34 0.00
assign ((segid A and resid 10 and name HB# )) ((segid A and resid 37 and name HG# )) 5.18 3.38 0.00
assign ((segid A and resid 10 and name HB# )) ((segid A and resid 59 and name HD# )) 4.40 2.60 0.00
assign ((segid A and resid 10 and name HB# )) ((segid A and resid 62 and name HN )) 5.81 4.01 0.00
assign ((segid A and resid 10 and name HB# )) ((segid A and resid 62 and name HB# )) 4.94 3.14 0.00
assign ((segid A and resid 10 and name HE# )) ((segid A and resid 37 and name HG# )) 3.51 1.71 0.00
assign ((segid A and resid 10 and name HE# )) ((segid A and resid 59 and name HD# )) 3.82 2.02 0.00
assign ((segid A and resid 10 and name HD# )) ((segid A and resid 37 and name HG# )) 3.82 2.02 0.00
assign ((segid A and resid 10 and name HZ )) ((segid A and resid 37 and name HG# )) 3.15 1.35 0.00
assign ((segid A and resid 11 and name HN )) ((segid A and resid 11 and name HG1# )) 4.15 2.35 0.00
assign ((segid A and resid 11 and name HN )) ((segid A and resid 61 and name HG# )) 5.81 4.01 0.00
assign ((segid A and resid 11 and name HB )) ((segid A and resid 61 and name HG# )) 5.81 4.01 0.00
assign ((segid A and resid 11 and name HG2# )) ((segid A and resid 12 and name HG# )) 4.76 2.96 0.00
assign ((segid A and resid 11 and name HG2# )) ((segid A and resid 14 and name HD# )) 3.52 1.72 0.00
assign ((segid A and resid 11 and name HG1# )) ((segid A and resid 13 and name HN )) 5.81 4.01 0.00
assign ((segid A and resid 11 and name HG1# )) ((segid A and resid 61 and name HG# )) 4.17 2.37 0.00
assign ((segid A and resid 11 and name HG1# )) ((segid A and resid 62 and name HN )) 3.89 2.09 0.00
assign ((segid A and resid 11 and name HG1# )) ((segid A and resid 62 and name HA )) 3.48 1.68 0.00
assign ((segid A and resid 11 and name HD1# )) ((segid A and resid 14 and name HD# )) 4.10 2.30 0.00
assign ((segid A and resid 11 and name HD1# )) ((segid A and resid 61 and name HB# )) 4.40 2.60 0.00
assign ((segid A and resid 11 and name HD1# )) ((segid A and resid 61 and name HG# )) 4.07 2.27 0.00
assign ((segid A and resid 12 and name HN )) ((segid A and resid 12 and name HB# )) 3.32 1.52 0.00
assign ((segid A and resid 12 and name HN )) ((segid A and resid 12 and name HG# )) 3.70 1.90 0.00
assign ((segid A and resid 12 and name HN )) ((segid A and resid 13 and name HB# )) 5.03 3.23 0.00
assign ((segid A and resid 12 and name HN )) ((segid A and resid 14 and name HD# )) 4.62 2.82 0.00
assign ((segid A and resid 12 and name HA )) ((segid A and resid 12 and name HG# )) 3.35 1.55 0.00
assign ((segid A and resid 12 and name HG# )) ((segid A and resid 13 and name HN )) 5.81 4.01 0.00
assign ((segid A and resid 13 and name HN )) ((segid A and resid 14 and name HD# )) 5.38 3.58 0.00
assign ((segid A and resid 13 and name HB# )) ((segid A and resid 15 and name HN )) 5.81 4.01 0.00
assign ((segid A and resid 13 and name HB# )) ((segid A and resid 33 and name HE# )) 4.45 2.65 0.00
assign ((segid A and resid 14 and name HN )) ((segid A and resid 14 and name HB# )) 3.54 1.74 0.00
assign ((segid A and resid 14 and name HN )) ((segid A and resid 14 and name HD# )) 3.38 1.58 0.00
assign ((segid A and resid 14 and name HN )) ((segid A and resid 33 and name HB# )) 5.81 4.01 0.00
assign ((segid A and resid 14 and name HA )) ((segid A and resid 33 and name HB# )) 5.69 3.89 0.00
assign ((segid A and resid 14 and name HB# )) ((segid A and resid 18 and name HG2# )) 3.94 2.14 0.00
assign ((segid A and resid 14 and name HB# )) ((segid A and resid 18 and name HD1# )) 4.85 3.05 0.00
assign ((segid A and resid 14 and name HD# )) ((segid A and resid 15 and name HN )) 3.80 2.00 0.00
assign ((segid A and resid 14 and name HD# )) ((segid A and resid 17 and name HN )) 5.92 4.12 0.00
assign ((segid A and resid 14 and name HD# )) ((segid A and resid 29 and name HN )) 5.92 4.12 0.00
assign ((segid A and resid 14 and name HD# )) ((segid A and resid 29 and name HB# )) 5.92 4.12 0.00
assign ((segid A and resid 14 and name HD# )) ((segid A and resid 30 and name HN )) 5.92 4.12 0.00
assign ((segid A and resid 14 and name HD# )) ((segid A and resid 30 and name HA )) 4.69 2.89 0.00
assign ((segid A and resid 14 and name HD# )) ((segid A and resid 33 and name HN )) 5.92 4.12 0.00
assign ((segid A and resid 14 and name HD# )) ((segid A and resid 33 and name HB# )) 5.07 3.27 0.00
assign ((segid A and resid 14 and name HD# )) ((segid A and resid 33 and name HE# )) 3.89 2.09 0.00
assign ((segid A and resid 14 and name HD# )) ((segid A and resid 33 and name HD# )) 3.78 1.98 0.00
assign ((segid A and resid 14 and name HD# )) ((segid A and resid 34 and name HN )) 5.87 4.07 0.00
assign ((segid A and resid 14 and name HD# )) ((segid A and resid 59 and name HD# )) 4.52 2.72 0.00
assign ((segid A and resid 14 and name HD# )) ((segid A and resid 61 and name HG# )) 5.73 3.93 0.00
assign ((segid A and resid 14 and name HD# )) ((segid A and resid 62 and name HA )) 3.88 2.08 0.00
assign ((segid A and resid 14 and name HD# )) ((segid A and resid 62 and name HB# )) 2.93 1.13 0.00
assign ((segid A and resid 14 and name HD# )) ((segid A and resid 63 and name HN )) 5.44 3.64 0.00
assign ((segid A and resid 14 and name HD# )) ((segid A and resid 63 and name HA )) 4.16 2.36 0.00
assign ((segid A and resid 14 and name HD# )) ((segid A and resid 66 and name HN )) 5.92 4.12 0.00
assign ((segid A and resid 14 and name HD# )) ((segid A and resid 66 and name HB )) 3.94 2.14 0.00
assign ((segid A and resid 15 and name HN )) ((segid A and resid 15 and name HB# )) 3.39 1.59 0.00
assign ((segid A and resid 15 and name HB# )) ((segid A and resid 16 and name HN )) 3.15 1.35 0.00
assign ((segid A and resid 16 and name HA )) ((segid A and resid 16 and name HG# )) 3.17 1.37 0.00
assign ((segid A and resid 16 and name HG# )) ((segid A and resid 17 and name HA )) 4.44 2.64 0.00
assign ((segid A and resid 16 and name HG# )) ((segid A and resid 18 and name HN )) 5.81 4.01 0.00
assign ((segid A and resid 16 and name HG# )) ((segid A and resid 20 and name HN )) 4.50 2.70 0.00
assign ((segid A and resid 16 and name HG# )) ((segid A and resid 21 and name HN )) 4.45 2.65 0.00
assign ((segid A and resid 16 and name HG# )) ((segid A and resid 22 and name HN )) 4.06 2.26 0.00
assign ((segid A and resid 17 and name HA )) ((segid A and resid 20 and name HG# )) 5.81 4.01 0.00
assign ((segid A and resid 17 and name HB# )) ((segid A and resid 18 and name HG2# )) 4.06 2.26 0.00
assign ((segid A and resid 17 and name HB# )) ((segid A and resid 19 and name HN )) 4.75 2.95 0.00
assign ((segid A and resid 18 and name HA )) ((segid A and resid 26 and name HB# )) 4.18 2.38 0.00
assign ((segid A and resid 18 and name HA )) ((segid A and resid 26 and name HG# )) 4.77 2.97 0.00
assign ((segid A and resid 18 and name HG2# )) ((segid A and resid 24 and name HG# )) 5.29 3.49 0.00
assign ((segid A and resid 18 and name HG2# )) ((segid A and resid 26 and name HG# )) 3.37 1.57 0.00
assign ((segid A and resid 18 and name HD1# )) ((segid A and resid 26 and name HG# )) 4.13 2.33 0.00
assign ((segid A and resid 18 and name HD1# )) ((segid A and resid 70 and name HG# )) 4.94 3.14 0.00
assign ((segid A and resid 19 and name HN )) ((segid A and resid 19 and name HG# )) 4.65 2.85 0.00
assign ((segid A and resid 19 and name HN )) ((segid A and resid 26 and name HG# )) 5.81 4.01 0.00
assign ((segid A and resid 19 and name HA )) ((segid A and resid 19 and name HG# )) 3.55 1.75 0.00
assign ((segid A and resid 20 and name HN )) ((segid A and resid 21 and name HB# )) 5.81 4.01 0.00
assign ((segid A and resid 20 and name HA )) ((segid A and resid 20 and name HG# )) 3.33 1.53 0.00
assign ((segid A and resid 20 and name HB# )) ((segid A and resid 21 and name HN )) 3.28 1.48 0.00
assign ((segid A and resid 20 and name HB# )) ((segid A and resid 22 and name HN )) 4.31 2.51 0.00
assign ((segid A and resid 21 and name HN )) ((segid A and resid 21 and name HB# )) 3.28 1.48 0.00
assign ((segid A and resid 21 and name HN )) ((segid A and resid 22 and name HB# )) 4.16 2.36 0.00
assign ((segid A and resid 21 and name HA )) ((segid A and resid 22 and name HB# )) 5.81 4.01 0.00
assign ((segid A and resid 21 and name HB# )) ((segid A and resid 24 and name HN )) 5.81 4.01 0.00
assign ((segid A and resid 22 and name HN )) ((segid A and resid 22 and name HB# )) 3.34 1.54 0.00
assign ((segid A and resid 22 and name HN )) ((segid A and resid 22 and name HG# )) 4.88 3.08 0.00
assign ((segid A and resid 22 and name HA )) ((segid A and resid 22 and name HG# )) 3.32 1.52 0.00
assign ((segid A and resid 22 and name HB# )) ((segid A and resid 23 and name HN )) 3.59 1.79 0.00
assign ((segid A and resid 22 and name HG# )) ((segid A and resid 23 and name HN )) 4.37 2.57 0.00
assign ((segid A and resid 24 and name HN )) ((segid A and resid 24 and name HG# )) 4.41 2.61 0.00
assign ((segid A and resid 24 and name HA )) ((segid A and resid 24 and name HG# )) 3.57 1.77 0.00
assign ((segid A and resid 24 and name HG# )) ((segid A and resid 29 and name HN )) 4.36 2.56 0.00
assign ((segid A and resid 24 and name HG# )) ((segid A and resid 29 and name HB# )) 3.65 1.85 0.00
assign ((segid A and resid 24 and name HG# )) ((segid A and resid 32 and name HN )) 5.05 3.25 0.00
assign ((segid A and resid 24 and name HG# )) ((segid A and resid 33 and name HN )) 4.31 2.51 0.00
assign ((segid A and resid 25 and name HA )) ((segid A and resid 26 and name HB# )) 5.30 3.50 0.00
assign ((segid A and resid 26 and name HN )) ((segid A and resid 26 and name HB# )) 3.56 1.76 0.00
assign ((segid A and resid 26 and name HN )) ((segid A and resid 26 and name HG# )) 4.42 2.62 0.00
assign ((segid A and resid 26 and name HA )) ((segid A and resid 26 and name HG# )) 3.65 1.85 0.00
assign ((segid A and resid 26 and name HB# )) ((segid A and resid 27 and name HG1# )) 5.61 3.81 0.00
assign ((segid A and resid 26 and name HB# )) ((segid A and resid 29 and name HN )) 5.81 4.01 0.00
assign ((segid A and resid 26 and name HG# )) ((segid A and resid 27 and name HN )) 4.41 2.61 0.00
assign ((segid A and resid 26 and name HG# )) ((segid A and resid 29 and name HN )) 5.81 4.01 0.00
assign ((segid A and resid 27 and name HN )) ((segid A and resid 27 and name HG1# )) 5.81 4.01 0.00
assign ((segid A and resid 27 and name HA )) ((segid A and resid 66 and name HG# )) 5.34 3.54 0.00
assign ((segid A and resid 27 and name HA )) ((segid A and resid 67 and name HD# )) 4.95 3.15 0.00
assign ((segid A and resid 27 and name HG2# )) ((segid A and resid 27 and name HG1# )) 3.17 1.37 0.00
assign ((segid A and resid 27 and name HG2# )) ((segid A and resid 67 and name HD# )) 5.75 3.95 0.00
assign ((segid A and resid 27 and name HG1# )) ((segid A and resid 71 and name HN )) 5.81 4.01 0.00
assign ((segid A and resid 27 and name HD1# )) ((segid A and resid 67 and name HD# )) 2.68 0.88 0.00
assign ((segid A and resid 27 and name HD1# )) ((segid A and resid 70 and name HB# )) 5.40 3.60 0.00
assign ((segid A and resid 27 and name HD1# )) ((segid A and resid 70 and name HG# )) 5.81 4.01 0.00
assign ((segid A and resid 29 and name HN )) ((segid A and resid 66 and name HG# )) 4.13 2.33 0.00
assign ((segid A and resid 29 and name HB# )) ((segid A and resid 33 and name HB# )) 4.28 2.48 0.00
assign ((segid A and resid 30 and name HN )) ((segid A and resid 33 and name HB# )) 4.41 2.61 0.00
assign ((segid A and resid 30 and name HN )) ((segid A and resid 67 and name HD# )) 5.77 3.97 0.00
assign ((segid A and resid 30 and name HA )) ((segid A and resid 33 and name HB# )) 5.81 4.01 0.00
assign ((segid A and resid 30 and name HA )) ((segid A and resid 66 and name HG# )) 4.12 2.32 0.00
assign ((segid A and resid 31 and name HN )) ((segid A and resid 33 and name HB# )) 5.03 3.23 0.00
assign ((segid A and resid 32 and name HN )) ((segid A and resid 33 and name HB# )) 5.81 4.01 0.00
assign ((segid A and resid 33 and name HN )) ((segid A and resid 33 and name HB# )) 3.30 1.50 0.00
assign ((segid A and resid 33 and name HB# )) ((segid A and resid 34 and name HN )) 4.05 2.25 0.00
assign ((segid A and resid 33 and name HB# )) ((segid A and resid 35 and name HN )) 5.72 3.92 0.00
assign ((segid A and resid 33 and name HE# )) ((segid A and resid 37 and name HG# )) 3.92 2.12 0.00
assign ((segid A and resid 33 and name HE# )) ((segid A and resid 59 and name HD# )) 3.35 1.55 0.00
assign ((segid A and resid 33 and name HD# )) ((segid A and resid 37 and name HG# )) 4.43 2.63 0.00
assign ((segid A and resid 33 and name HD# )) ((segid A and resid 59 and name HD# )) 4.17 2.37 0.00
assign ((segid A and resid 34 and name HN )) ((segid A and resid 38 and name HG# )) 5.63 3.83 0.00
assign ((segid A and resid 34 and name HB# )) ((segid A and resid 38 and name HG# )) 5.38 3.58 0.00
assign ((segid A and resid 34 and name HE# )) ((segid A and resid 37 and name HG# )) 5.11 3.31 0.00
assign ((segid A and resid 34 and name HE# )) ((segid A and resid 38 and name HG# )) 4.34 2.54 0.00
assign ((segid A and resid 34 and name HE# )) ((segid A and resid 60 and name HB# )) 5.34 3.54 0.00
assign ((segid A and resid 34 and name HE# )) ((segid A and resid 63 and name HB# )) 4.23 2.43 0.00
assign ((segid A and resid 34 and name HD# )) ((segid A and resid 38 and name HG# )) 5.60 3.80 0.00
assign ((segid A and resid 34 and name HD# )) ((segid A and resid 59 and name HD# )) 5.13 3.33 0.00
assign ((segid A and resid 34 and name HD# )) ((segid A and resid 63 and name HB# )) 4.17 2.37 0.00
assign ((segid A and resid 35 and name HN )) ((segid A and resid 35 and name HB# )) 3.14 1.34 0.00
assign ((segid A and resid 36 and name HN )) ((segid A and resid 37 and name HG# )) 4.85 3.05 0.00
assign ((segid A and resid 37 and name HN )) ((segid A and resid 37 and name HG# )) 3.40 1.60 0.00
assign ((segid A and resid 37 and name HN )) ((segid A and resid 38 and name HG# )) 4.65 2.85 0.00
assign ((segid A and resid 37 and name HN )) ((segid A and resid 59 and name HD# )) 4.49 2.69 0.00
assign ((segid A and resid 37 and name HA )) ((segid A and resid 37 and name HG# )) 2.77 0.97 0.00
assign ((segid A and resid 37 and name HB )) ((segid A and resid 38 and name HG# )) 4.93 3.13 0.00
assign ((segid A and resid 37 and name HB )) ((segid A and resid 59 and name HD# )) 4.41 2.61 0.00
assign ((segid A and resid 37 and name HG# )) ((segid A and resid 38 and name HN )) 3.19 1.39 0.00
assign ((segid A and resid 37 and name HG# )) ((segid A and resid 38 and name HA )) 3.47 1.67 0.00
assign ((segid A and resid 37 and name HG# )) ((segid A and resid 40 and name HB )) 5.24 3.44 0.00
assign ((segid A and resid 37 and name HG# )) ((segid A and resid 40 and name HG2# )) 4.87 3.07 0.00
assign ((segid A and resid 37 and name HG# )) ((segid A and resid 41 and name HN )) 3.85 2.05 0.00
assign ((segid A and resid 37 and name HG# )) ((segid A and resid 41 and name HB# )) 3.95 2.15 0.00
assign ((segid A and resid 37 and name HG# )) ((segid A and resid 41 and name HD# )) 2.99 1.19 0.00
assign ((segid A and resid 37 and name HG# )) ((segid A and resid 42 and name HN )) 5.25 3.45 0.00
assign ((segid A and resid 37 and name HG# )) ((segid A and resid 56 and name HA )) 5.51 3.71 0.00
assign ((segid A and resid 37 and name HG# )) ((segid A and resid 59 and name HN )) 5.52 3.72 0.00
assign ((segid A and resid 37 and name HG# )) ((segid A and resid 59 and name HD# )) 3.18 1.38 0.00
assign ((segid A and resid 38 and name HN )) ((segid A and resid 38 and name HG# )) 3.11 1.31 0.00
assign ((segid A and resid 38 and name HN )) ((segid A and resid 39 and name HB# )) 5.04 3.24 0.00
assign ((segid A and resid 38 and name HN )) ((segid A and resid 41 and name HD# )) 5.18 3.38 0.00
assign ((segid A and resid 38 and name HA )) ((segid A and resid 41 and name HD# )) 4.41 2.61 0.00
assign ((segid A and resid 38 and name HA )) ((segid A and resid 42 and name HG# )) 5.92 4.12 0.00
assign ((segid A and resid 38 and name HG# )) ((segid A and resid 39 and name HN )) 5.92 4.12 0.00
assign ((segid A and resid 38 and name HG# )) ((segid A and resid 59 and name HB2 )) 5.57 3.77 0.00
assign ((segid A and resid 38 and name HG# )) ((segid A and resid 59 and name HD# )) 3.42 1.62 0.00
assign ((segid A and resid 39 and name HN )) ((segid A and resid 39 and name HB# )) 3.32 1.52 0.00
assign ((segid A and resid 39 and name HN )) ((segid A and resid 42 and name HG# )) 5.92 4.12 0.00
assign ((segid A and resid 39 and name HA )) ((segid A and resid 42 and name HG# )) 5.92 4.12 0.00
assign ((segid A and resid 39 and name HB# )) ((segid A and resid 40 and name HN )) 3.30 1.50 0.00
assign ((segid A and resid 39 and name HB# )) ((segid A and resid 40 and name HG2# )) 5.81 4.01 0.00
assign ((segid A and resid 39 and name HB# )) ((segid A and resid 41 and name HN )) 5.78 3.98 0.00
assign ((segid A and resid 39 and name HB# )) ((segid A and resid 42 and name HN )) 5.81 4.01 0.00
assign ((segid A and resid 40 and name HN )) ((segid A and resid 41 and name HB# )) 5.33 3.53 0.00
assign ((segid A and resid 40 and name HN )) ((segid A and resid 41 and name HD# )) 5.35 3.55 0.00
assign ((segid A and resid 40 and name HN )) ((segid A and resid 42 and name HG# )) 5.56 3.76 0.00
assign ((segid A and resid 40 and name HN )) ((segid A and resid 43 and name HB# )) 5.84 4.04 0.00
assign ((segid A and resid 40 and name HB )) ((segid A and resid 41 and name HD# )) 5.92 4.12 0.00
assign ((segid A and resid 40 and name HG2# )) ((segid A and resid 41 and name HB# )) 4.60 2.80 0.00
assign ((segid A and resid 40 and name HG2# )) ((segid A and resid 41 and name HD# )) 5.10 3.30 0.00
assign ((segid A and resid 40 and name HG2# )) ((segid A and resid 43 and name HB# )) 5.81 4.01 0.00
assign ((segid A and resid 40 and name HG2# )) ((segid A and resid 43 and name HG# )) 5.81 4.01 0.00
assign ((segid A and resid 41 and name HN )) ((segid A and resid 41 and name HB# )) 3.06 1.26 0.00
assign ((segid A and resid 41 and name HN )) ((segid A and resid 41 and name HD# )) 3.85 2.05 0.00
assign ((segid A and resid 41 and name HA )) ((segid A and resid 41 and name HD# )) 3.95 2.15 0.00
assign ((segid A and resid 41 and name HG )) ((segid A and resid 59 and name HD# )) 5.92 4.12 0.00
assign ((segid A and resid 41 and name HD# )) ((segid A and resid 42 and name HN )) 4.31 2.51 0.00
assign ((segid A and resid 41 and name HD# )) ((segid A and resid 52 and name HA )) 5.92 4.12 0.00
assign ((segid A and resid 41 and name HD# )) ((segid A and resid 55 and name HN )) 5.10 3.30 0.00
assign ((segid A and resid 41 and name HD# )) ((segid A and resid 55 and name HA )) 4.81 3.01 0.00
assign ((segid A and resid 41 and name HD# )) ((segid A and resid 56 and name HN )) 5.92 4.12 0.00
assign ((segid A and resid 42 and name HN )) ((segid A and resid 42 and name HG# )) 3.52 1.72 0.00
assign ((segid A and resid 42 and name HN )) ((segid A and resid 44 and name HB# )) 5.81 4.01 0.00
assign ((segid A and resid 42 and name HA )) ((segid A and resid 42 and name HG# )) 3.12 1.32 0.00
assign ((segid A and resid 42 and name HA )) ((segid A and resid 52 and name HG# )) 4.50 2.70 0.00
assign ((segid A and resid 42 and name HB )) ((segid A and resid 52 and name HG# )) 5.92 4.12 0.00
assign ((segid A and resid 42 and name HG# )) ((segid A and resid 43 and name HN )) 3.96 2.16 0.00
assign ((segid A and resid 42 and name HG# )) ((segid A and resid 43 and name HB# )) 5.73 3.93 0.00
assign ((segid A and resid 42 and name HG# )) ((segid A and resid 43 and name HG# )) 5.73 3.93 0.00
assign ((segid A and resid 42 and name HG# )) ((segid A and resid 44 and name HN )) 4.47 2.67 0.00
assign ((segid A and resid 42 and name HG# )) ((segid A and resid 44 and name HB# )) 5.73 3.93 0.00
assign ((segid A and resid 42 and name HG# )) ((segid A and resid 52 and name HN )) 5.92 4.12 0.00
assign ((segid A and resid 42 and name HG# )) ((segid A and resid 56 and name HG1# )) 5.66 3.86 0.00
assign ((segid A and resid 43 and name HN )) ((segid A and resid 44 and name HB# )) 5.33 3.53 0.00
assign ((segid A and resid 43 and name HA )) ((segid A and resid 43 and name HG# )) 3.55 1.75 0.00
assign ((segid A and resid 44 and name HN )) ((segid A and resid 44 and name HB# )) 3.23 1.43 0.00
assign ((segid A and resid 44 and name HB# )) ((segid A and resid 46 and name HN )) 5.87 4.07 0.00
assign ((segid A and resid 45 and name HN )) ((segid A and resid 45 and name HG# )) 4.73 2.93 0.00
assign ((segid A and resid 45 and name HN )) ((segid A and resid 46 and name HB# )) 5.81 4.01 0.00
assign ((segid A and resid 45 and name HA )) ((segid A and resid 45 and name HG# )) 3.71 1.91 0.00
assign ((segid A and resid 45 and name HG# )) ((segid A and resid 46 and name HN )) 4.21 2.41 0.00
assign ((segid A and resid 45 and name HG# )) ((segid A and resid 46 and name HA )) 4.40 2.60 0.00
assign ((segid A and resid 45 and name HG# )) ((segid A and resid 46 and name HB# )) 5.61 3.81 0.00
assign ((segid A and resid 45 and name HG# )) ((segid A and resid 46 and name HG )) 5.81 4.01 0.00
assign ((segid A and resid 45 and name HG# )) ((segid A and resid 47 and name HN )) 3.73 1.93 0.00
assign ((segid A and resid 45 and name HG# )) ((segid A and resid 48 and name HN )) 5.81 4.01 0.00
assign ((segid A and resid 45 and name HG# )) ((segid A and resid 51 and name HG# )) 4.15 2.35 0.00
assign ((segid A and resid 45 and name HG# )) ((segid A and resid 52 and name HN )) 5.84 4.04 0.00
assign ((segid A and resid 46 and name HN )) ((segid A and resid 46 and name HB# )) 3.08 1.28 0.00
assign ((segid A and resid 46 and name HA )) ((segid A and resid 46 and name HD# )) 3.78 1.98 0.00
assign ((segid A and resid 46 and name HB# )) ((segid A and resid 47 and name HA )) 5.81 4.01 0.00
assign ((segid A and resid 46 and name HD# )) ((segid A and resid 48 and name HN )) 5.47 3.67 0.00
assign ((segid A and resid 47 and name HN )) ((segid A and resid 51 and name HG# )) 3.72 1.92 0.00
assign ((segid A and resid 47 and name HA )) ((segid A and resid 48 and name HB# )) 5.81 4.01 0.00
assign ((segid A and resid 47 and name HA )) ((segid A and resid 51 and name HG# )) 4.87 3.07 0.00
assign ((segid A and resid 47 and name HB )) ((segid A and resid 51 and name HG# )) 4.45 2.65 0.00
assign ((segid A and resid 47 and name HB )) ((segid A and resid 52 and name HG# )) 5.52 3.72 0.00
assign ((segid A and resid 47 and name HG2# )) ((segid A and resid 48 and name HB# )) 3.72 1.92 0.00
assign ((segid A and resid 47 and name HG2# )) ((segid A and resid 51 and name HG# )) 2.41 0.61 0.00
assign ((segid A and resid 48 and name HN )) ((segid A and resid 48 and name HB# )) 3.41 1.61 0.00
assign ((segid A and resid 48 and name HN )) ((segid A and resid 48 and name HG# )) 4.08 2.28 0.00
assign ((segid A and resid 48 and name HN )) ((segid A and resid 51 and name HG# )) 4.66 2.86 0.00
assign ((segid A and resid 48 and name HN )) ((segid A and resid 52 and name HG# )) 4.73 2.93 0.00
assign ((segid A and resid 48 and name HB# )) ((segid A and resid 49 and name HN )) 3.64 1.84 0.00
assign ((segid A and resid 48 and name HB# )) ((segid A and resid 50 and name HN )) 3.96 2.16 0.00
assign ((segid A and resid 48 and name HB# )) ((segid A and resid 50 and name HB# )) 4.57 2.77 0.00
assign ((segid A and resid 48 and name HB# )) ((segid A and resid 51 and name HG# )) 3.02 1.22 0.00
assign ((segid A and resid 48 and name HG# )) ((segid A and resid 49 and name HN )) 3.83 2.03 0.00
assign ((segid A and resid 48 and name HG# )) ((segid A and resid 50 and name HN )) 4.76 2.96 0.00
assign ((segid A and resid 48 and name HG# )) ((segid A and resid 50 and name HA )) 4.54 2.74 0.00
assign ((segid A and resid 48 and name HG# )) ((segid A and resid 51 and name HN )) 4.24 2.44 0.00
assign ((segid A and resid 48 and name HG# )) ((segid A and resid 51 and name HG# )) 3.37 1.57 0.00
assign ((segid A and resid 48 and name HG# )) ((segid A and resid 52 and name HN )) 4.79 2.99 0.00
assign ((segid A and resid 48 and name HE# )) ((segid A and resid 49 and name HN )) 5.81 4.01 0.00
assign ((segid A and resid 48 and name HE# )) ((segid A and resid 49 and name HA )) 5.81 4.01 0.00
assign ((segid A and resid 48 and name HE# )) ((segid A and resid 50 and name HN )) 4.16 2.36 0.00
assign ((segid A and resid 48 and name HE# )) ((segid A and resid 50 and name HB# )) 4.62 2.82 0.00
assign ((segid A and resid 49 and name HN )) ((segid A and resid 49 and name HB# )) 3.53 1.73 0.00
assign ((segid A and resid 49 and name HA )) ((segid A and resid 52 and name HG# )) 5.15 3.35 0.00
assign ((segid A and resid 49 and name HA )) ((segid A and resid 53 and name HB# )) 4.67 2.87 0.00
assign ((segid A and resid 49 and name HB# )) ((segid A and resid 50 and name HN )) 3.56 1.76 0.00
assign ((segid A and resid 49 and name HB# )) ((segid A and resid 50 and name HB# )) 5.09 3.29 0.00
assign ((segid A and resid 49 and name HB# )) ((segid A and resid 51 and name HN )) 5.81 4.01 0.00
assign ((segid A and resid 49 and name HB# )) ((segid A and resid 52 and name HG# )) 4.89 3.09 0.00
assign ((segid A and resid 49 and name HB# )) ((segid A and resid 53 and name HN )) 5.81 4.01 0.00
assign ((segid A and resid 49 and name HB# )) ((segid A and resid 53 and name HD# )) 4.19 2.39 0.00
assign ((segid A and resid 50 and name HN )) ((segid A and resid 51 and name HG# )) 4.91 3.11 0.00
assign ((segid A and resid 50 and name HN )) ((segid A and resid 52 and name HG# )) 4.38 2.58 0.00
assign ((segid A and resid 50 and name HB# )) ((segid A and resid 51 and name HG# )) 3.62 1.82 0.00
assign ((segid A and resid 51 and name HN )) ((segid A and resid 51 and name HG# )) 3.21 1.41 0.00
assign ((segid A and resid 51 and name HN )) ((segid A and resid 52 and name HG# )) 4.77 2.97 0.00
assign ((segid A and resid 51 and name HA )) ((segid A and resid 51 and name HG# )) 2.78 0.98 0.00
assign ((segid A and resid 51 and name HA )) ((segid A and resid 54 and name HB# )) 4.85 3.05 0.00
assign ((segid A and resid 51 and name HB )) ((segid A and resid 52 and name HG# )) 4.56 2.76 0.00
assign ((segid A and resid 51 and name HG# )) ((segid A and resid 52 and name HN )) 3.46 1.66 0.00
assign ((segid A and resid 51 and name HG# )) ((segid A and resid 52 and name HA )) 5.92 4.12 0.00
assign ((segid A and resid 51 and name HG# )) ((segid A and resid 54 and name HN )) 5.92 4.12 0.00
assign ((segid A and resid 51 and name HG# )) ((segid A and resid 54 and name HG# )) 5.06 3.26 0.00
assign ((segid A and resid 51 and name HG# )) ((segid A and resid 54 and name HE# )) 5.92 4.12 0.00
assign ((segid A and resid 52 and name HN )) ((segid A and resid 52 and name HG# )) 3.63 1.83 0.00
assign ((segid A and resid 52 and name HN )) ((segid A and resid 53 and name HB# )) 5.14 3.34 0.00
assign ((segid A and resid 52 and name HN )) ((segid A and resid 54 and name HD# )) 5.60 3.80 0.00
assign ((segid A and resid 52 and name HA )) ((segid A and resid 52 and name HG# )) 3.02 1.22 0.00
assign ((segid A and resid 52 and name HG# )) ((segid A and resid 53 and name HN )) 3.44 1.64 0.00
assign ((segid A and resid 52 and name HG# )) ((segid A and resid 53 and name HA )) 3.63 1.83 0.00
assign ((segid A and resid 52 and name HG# )) ((segid A and resid 53 and name HB# )) 3.96 2.16 0.00
assign ((segid A and resid 52 and name HG# )) ((segid A and resid 54 and name HN )) 4.78 2.98 0.00
assign ((segid A and resid 52 and name HG# )) ((segid A and resid 54 and name HD# )) 5.34 3.54 0.00
assign ((segid A and resid 52 and name HG# )) ((segid A and resid 55 and name HN )) 4.81 3.01 0.00
assign ((segid A and resid 52 and name HG# )) ((segid A and resid 56 and name HB )) 5.92 4.12 0.00
assign ((segid A and resid 53 and name HN )) ((segid A and resid 53 and name HB# )) 3.19 1.39 0.00
assign ((segid A and resid 53 and name HN )) ((segid A and resid 53 and name HD# )) 3.88 2.08 0.00
assign ((segid A and resid 53 and name HN )) ((segid A and resid 54 and name HD# )) 5.39 3.59 0.00
assign ((segid A and resid 53 and name HN )) ((segid A and resid 56 and name HG1# )) 4.64 2.84 0.00
assign ((segid A and resid 53 and name HA )) ((segid A and resid 53 and name HD# )) 3.45 1.65 0.00
assign ((segid A and resid 53 and name HB# )) ((segid A and resid 53 and name HD# )) 2.67 0.87 0.00
assign ((segid A and resid 53 and name HD# )) ((segid A and resid 54 and name HN )) 5.27 3.47 0.00
assign ((segid A and resid 53 and name HD# )) ((segid A and resid 56 and name HA )) 5.92 4.12 0.00
assign ((segid A and resid 53 and name HD# )) ((segid A and resid 56 and name HB )) 4.14 2.34 0.00
assign ((segid A and resid 53 and name HD# )) ((segid A and resid 56 and name HG1# )) 3.97 2.17 0.00
assign ((segid A and resid 53 and name HD# )) ((segid A and resid 57 and name HA )) 4.80 3.00 0.00
assign ((segid A and resid 53 and name HD# )) ((segid A and resid 57 and name HG# )) 3.70 1.90 0.00
assign ((segid A and resid 54 and name HN )) ((segid A and resid 54 and name HG# )) 3.73 1.93 0.00
assign ((segid A and resid 54 and name HN )) ((segid A and resid 56 and name HG1# )) 5.81 4.01 0.00
assign ((segid A and resid 54 and name HN )) ((segid A and resid 57 and name HG# )) 4.90 3.10 0.00
assign ((segid A and resid 54 and name HA )) ((segid A and resid 54 and name HG# )) 3.63 1.83 0.00
assign ((segid A and resid 54 and name HA )) ((segid A and resid 57 and name HG# )) 5.81 4.01 0.00
assign ((segid A and resid 54 and name HB# )) ((segid A and resid 54 and name HE# )) 4.64 2.84 0.00
assign ((segid A and resid 54 and name HG# )) ((segid A and resid 54 and name HE# )) 2.95 1.15 0.00
assign ((segid A and resid 54 and name HG# )) ((segid A and resid 55 and name HN )) 5.81 4.01 0.00
assign ((segid A and resid 55 and name HN )) ((segid A and resid 57 and name HG# )) 5.31 3.51 0.00
assign ((segid A and resid 56 and name HA )) ((segid A and resid 59 and name HD# )) 5.92 4.12 0.00
assign ((segid A and resid 56 and name HG2# )) ((segid A and resid 57 and name HG# )) 5.81 4.01 0.00
assign ((segid A and resid 56 and name HG2# )) ((segid A and resid 59 and name HD# )) 4.29 2.49 0.00
assign ((segid A and resid 56 and name HG1# )) ((segid A and resid 57 and name HN )) 4.04 2.24 0.00
assign ((segid A and resid 59 and name HN )) ((segid A and resid 59 and name HD# )) 4.16 2.36 0.00
assign ((segid A and resid 59 and name HA )) ((segid A and resid 59 and name HD# )) 3.81 2.01 0.00
assign ((segid A and resid 59 and name HB2 )) ((segid A and resid 61 and name HG# )) 5.81 4.01 0.00
assign ((segid A and resid 59 and name HD# )) ((segid A and resid 60 and name HN )) 4.82 3.02 0.00
assign ((segid A and resid 59 and name HD# )) ((segid A and resid 62 and name HB# )) 4.09 2.29 0.00
assign ((segid A and resid 60 and name HN )) ((segid A and resid 61 and name HB# )) 5.78 3.98 0.00
assign ((segid A and resid 60 and name HN )) ((segid A and resid 61 and name HG# )) 5.81 4.01 0.00
assign ((segid A and resid 60 and name HB# )) ((segid A and resid 61 and name HG# )) 5.61 3.81 0.00
assign ((segid A and resid 60 and name HB# )) ((segid A and resid 62 and name HN )) 5.81 4.01 0.00
assign ((segid A and resid 60 and name HB# )) ((segid A and resid 63 and name HN )) 5.81 4.01 0.00
assign ((segid A and resid 60 and name HB# )) ((segid A and resid 63 and name HE# )) 5.06 3.26 0.00
assign ((segid A and resid 60 and name HB# )) ((segid A and resid 63 and name HD# )) 5.18 3.38 0.00
assign ((segid A and resid 61 and name HN )) ((segid A and resid 61 and name HB# )) 3.41 1.61 0.00
assign ((segid A and resid 61 and name HN )) ((segid A and resid 61 and name HG# )) 3.49 1.69 0.00
assign ((segid A and resid 61 and name HA )) ((segid A and resid 61 and name HG# )) 3.56 1.76 0.00
assign ((segid A and resid 61 and name HB# )) ((segid A and resid 62 and name HN )) 4.03 2.23 0.00
assign ((segid A and resid 61 and name HG# )) ((segid A and resid 62 and name HN )) 4.10 2.30 0.00
assign ((segid A and resid 61 and name HG# )) ((segid A and resid 65 and name HB# )) 5.61 3.81 0.00
assign ((segid A and resid 62 and name HN )) ((segid A and resid 63 and name HB# )) 5.81 4.01 0.00
assign ((segid A and resid 62 and name HN )) ((segid A and resid 65 and name HB# )) 5.32 3.52 0.00
assign ((segid A and resid 62 and name HA )) ((segid A and resid 65 and name HB# )) 5.41 3.61 0.00
assign ((segid A and resid 63 and name HN )) ((segid A and resid 64 and name HB# )) 5.81 4.01 0.00
assign ((segid A and resid 63 and name HN )) ((segid A and resid 65 and name HB# )) 5.81 4.01 0.00
assign ((segid A and resid 63 and name HB# )) ((segid A and resid 64 and name HN )) 3.57 1.77 0.00
assign ((segid A and resid 63 and name HE# )) ((segid A and resid 67 and name HD# )) 4.83 3.03 0.00
assign ((segid A and resid 65 and name HN )) ((segid A and resid 66 and name HG# )) 5.61 3.81 0.00
assign ((segid A and resid 66 and name HN )) ((segid A and resid 66 and name HG# )) 3.10 1.30 0.00
assign ((segid A and resid 66 and name HA )) ((segid A and resid 66 and name HG# )) 2.95 1.15 0.00
assign ((segid A and resid 66 and name HG# )) ((segid A and resid 67 and name HN )) 3.71 1.91 0.00
assign ((segid A and resid 66 and name HG# )) ((segid A and resid 67 and name HA )) 4.23 2.43 0.00
assign ((segid A and resid 66 and name HG# )) ((segid A and resid 69 and name HN )) 5.92 4.12 0.00
assign ((segid A and resid 66 and name HG# )) ((segid A and resid 70 and name HG# )) 3.12 1.32 0.00
assign ((segid A and resid 67 and name HN )) ((segid A and resid 70 and name HB# )) 5.49 3.69 0.00
assign ((segid A and resid 67 and name HN )) ((segid A and resid 70 and name HG# )) 5.81 4.01 0.00
assign ((segid A and resid 67 and name HA )) ((segid A and resid 67 and name HD# )) 4.81 3.01 0.00
assign ((segid A and resid 67 and name HB2 )) ((segid A and resid 68 and name HD# )) 5.81 4.01 0.00
assign ((segid A and resid 67 and name HD# )) ((segid A and resid 70 and name HN )) 5.81 4.01 0.00
assign ((segid A and resid 68 and name HN )) ((segid A and resid 68 and name HG# )) 3.64 1.84 0.00
assign ((segid A and resid 68 and name HN )) ((segid A and resid 68 and name HD# )) 4.99 3.19 0.00
assign ((segid A and resid 68 and name HN )) ((segid A and resid 70 and name HB# )) 5.81 4.01 0.00
assign ((segid A and resid 68 and name HA )) ((segid A and resid 68 and name HG# )) 3.03 1.23 0.00
assign ((segid A and resid 68 and name HA )) ((segid A and resid 68 and name HD# )) 5.43 3.63 0.00
assign ((segid A and resid 68 and name HB1 )) ((segid A and resid 68 and name HD# )) 3.39 1.59 0.00
assign ((segid A and resid 68 and name HB1 )) ((segid A and resid 69 and name HA# )) 4.46 2.66 0.00
assign ((segid A and resid 68 and name HG# )) ((segid A and resid 69 and name HN )) 4.68 2.88 0.00
assign ((segid A and resid 69 and name HN )) ((segid A and resid 69 and name HA# )) 2.58 0.78 0.00
assign ((segid A and resid 69 and name HN )) ((segid A and resid 70 and name HB# )) 5.81 4.01 0.00
assign ((segid A and resid 69 and name HN )) ((segid A and resid 70 and name HG# )) 4.75 2.95 0.00
assign ((segid A and resid 69 and name HA# )) ((segid A and resid 70 and name HN )) 3.10 1.30 0.00
assign ((segid A and resid 69 and name HA# )) ((segid A and resid 70 and name HA )) 5.24 3.44 0.00
assign ((segid A and resid 70 and name HN )) ((segid A and resid 70 and name HB# )) 3.05 1.25 0.00
assign ((segid A and resid 70 and name HN )) ((segid A and resid 70 and name HG# )) 3.81 2.01 0.00
assign ((segid A and resid 70 and name HA )) ((segid A and resid 70 and name HG# )) 3.08 1.28 0.00
assign ((segid A and resid 71 and name HN )) ((segid A and resid 71 and name HB# )) 3.06 1.26 0.00
assign ((segid A and resid 71 and name HN )) ((segid A and resid 72 and name HG# )) 5.81 4.01 0.00
assign ((segid A and resid 71 and name HA )) ((segid A and resid 72 and name HB# )) 4.87 3.07 0.00
assign ((segid A and resid 71 and name HB# )) ((segid A and resid 72 and name HN )) 3.89 2.09 0.00
assign ((segid A and resid 71 and name HB# )) ((segid A and resid 73 and name HN )) 5.61 3.81 0.00
assign ((segid A and resid 72 and name HN )) ((segid A and resid 72 and name HB# )) 3.48 1.68 0.00
assign ((segid A and resid 72 and name HN )) ((segid A and resid 72 and name HG# )) 4.16 2.36 0.00
assign ((segid A and resid 72 and name HA )) ((segid A and resid 72 and name HG# )) 3.61 1.81 0.00
assign ((segid A and resid 72 and name HA )) ((segid A and resid 72 and name HD# )) 5.81 4.01 0.00
assign ((segid A and resid 72 and name HB# )) ((segid A and resid 73 and name HN )) 3.90 2.10 0.00
assign ((segid A and resid 72 and name HB# )) ((segid A and resid 73 and name HD# )) 5.73 3.93 0.00
assign ((segid A and resid 73 and name HN )) ((segid A and resid 73 and name HB# )) 3.00 1.20 0.00
assign ((segid A and resid 73 and name HN )) ((segid A and resid 73 and name HD# )) 4.53 2.73 0.00
assign ((segid A and resid 73 and name HN )) ((segid A and resid 74 and name HG# )) 5.81 4.01 0.00
assign ((segid A and resid 73 and name HA )) ((segid A and resid 73 and name HD# )) 3.48 1.68 0.00
assign ((segid A and resid 73 and name HA )) ((segid A and resid 74 and name HB# )) 5.34 3.54 0.00
assign ((segid A and resid 73 and name HB# )) ((segid A and resid 73 and name HD# )) 2.70 0.90 0.00
assign ((segid A and resid 73 and name HB# )) ((segid A and resid 74 and name HN )) 4.12 2.32 0.00
assign ((segid A and resid 73 and name HB# )) ((segid A and resid 74 and name HG# )) 5.10 3.30 0.00
assign ((segid A and resid 73 and name HD# )) ((segid A and resid 74 and name HN )) 4.67 2.87 0.00
assign ((segid A and resid 73 and name HD# )) ((segid A and resid 75 and name HN )) 5.80 4.00 0.00
assign ((segid A and resid 73 and name HD# )) ((segid A and resid 75 and name HB# )) 5.73 3.93 0.00
assign ((segid A and resid 74 and name HN )) ((segid A and resid 74 and name HB# )) 3.19 1.39 0.00
assign ((segid A and resid 74 and name HN )) ((segid A and resid 74 and name HG# )) 3.71 1.91 0.00
assign ((segid A and resid 74 and name HA )) ((segid A and resid 74 and name HG# )) 3.53 1.73 0.00
assign ((segid A and resid 74 and name HB# )) ((segid A and resid 75 and name HA )) 5.81 4.01 0.00
list of removed NOE constraints
21-> ILE A 56 HA - ILE A 56 HG2# 1.80 4.05 # NoRestrctn I [2.63 3.78] -- intra
22-> LEU A 46 HN - THR A 47 HA 1.80 5.53 # NoRestrctn S [2.00 3.99] -- sequential
29-> ASP A 25 HN - LYS A 26 HA 1.80 6.00 # NoRestrctn S [2.00 3.99] -- sequential
36-> ASP A 60 HN - GLU A 61 HA 1.80 5.50 # NoRestrctn S [2.00 3.99] -- sequential
127-> ALA A 62 HN - ALA A 62 HB# 1.80 4.09 # NoRestrctn I [2.66 3.68] -- intra
143-> ILE A 56 HN - ILE A 56 HG2# 1.80 6.00 # NoRestrctn I [2.04 4.91] -- intra
151-> ALA A 29 HN - ALA A 29 HB# 1.80 3.86 # NoRestrctn I [2.66 3.68] -- intra
161-> ILE A 18 HA - ILE A 18 HG2# 1.80 3.81 # NoRestrctn I [2.63 3.78] -- intra
162-> ILE A 18 HG2# - ILE A 18 HG1# 1.80 3.19 # Same atoms-diff bounds ( 156)
166-> ILE A 11 HA - ILE A 11 HG2# 1.80 4.13 # NoRestrctn I [2.63 3.78] -- intra
169-> ILE A 11 HA - ILE A 11 HD1# 1.80 6.00 # NoRestrctn I [2.11 5.99] -- intra
178-> ILE A 56 HA - ILE A 56 HD1# 1.80 6.00 # NoRestrctn I [2.11 5.99] -- intra
211-> LEU A 14 HN - LYS A 15 HA 1.80 5.96 # NoRestrctn S [2.00 3.99] -- sequential
246-> ASN A 49 HN - ALA A 50 HA 1.80 5.70 # NoRestrctn S [2.00 3.99] -- sequential
281-> VAL A 52 HN - LEU A 53 HA 1.80 6.00 # NoRestrctn S [2.00 3.99] -- sequential
317-> ALA A 50 HN - VAL A 51 HA 1.80 6.00 # NoRestrctn S [2.00 3.99] -- sequential
336-> ASN A 21 HN - GLN A 22 HA 1.80 4.48 # NoRestrctn S [2.00 3.99] -- sequential
352-> TYR A 63 HN - ASN A 64 HA 1.80 6.00 # NoRestrctn S [2.00 3.99] -- sequential
353-> GLU A 61 HN - ALA A 62 HA 1.80 6.00 # NoRestrctn S [2.00 3.99] -- sequential
436-> MET A 20 HN - ASN A 21 HA 1.80 6.00 # NoRestrctn S [2.00 3.99] -- sequential
442-> SER A 23 HN - GLU A 24 HA 1.80 5.60 # NoRestrctn S [2.00 3.99] -- sequential
461-> GLY A 69 HN - GLU A 70 HA 1.80 5.04 # NoRestrctn S [2.00 3.99] -- sequential
713-> GLN A 43 HA - GLN A 43 HB# 1.80 2.44 # FixedDistn I [0.00 0.00] -- intra
730-> LEU A 46 HB# - LEU A 46 HD# 1.80 2.44 # TooRestrct I [2.57 2.89] -- intra
748-> LYS A 48 HG# - LYS A 48 HE# 1.80 2.38 # TooRestrct I [2.52 3.73] -- intra
812-> LYS A 54 HD# - LYS A 54 HE# 1.80 2.05 # TooRestrct I [2.27 2.51] -- intra
857-> ARG A 68 HN - GLY A 69 HA# 1.80 4.33 # NoRestrctn S [2.00 3.55] -- sequential
871-> GLU A 70 HA - GLU A 70 HB# 1.80 2.60 # FixedDistn I [0.00 0.00] -- intra
====== TOTAL ======: 28
table of distance constraints violations
Residual Violations greater than 0.10
7-> VAL A 38 HA - LEU A 41 HG [ 1.80 4.15] 0.00 0.08 0.02 0.00 0.00 0.00 0.00 0.38 0.00 0.00 0.15 0.00 0.03 0.00 0.08 0.00 0.02 0.13 0.00 0.00 - 8 [ 0.02 .. 0.38]
13-> PRO A 8 HA - ILE A 11 HG2* [ 1.80 4.48] 0.00 0.00 0.00 0.00 0.00 0.00 0.00 0.00 0.00 0.00 0.00 0.00 0.00 0.00 0.12 0.00 0.03 0.11 0.02 0.00 - 4 [ 0.02 .. 0.12]
17-> VAL A 37 HA - LEU A 41 HG [ 1.80 6.00] 0.00 0.00 0.00 0.00 0.00 0.00 0.00 0.00 0.00 0.00 0.03 0.00 0.00 0.42 0.00 0.00 0.19 0.00 0.31 0.00 - 4 [ 0.03 .. 0.42]
22-> LEU A 46 HN - THR A 47 HA [ 1.80 5.53] 0.00 0.00 0.00 0.00 0.00 0.00 0.00 0.00 0.00 0.00 0.00 0.00 0.00 0.00 0.00 0.00 0.12 0.00 0.00 0.00 - 1 [ 0.12 .. 0.12]
24-> ILE A 27 HA - ILE A 27 HD1* [ 1.80 3.32] 0.35 0.36 0.40 0.00 0.00 0.00 0.37 0.40 0.21 0.36 0.25 0.38 0.00 0.00 0.00 0.34 0.15 0.31 0.37 0.00 - 13 [ 0.15 .. 0.40]
32-> ILE A 18 HD1* - LYS A 26 HA [ 1.80 4.87] 0.18 0.07 0.23 0.00 0.15 0.19 0.09 0.10 0.16 0.16 0.18 0.18 0.17 0.09 0.10 0.10 0.18 0.35 0.21 0.10 - 19 [ 0.07 .. 0.35]
40-> LYS A 67 HA - LYS A 67 HG3 [ 1.80 3.44] 0.19 0.28 0.12 0.14 0.03 0.22 0.25 0.10 0.21 0.13 0.23 0.28 0.16 0.23 0.22 0.25 0.01 0.04 0.22 0.12 - 20 [ 0.01 .. 0.28]
43-> ILE A 18 HB - GLU A 19 HA [ 1.80 4.76] 0.00 0.00 0.00 0.05 0.00 0.00 0.00 0.00 0.00 0.00 0.00 0.00 0.00 0.00 0.11 0.00 0.00 0.00 0.00 0.00 - 2 [ 0.05 .. 0.11]
44-> LEU A 53 HA - ILE A 56 HD1* [ 1.80 4.44] 0.00 0.00 0.00 0.00 0.00 0.00 0.00 0.33 0.00 0.25 0.15 0.00 0.00 0.02 0.00 0.00 0.41 0.00 0.00 0.00 - 5 [ 0.02 .. 0.41]
51-> LEU A 46 HA - VAL A 52 HN [ 1.80 5.69] 0.00 0.00 0.00 0.00 0.00 0.10 0.00 0.03 0.00 0.00 0.00 0.08 0.05 0.00 0.03 0.30 0.43 0.00 0.00 0.00 - 7 [ 0.03 .. 0.43]
55-> LEU A 73 HA - LEU A 73 HG [ 1.80 3.51] 0.00 0.00 0.00 0.13 0.00 0.00 0.00 0.00 0.00 0.00 0.00 0.00 0.00 0.00 0.00 0.07 0.00 0.00 0.07 0.00 - 3 [ 0.07 .. 0.13]
57-> ASP A 25 HB2 - MET A 30 HN [ 1.80 5.83] 0.13 0.24 0.22 0.30 0.25 0.22 0.13 0.09 0.20 0.21 0.17 0.21 0.33 0.01 0.00 0.00 0.28 0.05 0.16 0.02 - 18 [ 0.01 .. 0.33]
60-> LYS A 54 HD2 - LYS A 54 HE* [ 1.80 2.36] 0.02 0.04 0.00 0.00 0.06 0.00 0.00 0.00 0.04 0.11 0.00 0.00 0.00 0.00 0.06 0.01 0.00 0.03 0.04 0.00 - 9 [ 0.01 .. 0.11]
65-> LEU A 53 HD1* - ILE A 56 HB [ 1.80 5.17] 0.00 0.00 0.00 0.24 0.00 0.00 0.00 0.00 0.00 0.00 0.00 0.00 0.00 0.00 0.00 0.00 0.00 0.00 0.09 0.03 - 3 [ 0.03 .. 0.24]
66-> VAL A 7 HG2* - GLU A 61 HG2 [ 1.80 6.00] 0.00 0.00 0.00 0.00 0.00 0.00 0.00 0.00 0.00 0.00 0.00 0.01 0.00 0.00 0.12 0.00 0.00 0.00 0.00 0.00 - 2 [ 0.01 .. 0.12]
67-> HIS A 17 HN - GLU A 24 HG2 [ 1.80 6.00] 0.00 0.00 0.00 0.27 0.00 0.00 0.00 0.06 0.00 0.00 0.07 0.00 0.00 0.00 0.13 0.00 0.00 0.00 0.04 0.01 - 6 [ 0.01 .. 0.27]
69-> GLN A 16 HG3 - ASN A 21 HN [ 1.80 5.17] 0.04 0.00 0.04 0.12 0.13 0.03 0.06 0.00 0.12 0.10 0.14 0.07 0.08 0.00 0.14 0.00 0.04 0.21 0.04 0.06 - 16 [ 0.03 .. 0.21]
81-> LYS A 67 HN - ARG A 68 HB3 [ 1.80 6.00] 0.06 0.14 0.00 0.00 0.00 0.00 0.00 0.00 0.07 0.00 0.00 0.00 0.09 0.09 0.00 0.00 0.00 0.00 0.00 0.02 - 6 [ 0.02 .. 0.14]
82-> GLN A 22 HB2 - SER A 23 HN [ 1.80 4.16] 0.37 0.00 0.00 0.00 0.00 0.00 0.00 0.00 0.40 0.00 0.00 0.00 0.42 0.00 0.00 0.00 0.00 0.00 0.00 0.00 - 3 [ 0.37 .. 0.42]
83-> LEU A 46 HG - LYS A 48 HN [ 1.80 6.00] 0.00 0.07 0.11 0.10 0.00 0.11 0.09 0.00 0.00 0.00 0.00 0.00 0.00 0.10 0.22 0.00 0.06 0.00 0.00 0.02 - 10 [ 0.00 .. 0.22]
85-> LEU A 73 HG - GLU A 74 HN [ 1.80 4.46] 0.00 0.00 0.00 0.29 0.00 0.00 0.25 0.02 0.00 0.00 0.00 0.36 0.00 0.06 0.21 0.00 0.00 0.45 0.00 0.00 - 7 [ 0.02 .. 0.45]
86-> LEU A 73 HN - LEU A 73 HG [ 1.80 4.49] 0.00 0.00 0.14 0.00 0.00 0.07 0.00 0.11 0.00 0.00 0.14 0.00 0.00 0.00 0.08 0.00 0.00 0.00 0.00 0.00 - 7 [ 0.00 .. 0.14]
87-> LEU A 14 HD1* - ALA A 62 HN [ 1.80 6.00] 0.00 0.00 0.00 0.00 0.00 0.00 0.00 0.00 0.00 0.00 0.00 0.00 0.00 0.00 0.00 0.00 0.00 0.00 0.25 0.00 - 1 [ 0.25 .. 0.25]
89-> MET A 30 HN - VAL A 66 HG2* [ 1.80 3.96] 0.00 0.00 0.00 0.14 0.00 0.00 0.00 0.00 0.00 0.00 0.00 0.00 0.04 0.49 0.00 0.00 0.00 0.15 0.00 0.00 - 5 [ 0.00 .. 0.49]
97-> LEU A 53 HD2* - ILE A 56 HB [ 1.80 5.17] 0.00 0.00 0.00 0.00 0.00 0.00 0.00 0.00 0.00 0.28 0.28 0.00 0.04 0.00 0.04 0.00 0.32 0.00 0.00 0.00 - 5 [ 0.04 .. 0.32]
101-> LEU A 4 HD2* - TYR A 9 HE* [ 1.80 4.75] 0.00 0.00 0.00 0.00 0.00 0.00 0.00 0.00 0.00 0.00 0.00 0.00 0.18 0.00 0.00 0.00 0.00 0.00 0.00 0.00 - 1 [ 0.18 .. 0.18]
103-> LEU A 4 HN - LEU A 4 HD2* [ 1.80 6.00] 0.00 0.00 0.00 0.00 0.00 0.00 0.00 0.00 0.00 0.00 0.00 0.12 0.00 0.00 0.00 0.00 0.00 0.00 0.00 0.00 - 1 [ 0.12 .. 0.12]
109-> VAL A 7 HN - VAL A 7 HG2* [ 1.80 4.85] 0.02 0.02 0.04 0.04 0.00 0.00 0.00 0.00 0.02 0.00 0.04 0.01 0.05 0.00 0.01 0.06 0.01 0.13 0.00 0.00 - 12 [ 0.01 .. 0.13]
110-> LEU A 46 HN - THR A 47 HG2* [ 1.80 6.00] 0.00 0.00 0.11 0.00 0.00 0.00 0.00 0.00 0.00 0.00 0.00 0.00 0.00 0.00 0.00 0.00 0.00 0.00 0.25 0.00 - 2 [ 0.11 .. 0.25]
111-> THR A 47 HG2* - VAL A 52 HN [ 1.80 4.99] 0.00 0.00 0.13 0.00 0.00 0.00 0.00 0.00 0.00 0.00 0.34 0.00 0.49 0.00 0.00 0.38 0.00 0.00 0.17 0.00 - 5 [ 0.13 .. 0.49]
123-> THR A 47 HN - VAL A 51 HG2* [ 1.80 4.92] 0.10 0.12 0.25 0.10 0.03 0.07 0.18 0.00 0.00 0.00 0.16 0.00 0.11 0.13 0.02 0.00 0.00 0.00 0.17 0.12 - 13 [ 0.02 .. 0.25]
134-> VAL A 37 HN - LEU A 59 HD1* [ 1.80 5.39] 0.00 0.12 0.00 0.00 0.11 0.00 0.00 0.00 0.09 0.00 0.03 0.00 0.07 0.21 0.00 0.06 0.00 0.02 0.03 0.00 - 9 [ 0.02 .. 0.21]
153-> ILE A 27 HG2* - ALA A 29 HB* [ 1.80 5.40] 0.00 0.00 0.00 0.12 0.00 0.00 0.00 0.00 0.00 0.00 0.00 0.00 0.00 0.00 0.00 0.00 0.00 0.00 0.00 0.00 - 1 [ 0.12 .. 0.12]
155-> LYS A 26 HN - ILE A 27 HG2* [ 1.80 6.00] 0.18 0.00 0.00 0.00 0.00 0.00 0.00 0.00 0.00 0.00 0.26 0.00 0.00 0.00 0.12 0.22 0.00 0.00 0.00 0.00 - 5 [ 0.00 .. 0.26]
157-> LYS A 15 HN - ILE A 18 HG2* [ 1.80 5.06] 0.13 0.13 0.18 0.00 0.17 0.18 0.00 0.10 0.08 0.19 0.15 0.21 0.11 0.11 0.00 0.07 0.12 0.10 0.00 0.12 - 16 [ 0.07 .. 0.21]
167-> ILE A 11 HG2* - ALA A 62 HB* [ 1.80 3.41] 0.00 0.00 0.00 0.00 0.35 0.30 0.22 0.01 0.00 0.00 0.06 0.00 0.31 0.00 0.00 0.00 0.01 0.08 0.00 0.13 - 9 [ 0.01 .. 0.35]
174-> ILE A 18 HD1* - ALA A 29 HB* [ 1.80 4.55] 0.00 0.00 0.00 0.05 0.00 0.03 0.02 0.00 0.02 0.00 0.08 0.00 0.00 0.00 0.13 0.07 0.00 0.00 0.00 0.00 - 7 [ 0.02 .. 0.13]
182-> ILE A 27 HD1* - SER A 71 HN [ 1.80 4.98] 0.00 0.00 0.00 0.00 0.00 0.00 0.00 0.17 0.00 0.00 0.00 0.00 0.00 0.00 0.00 0.05 0.00 0.00 0.00 0.00 - 2 [ 0.05 .. 0.17]
184-> ILE A 27 HD1* - TYR A 63 HE* [ 1.80 6.00] 0.00 0.00 0.00 0.00 0.00 0.00 0.00 0.30 0.00 0.00 0.07 0.00 0.00 0.00 0.00 0.08 0.00 0.00 0.00 0.00 - 3 [ 0.07 .. 0.30]
186-> ASN A 21 HA - GLU A 24 HB2 [ 1.80 6.00] 0.00 0.00 0.00 0.00 0.00 0.00 0.00 0.00 0.00 0.00 0.00 0.00 0.00 0.00 0.31 0.00 0.00 0.00 0.00 0.00 - 1 [ 0.31 .. 0.31]
189-> LEU A 14 HD2* - ALA A 62 HB* [ 1.80 3.91] 0.12 0.14 0.11 0.00 0.09 0.00 0.00 0.03 0.11 0.00 0.07 0.00 0.06 0.00 0.16 0.00 0.06 0.16 0.00 0.05 - 12 [ 0.03 .. 0.16]
192-> THR A 47 HG2* - VAL A 51 HG1* [ 1.80 2.79] 0.00 0.00 0.12 0.00 0.00 0.00 0.00 0.00 0.00 0.00 0.00 0.00 0.07 0.00 0.00 0.00 0.00 0.00 0.32 0.00 - 3 [ 0.07 .. 0.32]
196-> GLU A 19 HN - MET A 20 HE* [ 1.80 5.85] 0.00 0.00 0.00 0.00 0.00 0.13 0.19 0.00 0.00 0.19 0.00 0.00 0.00 0.23 0.00 0.00 0.00 0.00 0.00 0.00 - 4 [ 0.13 .. 0.23]
197-> HIS A 17 HN - MET A 20 HE* [ 1.80 6.00] 0.11 0.14 0.07 0.08 0.20 0.23 0.18 0.00 0.00 0.24 0.00 0.08 0.00 0.00 0.00 0.11 0.10 0.20 0.02 0.09 - 14 [ 0.02 .. 0.24]
202-> THR A 47 HN - LYS A 48 HN [ 1.80 5.37] 0.00 0.00 0.00 0.00 0.00 0.00 0.00 0.00 0.00 0.00 0.00 0.00 0.00 0.00 0.00 0.00 0.13 0.00 0.00 0.00 - 1 [ 0.13 .. 0.13]
203-> THR A 47 HA - LYS A 48 HN [ 1.80 3.28] 0.20 0.20 0.23 0.22 0.20 0.18 0.20 0.17 0.20 0.18 0.24 0.25 0.25 0.22 0.21 0.28 0.18 0.22 0.19 0.19 - 20 [ 0.17 .. 0.28]
210-> GLN A 45 HN - LEU A 46 HG [ 1.80 6.00] 0.00 0.00 0.00 0.00 0.00 0.00 0.00 0.00 0.00 0.00 0.20 0.09 0.00 0.00 0.00 0.11 0.00 0.00 0.22 0.00 - 4 [ 0.09 .. 0.22]
215-> LEU A 14 HN - LEU A 14 HD2* [ 1.80 3.96] 0.00 0.00 0.00 0.14 0.00 0.00 0.00 0.00 0.00 0.12 0.00 0.00 0.00 0.00 0.00 0.00 0.00 0.00 0.00 0.00 - 2 [ 0.12 .. 0.14]
227-> ILE A 56 HD1* - LEU A 59 HN [ 1.80 6.00] 0.39 0.00 0.40 0.34 0.00 0.30 0.00 0.00 0.46 0.00 0.00 0.41 0.40 0.00 0.26 0.38 0.00 0.00 0.35 0.00 - 10 [ 0.26 .. 0.46]
229-> SER A 71 HA - LYS A 72 HN [ 1.80 3.07] 0.00 0.00 0.12 0.00 0.00 0.00 0.00 0.00 0.45 0.00 0.00 0.00 0.00 0.00 0.00 0.00 0.00 0.08 0.00 0.00 - 3 [ 0.08 .. 0.45]
230-> LYS A 72 HN - GLU A 74 HG3 [ 1.80 6.00] 0.17 0.02 0.00 0.00 0.00 0.00 0.00 0.01 0.04 0.00 0.00 0.00 0.00 0.00 0.00 0.00 0.00 0.42 0.19 0.00 - 6 [ 0.01 .. 0.42]
231-> LYS A 72 HN - GLU A 74 HG2 [ 1.80 6.00] 0.00 0.00 0.14 0.00 0.00 0.00 0.00 0.18 0.00 0.00 0.00 0.00 0.07 0.03 0.09 0.00 0.00 0.00 0.00 0.19 - 6 [ 0.03 .. 0.19]
238-> ASP A 25 HN - HIS A 28 HN [ 1.80 6.00] 0.00 0.00 0.00 0.00 0.00 0.00 0.00 0.00 0.00 0.00 0.00 0.00 0.00 0.06 0.06 0.13 0.00 0.04 0.00 0.00 - 4 [ 0.04 .. 0.13]
261-> VAL A 37 HN - LEU A 59 HD2* [ 1.80 5.39] 0.00 0.00 0.00 0.00 0.00 0.00 0.00 0.00 0.00 0.00 0.00 0.16 0.00 0.00 0.00 0.00 0.00 0.00 0.00 0.00 - 2 [ 0.00 .. 0.16]
262-> TYR A 33 HN - VAL A 37 HN [ 1.80 6.00] 0.15 0.14 0.18 0.01 0.00 0.00 0.07 0.13 0.11 0.06 0.00 0.15 0.04 0.26 0.15 0.06 0.11 0.15 0.18 0.03 - 17 [ 0.01 .. 0.26]
265-> SER A 23 HN - GLU A 24 HN [ 1.80 3.39] 0.00 0.00 0.00 0.12 0.00 0.00 0.00 0.00 0.00 0.00 0.00 0.00 0.00 0.00 0.00 0.00 0.00 0.00 0.00 0.00 - 1 [ 0.12 .. 0.12]
268-> LEU A 73 HA - GLU A 74 HN [ 1.80 3.22] 0.00 0.32 0.00 0.34 0.00 0.32 0.00 0.00 0.00 0.00 0.00 0.00 0.33 0.00 0.13 0.09 0.05 0.00 0.00 0.00 - 7 [ 0.05 .. 0.34]
278-> VAL A 66 HN - LYS A 67 HG3 [ 1.80 4.66] 0.02 0.45 0.20 0.00 0.00 0.22 0.32 0.00 0.02 0.08 0.33 0.37 0.05 0.20 0.49 0.37 0.00 0.16 0.13 0.00 - 15 [ 0.02 .. 0.49]
285-> ILE A 18 HN - ASP A 25 HN [ 1.80 6.00] 0.12 0.00 0.08 0.37 0.01 0.03 0.00 0.08 0.00 0.00 0.00 0.00 0.04 0.00 0.00 0.06 0.00 0.10 0.03 0.06 - 12 [ 0.00 .. 0.37]
292-> TYR A 33 HE* - VAL A 38 HN [ 1.80 6.00] 0.19 0.27 0.00 0.15 0.17 0.19 0.16 0.14 0.24 0.18 0.00 0.27 0.13 0.02 0.14 0.21 0.02 0.00 0.22 0.20 - 18 [ 0.00 .. 0.27]
293-> TYR A 33 HD* - VAL A 38 HN [ 1.80 6.00] 0.17 0.05 0.00 0.00 0.00 0.05 0.00 0.10 0.00 0.00 0.09 0.07 0.04 0.00 0.00 0.15 0.20 0.00 0.12 0.11 - 11 [ 0.04 .. 0.20]
299-> ARG A 68 HN - GLU A 70 HN [ 1.80 4.88] 0.00 0.00 0.00 0.00 0.00 0.00 0.00 0.00 0.00 0.00 0.00 0.00 0.00 0.00 0.11 0.00 0.00 0.00 0.00 0.00 - 1 [ 0.11 .. 0.11]
302-> VAL A 38 HN - THR A 40 HB [ 1.80 6.00] 0.00 0.00 0.00 0.00 0.00 0.00 0.00 0.00 0.00 0.00 0.00 0.00 0.00 0.00 0.00 0.00 0.00 0.00 0.12 0.00 - 1 [ 0.12 .. 0.12]
316-> ILE A 56 HD1* - GLN A 57 HN [ 1.80 4.52] 0.00 0.04 0.00 0.00 0.01 0.00 0.00 0.29 0.00 0.27 0.04 0.00 0.00 0.12 0.00 0.00 0.14 0.00 0.00 0.00 - 7 [ 0.01 .. 0.29]
319-> LEU A 46 HN - THR A 47 HN [ 1.80 3.23] 0.07 0.00 0.00 0.00 0.03 0.00 0.00 0.00 0.00 0.00 0.00 0.06 0.04 0.00 0.00 0.00 0.32 0.08 0.00 0.00 - 6 [ 0.03 .. 0.32]
323-> THR A 47 HN - VAL A 51 HG1* [ 1.80 4.92] 0.00 0.00 0.00 0.00 0.00 0.00 0.00 0.00 0.00 0.00 0.00 0.34 0.00 0.00 0.00 0.16 0.00 0.00 0.00 0.00 - 2 [ 0.16 .. 0.34]
327-> THR A 47 HG2* - ALA A 50 HN [ 1.80 6.00] 0.00 0.00 0.29 0.00 0.00 0.00 0.00 0.00 0.00 0.00 0.30 0.00 0.07 0.00 0.00 0.00 0.00 0.00 0.21 0.00 - 4 [ 0.07 .. 0.30]
328-> GLU A 74 HA - HIS A 75 HN [ 1.80 3.37] 0.00 0.00 0.00 0.00 0.14 0.00 0.13 0.19 0.00 0.00 0.00 0.10 0.17 0.00 0.00 0.00 0.00 0.15 0.00 0.06 - 7 [ 0.06 .. 0.19]
331-> VAL A 42 HN - ASP A 44 HN [ 1.80 5.16] 0.00 0.00 0.00 0.00 0.00 0.00 0.00 0.00 0.19 0.00 0.00 0.00 0.00 0.00 0.00 0.00 0.00 0.00 0.00 0.00 - 1 [ 0.19 .. 0.19]
332-> LEU A 41 HA - ASP A 44 HN [ 1.80 5.42] 0.00 0.00 0.00 0.00 0.00 0.00 0.00 0.00 0.13 0.00 0.00 0.00 0.00 0.00 0.00 0.00 0.00 0.00 0.00 0.00 - 1 [ 0.13 .. 0.13]
333-> VAL A 42 HB - ASP A 44 HN [ 1.80 5.64] 0.00 0.02 0.00 0.11 0.00 0.00 0.00 0.00 0.16 0.02 0.18 0.00 0.01 0.00 0.01 0.00 0.00 0.19 0.00 0.00 - 8 [ 0.01 .. 0.19]
334-> ASP A 25 HB2 - HIS A 28 HN [ 1.80 3.71] 0.09 0.04 0.06 0.00 0.00 0.07 0.09 0.09 0.00 0.01 0.02 0.06 0.00 0.10 0.17 0.13 0.05 0.16 0.03 0.00 - 16 [ 0.00 .. 0.17]
336-> ASN A 21 HN - GLN A 22 HA [ 1.80 4.48] 0.00 0.32 0.00 0.00 0.00 0.00 0.00 0.26 0.00 0.00 0.00 0.00 0.00 0.33 0.00 0.35 0.00 0.00 0.00 0.00 - 4 [ 0.26 .. 0.35]
340-> VAL A 52 HN - LYS A 54 HN [ 1.80 4.42] 0.00 0.00 0.01 0.00 0.00 0.02 0.00 0.00 0.11 0.00 0.02 0.00 0.00 0.00 0.00 0.00 0.06 0.00 0.00 0.00 - 6 [ 0.00 .. 0.11]
342-> VAL A 51 HA - LYS A 54 HN [ 1.80 4.33] 0.00 0.00 0.00 0.00 0.00 0.00 0.00 0.00 0.00 0.00 0.02 0.00 0.00 0.02 0.00 0.00 0.00 0.00 0.00 0.11 - 3 [ 0.02 .. 0.11]
354-> GLN A 22 HN - GLU A 24 HN [ 1.80 4.71] 0.00 0.07 0.08 0.00 0.07 0.03 0.06 0.12 0.00 0.01 0.00 0.03 0.00 0.03 0.04 0.12 0.00 0.00 0.06 0.13 - 13 [ 0.01 .. 0.13]
356-> GLN A 16 HA - GLN A 22 HN [ 1.80 6.00] 0.14 0.19 0.25 0.20 0.21 0.22 0.21 0.27 0.05 0.26 0.09 0.17 0.09 0.28 0.05 0.25 0.16 0.11 0.20 0.25 - 20 [ 0.05 .. 0.28]
359-> ILE A 11 HD1* - GLU A 61 HN [ 1.80 5.89] 0.00 0.00 0.00 0.00 0.00 0.17 0.08 0.00 0.00 0.00 0.00 0.03 0.22 0.08 0.00 0.00 0.00 0.00 0.00 0.09 - 6 [ 0.03 .. 0.22]
361-> VAL A 7 HA - TYR A 9 HN [ 1.80 3.73] 0.00 0.07 0.08 0.03 0.00 0.00 0.05 0.02 0.11 0.02 0.06 0.00 0.05 0.14 0.11 0.00 0.09 0.02 0.00 0.00 - 14 [ 0.00 .. 0.14]
362-> VAL A 7 HB - TYR A 9 HN [ 1.80 4.79] 0.08 0.13 0.13 0.09 0.06 0.03 0.23 0.10 0.12 0.01 0.08 0.06 0.00 0.20 0.10 0.03 0.13 0.00 0.11 0.11 - 18 [ 0.01 .. 0.23]
363-> MET A 30 HN - TYR A 33 HD* [ 1.80 6.00] 0.33 0.35 0.37 0.23 0.43 0.37 0.36 0.27 0.35 0.30 0.00 0.38 0.21 0.20 0.40 0.33 0.00 0.23 0.36 0.21 - 18 [ 0.20 .. 0.43]
375-> MET A 30 HN - VAL A 66 HB [ 1.80 4.48] 0.28 0.28 0.18 0.25 0.28 0.40 0.22 0.28 0.27 0.13 0.21 0.19 0.32 0.45 0.28 0.36 0.36 0.26 0.23 0.15 - 20 [ 0.13 .. 0.45]
378-> ILE A 56 HD1* - HIS A 58 HN [ 1.80 6.00] 0.26 0.00 0.25 0.26 0.00 0.26 0.00 0.00 0.19 0.00 0.00 0.28 0.22 0.00 0.25 0.29 0.00 0.00 0.23 0.00 - 10 [ 0.19 .. 0.29]
379-> PRO A 8 HA - GLU A 12 HN [ 1.80 4.13] 0.18 0.19 0.02 0.00 0.00 0.00 0.00 0.11 0.17 0.17 0.03 0.00 0.00 0.00 0.02 0.00 0.16 0.03 0.04 0.03 - 12 [ 0.02 .. 0.19]
383-> GLN A 43 HN - ASP A 44 HN [ 1.80 3.37] 0.00 0.13 0.00 0.02 0.00 0.28 0.06 0.00 0.34 0.00 0.00 0.00 0.00 0.00 0.16 0.00 0.00 0.06 0.00 0.20 - 8 [ 0.02 .. 0.34]
384-> LEU A 41 HA - GLN A 43 HN [ 1.80 4.56] 0.00 0.00 0.03 0.00 0.08 0.00 0.00 0.00 0.00 0.00 0.00 0.00 0.00 0.12 0.00 0.00 0.00 0.00 0.00 0.00 - 4 [ 0.00 .. 0.12]
401-> ASN A 64 HN - ARG A 68 HN [ 1.80 6.00] 0.00 0.00 0.00 0.00 0.05 0.07 0.17 0.00 0.00 0.00 0.09 0.16 0.01 0.00 0.04 0.10 0.03 0.00 0.00 0.09 - 10 [ 0.01 .. 0.17]
403-> ILE A 11 HD1* - ASN A 64 HN [ 1.80 6.00] 0.05 0.02 0.00 0.07 0.01 0.02 0.13 0.00 0.05 0.07 0.04 0.22 0.00 0.00 0.00 0.07 0.00 0.00 0.00 0.17 - 12 [ 0.01 .. 0.22]
407-> ASN A 13 HN - TYR A 33 HE* [ 1.80 5.63] 0.09 0.22 0.05 0.24 0.03 0.08 0.08 0.05 0.10 0.00 0.24 0.04 0.00 0.24 0.14 0.02 0.19 0.14 0.00 0.27 - 18 [ 0.00 .. 0.27]
413-> LYS A 15 HN - ILE A 18 HD1* [ 1.80 4.40] 0.00 0.05 0.05 0.16 0.00 0.06 0.16 0.07 0.03 0.08 0.00 0.04 0.02 0.16 0.07 0.19 0.00 0.09 0.23 0.02 - 16 [ 0.02 .. 0.23]
422-> MET A 30 HN - SER A 32 HN [ 1.80 4.48] 0.00 0.05 0.00 0.00 0.00 0.06 0.00 0.00 0.00 0.00 0.00 0.00 0.00 0.00 0.00 0.00 0.03 0.00 0.00 0.11 - 4 [ 0.03 .. 0.11]
424-> ILE A 27 HD1* - ARG A 68 HN [ 1.80 6.00] 0.13 0.00 0.00 0.00 0.00 0.00 0.25 0.00 0.09 0.00 0.11 0.00 0.00 0.00 0.00 0.07 0.04 0.07 0.04 0.00 - 8 [ 0.04 .. 0.25]
428-> ASN A 64 HA - ARG A 68 HN [ 1.80 4.04] 0.00 0.00 0.00 0.00 0.00 0.02 0.07 0.00 0.00 0.00 0.03 0.10 0.00 0.00 0.07 0.04 0.00 0.00 0.00 0.04 - 7 [ 0.02 .. 0.10]
432-> ARG A 68 HN - GLU A 70 HG3 [ 1.80 6.00] 0.00 0.00 0.00 0.26 0.24 0.00 0.00 0.08 0.46 0.26 0.00 0.00 0.46 0.22 0.00 0.00 0.37 0.09 0.18 0.12 - 11 [ 0.08 .. 0.46]
440-> GLN A 22 HB3 - SER A 23 HN [ 1.80 4.16] 0.03 0.00 0.00 0.00 0.00 0.00 0.00 0.00 0.11 0.00 0.00 0.00 0.12 0.00 0.00 0.00 0.00 0.00 0.00 0.00 - 3 [ 0.03 .. 0.12]
441-> SER A 23 HN - GLU A 24 HB2 [ 1.80 4.22] 0.32 0.00 0.00 0.00 0.00 0.00 0.00 0.00 0.38 0.00 0.00 0.00 0.33 0.00 0.00 0.00 0.00 0.00 0.00 0.00 - 3 [ 0.32 .. 0.38]
444-> LEU A 4 HN - SER A 6 HN [ 1.80 6.00] 0.00 0.00 0.00 0.00 0.00 0.00 0.00 0.00 0.00 0.00 0.00 0.00 0.24 0.00 0.00 0.00 0.00 0.00 0.00 0.00 - 1 [ 0.24 .. 0.24]
452-> SER A 39 HN - VAL A 42 HB [ 1.80 5.60] 0.00 0.00 0.00 0.00 0.00 0.00 0.00 0.17 0.01 0.08 0.00 0.00 0.00 0.00 0.00 0.00 0.00 0.00 0.00 0.00 - 3 [ 0.01 .. 0.17]
456-> ARG A 68 HB3 - GLY A 69 HN [ 1.80 3.52] 0.24 0.49 0.00 0.00 0.00 0.00 0.00 0.00 0.18 0.00 0.00 0.00 0.08 0.10 0.00 0.10 0.00 0.00 0.00 0.12 - 7 [ 0.08 .. 0.49]
457-> LYS A 67 HG3 - GLY A 69 HN [ 1.80 6.00] 0.08 0.29 0.00 0.04 0.00 0.21 0.25 0.00 0.16 0.13 0.17 0.24 0.29 0.29 0.21 0.22 0.00 0.00 0.05 0.13 - 15 [ 0.04 .. 0.29]
458-> ILE A 18 HD1* - GLY A 69 HN [ 1.80 6.00] 0.00 0.00 0.00 0.00 0.00 0.00 0.00 0.14 0.00 0.00 0.00 0.00 0.00 0.00 0.00 0.00 0.00 0.00 0.00 0.00 - 1 [ 0.14 .. 0.14]
459-> ILE A 27 HD1* - GLY A 69 HN [ 1.80 6.00] 0.14 0.19 0.14 0.00 0.00 0.00 0.24 0.00 0.31 0.25 0.20 0.07 0.00 0.00 0.00 0.00 0.27 0.26 0.17 0.00 - 11 [ 0.07 .. 0.31]
465-> PHE A 10 HZ - VAL A 37 HG2* [ 1.80 4.31] 0.00 0.00 0.12 0.00 0.00 0.00 0.00 0.00 0.00 0.00 0.00 0.00 0.00 0.00 0.00 0.00 0.00 0.34 0.00 0.00 - 2 [ 0.12 .. 0.34]
467-> TYR A 34 HD* - TYR A 63 HA [ 1.80 4.43] 0.29 0.34 0.12 0.08 0.13 0.06 0.11 0.05 0.05 0.13 0.31 0.13 0.05 0.16 0.27 0.12 0.22 0.11 0.23 0.00 - 19 [ 0.05 .. 0.34]
473-> LEU A 4 HN - TYR A 9 HD* [ 1.80 5.00] 0.00 0.00 0.02 0.00 0.00 0.01 0.03 0.00 0.00 0.00 0.00 0.26 0.12 0.00 0.00 0.00 0.01 0.00 0.03 0.00 - 7 [ 0.01 .. 0.26]
474-> TYR A 9 HD* - PHE A 10 HZ [ 1.80 6.00] 0.00 0.00 0.00 0.27 0.00 0.00 0.00 0.00 0.00 0.00 0.00 0.03 0.00 0.10 0.00 0.00 0.00 0.00 0.09 0.00 - 4 [ 0.03 .. 0.27]
481-> PHE A 10 HD* - VAL A 37 HB [ 1.80 6.00] 0.00 0.00 0.00 0.01 0.00 0.00 0.00 0.00 0.00 0.00 0.14 0.00 0.00 0.00 0.00 0.00 0.00 0.00 0.00 0.00 - 2 [ 0.01 .. 0.14]
483-> LEU A 4 HN - TYR A 9 HE* [ 1.80 5.26] 0.05 0.00 0.27 0.00 0.00 0.18 0.03 0.05 0.00 0.07 0.00 0.47 0.01 0.04 0.00 0.13 0.08 0.00 0.03 0.00 - 12 [ 0.01 .. 0.47]
484-> TYR A 9 HE* - PHE A 10 HE* [ 1.80 4.70] 0.01 0.00 0.00 0.00 0.00 0.00 0.01 0.07 0.00 0.00 0.11 0.00 0.00 0.00 0.06 0.00 0.03 0.00 0.00 0.00 - 6 [ 0.01 .. 0.11]
485-> LEU A 4 HA - TYR A 9 HE* [ 1.80 4.82] 0.00 0.00 0.00 0.00 0.00 0.00 0.00 0.00 0.00 0.00 0.00 0.15 0.00 0.00 0.00 0.00 0.00 0.00 0.00 0.00 - 1 [ 0.15 .. 0.15]
487-> LEU A 14 HG - TYR A 33 HE* [ 1.80 4.77] 0.00 0.02 0.00 0.00 0.00 0.00 0.00 0.00 0.00 0.00 0.00 0.00 0.00 0.00 0.15 0.00 0.00 0.00 0.00 0.06 - 3 [ 0.02 .. 0.15]
496-> GLU A 3 HA - LEU A 4 HB* [ 1.80 4.47] 0.15 0.00 0.10 0.13 0.00 0.12 0.05 0.00 0.08 0.00 0.12 0.30 0.21 0.09 0.06 0.00 0.18 0.00 0.00 0.00 - 12 [ 0.05 .. 0.30]
498-> GLU A 3 HG* - LEU A 4 HD* [ 1.80 5.79] 0.00 0.00 0.00 0.00 0.00 0.00 0.00 0.00 0.00 0.25 0.00 0.00 0.00 0.00 0.00 0.21 0.00 0.00 0.00 0.46 - 3 [ 0.21 .. 0.46]
500-> LEU A 4 HN - LEU A 4 HD* [ 1.80 4.50] 0.00 0.00 0.00 0.00 0.00 0.00 0.00 0.00 0.00 0.00 0.00 0.13 0.00 0.00 0.00 0.00 0.00 0.00 0.00 0.00 - 1 [ 0.13 .. 0.13]
503-> LEU A 4 HD* - TYR A 9 HE* [ 1.80 4.06] 0.00 0.00 0.00 0.00 0.00 0.00 0.00 0.00 0.00 0.01 0.00 0.00 0.27 0.00 0.00 0.00 0.00 0.00 0.04 0.03 - 4 [ 0.01 .. 0.27]
506-> VAL A 7 HN - VAL A 7 HG* [ 1.80 4.20] 0.02 0.02 0.05 0.04 0.00 0.00 0.00 0.00 0.02 0.00 0.04 0.02 0.06 0.00 0.01 0.06 0.01 0.13 0.00 0.00 - 12 [ 0.01 .. 0.13]
516-> PHE A 10 HB* - ALA A 62 HN [ 1.80 5.81] 0.01 0.03 0.00 0.34 0.13 0.00 0.00 0.24 0.03 0.18 0.02 0.15 0.00 0.34 0.00 0.13 0.21 0.00 0.21 0.05 - 14 [ 0.01 .. 0.34]
519-> PHE A 10 HE* - LEU A 59 HD* [ 1.80 3.82] 0.00 0.00 0.00 0.00 0.00 0.00 0.00 0.00 0.00 0.00 0.27 0.00 0.00 0.00 0.08 0.00 0.24 0.00 0.00 0.00 - 3 [ 0.08 .. 0.27]
521-> PHE A 10 HZ - VAL A 37 HG* [ 1.80 3.15] 0.30 0.26 0.14 0.12 0.22 0.28 0.37 0.38 0.36 0.29 0.42 0.09 0.32 0.17 0.33 0.24 0.39 0.13 0.21 0.37 - 20 [ 0.09 .. 0.42]
523-> ILE A 11 HN - GLU A 61 HG* [ 1.80 5.81] 0.06 0.00 0.00 0.10 0.26 0.02 0.12 0.26 0.05 0.00 0.05 0.04 0.00 0.18 0.00 0.00 0.09 0.00 0.00 0.15 - 12 [ 0.02 .. 0.26]
524-> ILE A 11 HB - GLU A 61 HG* [ 1.80 5.81] 0.10 0.00 0.11 0.00 0.17 0.07 0.09 0.02 0.09 0.00 0.04 0.06 0.30 0.10 0.10 0.00 0.00 0.05 0.00 0.13 - 14 [ 0.02 .. 0.30]
525-> ILE A 11 HG2* - GLU A 12 HG* [ 1.80 4.76] 0.00 0.00 0.00 0.18 0.17 0.16 0.06 0.00 0.00 0.00 0.00 0.00 0.00 0.00 0.12 0.20 0.00 0.00 0.00 0.00 - 6 [ 0.06 .. 0.20]
529-> ILE A 11 HG1* - ALA A 62 HN [ 1.80 3.89] 0.33 0.26 0.40 0.36 0.48 0.46 0.38 0.48 0.37 0.18 0.45 0.28 0.36 0.41 0.32 0.05 0.29 0.35 0.25 0.39 - 20 [ 0.05 .. 0.48]
531-> ILE A 11 HD1* - LEU A 14 HD* [ 1.80 4.10] 0.06 0.06 0.03 0.23 0.00 0.03 0.07 0.00 0.00 0.32 0.06 0.11 0.00 0.00 0.11 0.49 0.00 0.12 0.31 0.00 - 13 [ 0.03 .. 0.49]
535-> GLU A 12 HN - GLU A 12 HG* [ 1.80 3.70] 0.00 0.00 0.00 0.19 0.20 0.21 0.21 0.00 0.00 0.00 0.00 0.00 0.00 0.00 0.02 0.00 0.00 0.00 0.00 0.00 - 5 [ 0.02 .. 0.21]
537-> GLU A 12 HN - LEU A 14 HD* [ 1.80 4.62] 0.33 0.24 0.20 0.00 0.31 0.34 0.37 0.24 0.31 0.00 0.30 0.33 0.46 0.05 0.34 0.00 0.15 0.28 0.00 0.37 - 16 [ 0.05 .. 0.46]
545-> LEU A 14 HN - TYR A 33 HB* [ 1.80 5.81] 0.00 0.21 0.05 0.17 0.10 0.08 0.01 0.00 0.12 0.00 0.17 0.13 0.00 0.20 0.00 0.00 0.00 0.15 0.00 0.00 - 11 [ 0.01 .. 0.21]
549-> LEU A 14 HD* - LYS A 15 HN [ 1.80 3.80] 0.20 0.17 0.17 0.00 0.20 0.20 0.20 0.21 0.18 0.00 0.15 0.22 0.17 0.12 0.04 0.00 0.16 0.18 0.00 0.20 - 16 [ 0.04 .. 0.22]
551-> LEU A 14 HD* - ALA A 29 HN [ 1.80 5.92] 0.00 0.00 0.00 0.15 0.00 0.00 0.00 0.00 0.00 0.00 0.00 0.00 0.00 0.10 0.00 0.22 0.00 0.00 0.00 0.00 - 3 [ 0.10 .. 0.22]
567-> LEU A 14 HD* - VAL A 66 HB [ 1.80 3.94] 0.03 0.10 0.00 0.02 0.06 0.00 0.05 0.00 0.06 0.00 0.06 0.15 0.00 0.00 0.12 0.00 0.13 0.06 0.00 0.00 - 11 [ 0.02 .. 0.15]
574-> GLN A 16 HG* - ASN A 21 HN [ 1.80 4.45] 0.00 0.00 0.01 0.04 0.00 0.00 0.06 0.00 0.00 0.00 0.07 0.04 0.00 0.00 0.00 0.00 0.01 0.16 0.00 0.00 - 7 [ 0.01 .. 0.16]
576-> HIS A 17 HA - MET A 20 HG* [ 1.80 5.81] 0.00 0.00 0.00 0.00 0.02 0.14 0.17 0.14 0.13 0.11 0.15 0.00 0.05 0.00 0.00 0.00 0.00 0.00 0.00 0.00 - 8 [ 0.02 .. 0.17]
579-> ILE A 18 HA - LYS A 26 HB* [ 1.80 4.18] 0.09 0.11 0.00 0.43 0.00 0.01 0.02 0.19 0.02 0.05 0.00 0.00 0.02 0.02 0.10 0.16 0.00 0.05 0.15 0.19 - 16 [ 0.00 .. 0.43]
583-> ILE A 18 HD1* - LYS A 26 HG* [ 1.80 4.13] 0.08 0.04 0.10 0.12 0.09 0.09 0.02 0.00 0.13 0.09 0.16 0.17 0.15 0.07 0.00 0.00 0.19 0.38 0.08 0.13 - 17 [ 0.02 .. 0.38]
586-> GLU A 19 HN - LYS A 26 HG* [ 1.80 5.81] 0.24 0.36 0.07 0.00 0.15 0.18 0.23 0.28 0.24 0.24 0.21 0.24 0.30 0.29 0.45 0.27 0.29 0.22 0.22 0.24 - 19 [ 0.07 .. 0.45]
588-> MET A 20 HN - ASN A 21 HB* [ 1.80 5.81] 0.00 0.00 0.00 0.00 0.00 0.00 0.08 0.00 0.00 0.00 0.00 0.00 0.00 0.00 0.00 0.00 0.12 0.00 0.00 0.00 - 2 [ 0.08 .. 0.12]
590-> MET A 20 HB* - ASN A 21 HN [ 1.80 3.28] 0.02 0.00 0.00 0.05 0.00 0.00 0.00 0.00 0.06 0.00 0.07 0.00 0.07 0.00 0.06 0.00 0.00 0.12 0.01 0.05 - 11 [ 0.00 .. 0.12]
593-> ASN A 21 HN - GLN A 22 HB* [ 1.80 4.16] 0.03 0.04 0.00 0.00 0.00 0.00 0.17 0.04 0.00 0.00 0.00 0.00 0.00 0.04 0.00 0.02 0.00 0.00 0.00 0.00 - 6 [ 0.02 .. 0.17]
599-> GLN A 22 HB* - SER A 23 HN [ 1.80 3.59] 0.27 0.00 0.00 0.00 0.00 0.00 0.00 0.00 0.33 0.00 0.00 0.00 0.34 0.00 0.00 0.00 0.00 0.00 0.00 0.00 - 3 [ 0.27 .. 0.34]
603-> GLU A 24 HG* - ALA A 29 HN [ 1.80 4.36] 0.00 0.03 0.03 0.26 0.00 0.00 0.06 0.05 0.00 0.00 0.00 0.00 0.08 0.29 0.01 0.00 0.02 0.00 0.19 0.18 - 13 [ 0.00 .. 0.29]
606-> GLU A 24 HG* - TYR A 33 HN [ 1.80 4.31] 0.12 0.14 0.08 0.00 0.08 0.08 0.14 0.02 0.11 0.05 0.16 0.01 0.00 0.00 0.00 0.03 0.22 0.00 0.00 0.00 - 13 [ 0.01 .. 0.22]
617-> ILE A 27 HA - LYS A 67 HD* [ 1.80 4.95] 0.00 0.00 0.15 0.00 0.00 0.26 0.00 0.19 0.26 0.11 0.00 0.00 0.21 0.02 0.13 0.00 0.43 0.25 0.10 0.00 - 11 [ 0.02 .. 0.43]
620-> ILE A 27 HG1* - SER A 71 HN [ 1.80 5.81] 0.07 0.02 0.20 0.15 0.00 0.00 0.27 0.27 0.00 0.08 0.00 0.07 0.00 0.14 0.00 0.00 0.03 0.10 0.14 0.15 - 14 [ 0.00 .. 0.27]
624-> ALA A 29 HN - VAL A 66 HG* [ 1.80 4.13] 0.02 0.00 0.00 0.17 0.00 0.00 0.00 0.05 0.00 0.00 0.00 0.00 0.00 0.00 0.00 0.13 0.00 0.02 0.00 0.00 - 5 [ 0.02 .. 0.17]
626-> MET A 30 HN - TYR A 33 HB* [ 1.80 4.41] 0.28 0.19 0.20 0.14 0.19 0.21 0.19 0.22 0.13 0.18 0.14 0.20 0.14 0.14 0.10 0.22 0.19 0.04 0.26 0.33 - 20 [ 0.04 .. 0.33]
627-> MET A 30 HN - LYS A 67 HD* [ 1.80 5.77] 0.38 0.39 0.29 0.28 0.40 0.47 0.31 0.27 0.35 0.33 0.14 0.13 0.34 0.45 0.16 0.25 0.45 0.16 0.28 0.49 - 20 [ 0.13 .. 0.49]
629-> MET A 30 HA - VAL A 66 HG* [ 1.80 4.12] 0.00 0.00 0.00 0.00 0.00 0.00 0.00 0.00 0.00 0.00 0.00 0.00 0.00 0.15 0.00 0.00 0.00 0.00 0.00 0.00 - 1 [ 0.15 .. 0.15]
630-> ASN A 31 HN - TYR A 33 HB* [ 1.80 5.03] 0.03 0.00 0.01 0.00 0.03 0.00 0.00 0.00 0.05 0.10 0.00 0.07 0.14 0.20 0.00 0.01 0.00 0.03 0.01 0.00 - 11 [ 0.01 .. 0.20]
641-> TYR A 34 HE* - VAL A 37 HG* [ 1.80 5.11] 0.09 0.38 0.00 0.00 0.04 0.20 0.24 0.00 0.00 0.04 0.43 0.28 0.01 0.15 0.39 0.00 0.46 0.00 0.00 0.05 - 14 [ 0.00 .. 0.46]
652-> VAL A 37 HN - LEU A 59 HD* [ 1.80 4.49] 0.00 0.12 0.00 0.00 0.00 0.18 0.24 0.00 0.00 0.00 0.00 0.00 0.00 0.00 0.00 0.00 0.00 0.00 0.00 0.00 - 3 [ 0.12 .. 0.24]
660-> VAL A 37 HG* - LEU A 41 HN [ 1.80 3.85] 0.00 0.00 0.40 0.02 0.00 0.00 0.00 0.00 0.00 0.00 0.00 0.00 0.00 0.00 0.00 0.00 0.00 0.24 0.00 0.00 - 3 [ 0.02 .. 0.40]
663-> VAL A 37 HG* - VAL A 42 HN [ 1.80 5.25] 0.24 0.00 0.41 0.02 0.23 0.01 0.00 0.00 0.22 0.09 0.00 0.25 0.00 0.00 0.03 0.17 0.03 0.00 0.00 0.00 - 11 [ 0.01 .. 0.41]
665-> VAL A 37 HG* - LEU A 59 HN [ 1.80 5.52] 0.00 0.02 0.29 0.02 0.17 0.00 0.16 0.12 0.00 0.26 0.27 0.00 0.03 0.32 0.02 0.00 0.04 0.49 0.06 0.18 - 16 [ 0.00 .. 0.49]
668-> VAL A 38 HN - SER A 39 HB* [ 1.80 5.04] 0.00 0.00 0.00 0.00 0.05 0.10 0.00 0.00 0.00 0.00 0.00 0.00 0.00 0.05 0.00 0.00 0.00 0.05 0.00 0.00 - 4 [ 0.05 .. 0.10]
669-> VAL A 38 HN - LEU A 41 HD* [ 1.80 5.18] 0.00 0.00 0.00 0.00 0.00 0.00 0.00 0.00 0.00 0.00 0.24 0.00 0.00 0.00 0.00 0.00 0.22 0.00 0.00 0.00 - 2 [ 0.22 .. 0.24]
673-> VAL A 38 HG* - LEU A 59 HB2 [ 1.80 5.57] 0.00 0.00 0.00 0.04 0.00 0.00 0.00 0.24 0.00 0.00 0.00 0.00 0.00 0.00 0.00 0.00 0.00 0.00 0.00 0.00 - 2 [ 0.04 .. 0.24]
674-> VAL A 38 HG* - LEU A 59 HD* [ 1.80 3.42] 0.00 0.06 0.00 0.00 0.00 0.06 0.00 0.11 0.00 0.00 0.00 0.00 0.00 0.00 0.00 0.00 0.00 0.00 0.00 0.00 - 4 [ 0.00 .. 0.11]
678-> SER A 39 HB* - THR A 40 HN [ 1.80 3.30] 0.00 0.00 0.00 0.00 0.08 0.01 0.00 0.00 0.00 0.26 0.00 0.00 0.00 0.00 0.00 0.00 0.00 0.13 0.00 0.00 - 4 [ 0.01 .. 0.26]
683-> THR A 40 HN - LEU A 41 HD* [ 1.80 5.35] 0.00 0.00 0.00 0.00 0.00 0.00 0.00 0.03 0.00 0.00 0.00 0.00 0.00 0.18 0.00 0.00 0.00 0.00 0.09 0.00 - 3 [ 0.03 .. 0.18]
687-> THR A 40 HG2* - LEU A 41 HB* [ 1.80 4.60] 0.18 0.00 0.08 0.02 0.08 0.04 0.00 0.03 0.13 0.12 0.00 0.08 0.00 0.00 0.04 0.11 0.02 0.00 0.00 0.00 - 13 [ 0.00 .. 0.18]
694-> LEU A 41 HG - LEU A 59 HD* [ 1.80 5.92] 0.00 0.00 0.00 0.00 0.00 0.00 0.00 0.00 0.00 0.00 0.00 0.00 0.00 0.14 0.00 0.00 0.00 0.00 0.00 0.00 - 1 [ 0.14 .. 0.14]
697-> LEU A 41 HD* - ARG A 55 HN [ 1.80 5.10] 0.00 0.00 0.21 0.00 0.00 0.12 0.00 0.00 0.00 0.00 0.00 0.00 0.00 0.00 0.00 0.00 0.00 0.00 0.00 0.12 - 4 [ 0.00 .. 0.21]
698-> LEU A 41 HD* - ARG A 55 HA [ 1.80 4.81] 0.00 0.00 0.00 0.00 0.00 0.00 0.00 0.00 0.00 0.00 0.10 0.00 0.00 0.00 0.08 0.00 0.00 0.00 0.00 0.07 - 3 [ 0.07 .. 0.10]
701-> VAL A 42 HN - ASP A 44 HB* [ 1.80 5.81] 0.00 0.00 0.20 0.00 0.17 0.00 0.00 0.00 0.00 0.00 0.00 0.00 0.00 0.00 0.00 0.00 0.00 0.00 0.00 0.00 - 2 [ 0.17 .. 0.20]
708-> VAL A 42 HG* - ASP A 44 HN [ 1.80 4.47] 0.29 0.19 0.00 0.25 0.00 0.36 0.17 0.49 0.25 0.00 0.31 0.10 0.00 0.00 0.23 0.00 0.00 0.00 0.32 0.25 - 12 [ 0.10 .. 0.49]
709-> VAL A 42 HG* - ASP A 44 HB* [ 1.80 5.73] 0.07 0.18 0.20 0.20 0.03 0.07 0.17 0.00 0.00 0.00 0.00 0.00 0.00 0.00 0.28 0.00 0.00 0.00 0.00 0.10 - 9 [ 0.03 .. 0.28]
710-> VAL A 42 HG* - VAL A 52 HN [ 1.80 5.92] 0.00 0.00 0.00 0.00 0.19 0.00 0.00 0.00 0.11 0.14 0.00 0.09 0.05 0.11 0.05 0.04 0.12 0.01 0.00 0.00 - 10 [ 0.01 .. 0.19]
716-> ASP A 44 HB* - LEU A 46 HN [ 1.80 5.87] 0.42 0.00 0.00 0.00 0.00 0.00 0.00 0.00 0.00 0.00 0.00 0.00 0.20 0.00 0.00 0.00 0.00 0.06 0.00 0.00 - 3 [ 0.06 .. 0.42]
720-> GLN A 45 HG* - LEU A 46 HN [ 1.80 4.21] 0.00 0.00 0.00 0.00 0.22 0.00 0.00 0.00 0.00 0.00 0.00 0.00 0.00 0.00 0.00 0.00 0.00 0.18 0.00 0.00 - 2 [ 0.18 .. 0.22]
721-> GLN A 45 HG* - LEU A 46 HA [ 1.80 4.40] 0.00 0.00 0.00 0.00 0.19 0.00 0.00 0.00 0.00 0.00 0.00 0.00 0.00 0.00 0.00 0.00 0.00 0.09 0.00 0.00 - 2 [ 0.09 .. 0.19]
722-> GLN A 45 HG* - LEU A 46 HB* [ 1.80 5.61] 0.00 0.00 0.00 0.00 0.13 0.00 0.00 0.00 0.00 0.00 0.00 0.00 0.00 0.00 0.00 0.00 0.00 0.10 0.00 0.00 - 2 [ 0.10 .. 0.13]
723-> GLN A 45 HG* - LEU A 46 HG [ 1.80 5.81] 0.00 0.00 0.00 0.00 0.00 0.00 0.00 0.07 0.00 0.21 0.00 0.00 0.00 0.00 0.00 0.00 0.00 0.00 0.00 0.00 - 3 [ 0.00 .. 0.21]
727-> GLN A 45 HG* - VAL A 52 HN [ 1.80 5.84] 0.42 0.11 0.02 0.00 0.00 0.00 0.17 0.00 0.48 0.42 0.05 0.00 0.00 0.07 0.04 0.26 0.00 0.15 0.30 0.00 - 12 [ 0.02 .. 0.48]
728-> LEU A 46 HN - LEU A 46 HB* [ 1.80 3.08] 0.00 0.08 0.12 0.00 0.00 0.15 0.09 0.00 0.07 0.00 0.00 0.00 0.00 0.10 0.12 0.01 0.11 0.00 0.00 0.15 - 10 [ 0.01 .. 0.15]
733-> THR A 47 HN - VAL A 51 HG* [ 1.80 3.72] 0.28 0.17 0.26 0.18 0.11 0.16 0.28 0.00 0.10 0.15 0.01 0.00 0.00 0.16 0.20 0.00 0.00 0.12 0.21 0.16 - 15 [ 0.01 .. 0.28]
735-> THR A 47 HA - VAL A 51 HG* [ 1.80 4.87] 0.01 0.11 0.00 0.11 0.03 0.13 0.10 0.20 0.03 0.26 0.00 0.04 0.00 0.08 0.11 0.00 0.00 0.01 0.00 0.12 - 14 [ 0.01 .. 0.26]
737-> THR A 47 HB - VAL A 52 HG* [ 1.80 5.52] 0.01 0.00 0.00 0.00 0.00 0.00 0.00 0.00 0.00 0.00 0.00 0.15 0.00 0.00 0.00 0.00 0.16 0.06 0.00 0.00 - 4 [ 0.01 .. 0.16]
741-> LYS A 48 HN - LYS A 48 HG* [ 1.80 4.08] 0.22 0.03 0.10 0.21 0.10 0.24 0.10 0.01 0.10 0.18 0.11 0.10 0.06 0.22 0.23 0.18 0.21 0.06 0.12 0.24 - 20 [ 0.01 .. 0.24]
744-> LYS A 48 HB* - ASN A 49 HN [ 1.80 3.64] 0.11 0.08 0.00 0.06 0.08 0.03 0.06 0.08 0.00 0.13 0.00 0.05 0.00 0.04 0.10 0.07 0.11 0.08 0.00 0.04 - 15 [ 0.03 .. 0.13]
750-> LYS A 48 HG* - ALA A 50 HN [ 1.80 4.76] 0.00 0.00 0.11 0.00 0.07 0.00 0.00 0.04 0.06 0.00 0.02 0.18 0.08 0.00 0.00 0.07 0.03 0.00 0.12 0.00 - 11 [ 0.00 .. 0.18]
753-> LYS A 48 HG* - VAL A 51 HG* [ 1.80 3.37] 0.00 0.00 0.11 0.00 0.11 0.00 0.03 0.00 0.09 0.00 0.13 0.17 0.00 0.00 0.00 0.01 0.00 0.00 0.12 0.00 - 8 [ 0.01 .. 0.17]
754-> LYS A 48 HG* - VAL A 52 HN [ 1.80 4.79] 0.03 0.00 0.01 0.01 0.00 0.00 0.00 0.00 0.01 0.06 0.00 0.49 0.00 0.03 0.04 0.27 0.22 0.00 0.02 0.00 - 11 [ 0.01 .. 0.49]
756-> LYS A 48 HE* - ASN A 49 HA [ 1.80 5.81] 0.11 0.33 0.18 0.05 0.32 0.05 0.31 0.43 0.18 0.09 0.21 0.40 0.25 0.08 0.09 0.34 0.08 0.28 0.23 0.06 - 20 [ 0.05 .. 0.43]
762-> ASN A 49 HB* - ALA A 50 HN [ 1.80 3.56] 0.00 0.00 0.00 0.00 0.00 0.00 0.00 0.00 0.00 0.00 0.00 0.19 0.00 0.00 0.00 0.00 0.00 0.00 0.00 0.00 - 1 [ 0.19 .. 0.19]
769-> ALA A 50 HN - VAL A 52 HG* [ 1.80 4.38] 0.00 0.00 0.00 0.00 0.00 0.00 0.49 0.00 0.00 0.00 0.00 0.00 0.00 0.00 0.00 0.00 0.00 0.00 0.00 0.00 - 1 [ 0.49 .. 0.49]
776-> VAL A 51 HG* - VAL A 52 HN [ 1.80 3.46] 0.00 0.00 0.00 0.00 0.00 0.00 0.00 0.00 0.00 0.00 0.00 0.16 0.00 0.00 0.00 0.04 0.10 0.00 0.00 0.00 - 3 [ 0.04 .. 0.16]
787-> VAL A 52 HG* - LEU A 53 HB* [ 1.80 3.96] 0.00 0.00 0.00 0.00 0.00 0.00 0.20 0.00 0.00 0.00 0.00 0.00 0.00 0.00 0.00 0.00 0.00 0.00 0.00 0.00 - 1 [ 0.20 .. 0.20]
789-> VAL A 52 HG* - LYS A 54 HD* [ 1.80 5.34] 0.00 0.07 0.06 0.14 0.00 0.00 0.08 0.09 0.00 0.00 0.05 0.00 0.00 0.00 0.00 0.09 0.00 0.16 0.00 0.00 - 8 [ 0.05 .. 0.16]
794-> LEU A 53 HN - LYS A 54 HD* [ 1.80 5.39] 0.00 0.11 0.00 0.03 0.00 0.11 0.29 0.19 0.00 0.00 0.15 0.05 0.00 0.00 0.00 0.00 0.00 0.03 0.00 0.00 - 8 [ 0.03 .. 0.29]
795-> LEU A 53 HN - ILE A 56 HG1* [ 1.80 4.64] 0.31 0.00 0.29 0.17 0.00 0.25 0.00 0.00 0.17 0.00 0.00 0.00 0.24 0.00 0.22 0.23 0.00 0.00 0.23 0.00 - 9 [ 0.17 .. 0.31]
801-> LEU A 53 HD* - ILE A 56 HG1* [ 1.80 3.97] 0.04 0.00 0.00 0.17 0.03 0.43 0.00 0.00 0.00 0.00 0.00 0.21 0.04 0.00 0.05 0.11 0.00 0.00 0.23 0.08 - 10 [ 0.03 .. 0.43]
813-> ARG A 55 HN - GLN A 57 HG* [ 1.80 5.31] 0.14 0.06 0.12 0.28 0.00 0.03 0.02 0.13 0.05 0.12 0.02 0.00 0.07 0.11 0.18 0.22 0.08 0.09 0.06 0.04 - 18 [ 0.02 .. 0.28]
816-> ILE A 56 HG2* - LEU A 59 HD* [ 1.80 4.29] 0.00 0.24 0.00 0.00 0.00 0.17 0.20 0.00 0.00 0.00 0.00 0.00 0.00 0.00 0.00 0.00 0.00 0.00 0.00 0.00 - 3 [ 0.17 .. 0.24]
817-> ILE A 56 HG1* - GLN A 57 HN [ 1.80 4.04] 0.10 0.00 0.00 0.00 0.00 0.12 0.00 0.06 0.00 0.09 0.00 0.00 0.00 0.00 0.00 0.02 0.00 0.00 0.01 0.00 - 6 [ 0.01 .. 0.12]
820-> LEU A 59 HB2 - GLU A 61 HG* [ 1.80 5.81] 0.00 0.16 0.00 0.27 0.02 0.08 0.00 0.15 0.00 0.00 0.12 0.00 0.00 0.09 0.00 0.00 0.03 0.00 0.00 0.00 - 8 [ 0.02 .. 0.27]
849-> VAL A 66 HG* - GLU A 70 HG* [ 1.80 3.12] 0.19 0.00 0.00 0.00 0.00 0.00 0.00 0.00 0.00 0.00 0.00 0.00 0.00 0.00 0.49 0.13 0.00 0.00 0.00 0.00 - 3 [ 0.13 .. 0.49]
850-> LYS A 67 HN - GLU A 70 HB* [ 1.80 5.49] 0.05 0.00 0.00 0.00 0.00 0.36 0.00 0.00 0.00 0.00 0.00 0.00 0.00 0.00 0.00 0.00 0.00 0.00 0.00 0.00 - 2 [ 0.05 .. 0.36]
853-> LYS A 67 HB2 - ARG A 68 HD* [ 1.80 5.81] 0.00 0.00 0.00 0.23 0.21 0.05 0.06 0.07 0.00 0.00 0.00 0.00 0.00 0.00 0.00 0.00 0.13 0.17 0.10 0.00 - 8 [ 0.05 .. 0.23]
854-> LYS A 67 HD* - GLU A 70 HN [ 1.80 5.81] 0.26 0.00 0.09 0.26 0.42 0.15 0.00 0.36 0.18 0.07 0.09 0.00 0.19 0.00 0.41 0.05 0.41 0.30 0.03 0.35 - 16 [ 0.03 .. 0.42]
855-> ARG A 68 HN - ARG A 68 HG* [ 1.80 3.64] 0.00 0.00 0.40 0.29 0.28 0.28 0.30 0.29 0.00 0.40 0.32 0.42 0.00 0.00 0.41 0.34 0.27 0.27 0.27 0.00 - 14 [ 0.27 .. 0.42]
857-> ARG A 68 HN - GLY A 69 HA* [ 1.80 4.33] 0.25 0.00 0.21 0.21 0.26 0.10 0.06 0.15 0.23 0.21 0.00 0.00 0.23 0.26 0.02 0.00 0.27 0.24 0.22 0.25 - 17 [ 0.00 .. 0.27]
858-> ARG A 68 HN - GLU A 70 HB* [ 1.80 5.81] 0.00 0.00 0.00 0.00 0.00 0.49 0.13 0.00 0.00 0.00 0.02 0.00 0.00 0.00 0.20 0.00 0.00 0.00 0.00 0.00 - 4 [ 0.02 .. 0.49]
872-> GLU A 70 HA - GLU A 70 HG* [ 1.80 3.08] 0.00 0.34 0.00 0.00 0.00 0.23 0.34 0.00 0.00 0.00 0.33 0.34 0.00 0.00 0.20 0.33 0.00 0.00 0.00 0.00 - 7 [ 0.20 .. 0.34]
873-> SER A 71 HN - SER A 71 HB* [ 1.80 3.06] 0.05 0.00 0.17 0.16 0.10 0.00 0.11 0.00 0.00 0.00 0.00 0.10 0.00 0.00 0.00 0.00 0.00 0.00 0.00 0.12 - 7 [ 0.05 .. 0.17]
874-> SER A 71 HN - LYS A 72 HG* [ 1.80 5.81] 0.00 0.00 0.00 0.00 0.00 0.00 0.00 0.00 0.00 0.00 0.00 0.00 0.00 0.00 0.39 0.00 0.00 0.00 0.00 0.00 - 1 [ 0.39 .. 0.39]
875-> SER A 71 HA - LYS A 72 HB* [ 1.80 4.87] 0.00 0.00 0.00 0.00 0.00 0.00 0.00 0.00 0.00 0.00 0.00 0.00 0.00 0.00 0.00 0.00 0.10 0.39 0.00 0.00 - 2 [ 0.10 .. 0.39]
876-> SER A 71 HB* - LYS A 72 HN [ 1.80 3.89] 0.14 0.00 0.00 0.00 0.00 0.12 0.00 0.15 0.00 0.00 0.00 0.00 0.16 0.01 0.00 0.00 0.02 0.05 0.00 0.00 - 7 [ 0.01 .. 0.16]
879-> LYS A 72 HN - LYS A 72 HG* [ 1.80 4.16] 0.00 0.00 0.00 0.00 0.00 0.00 0.00 0.00 0.00 0.00 0.00 0.00 0.00 0.00 0.00 0.00 0.00 0.00 0.00 0.20 - 1 [ 0.20 .. 0.20]
882-> LYS A 72 HB* - LEU A 73 HN [ 1.80 3.90] 0.00 0.00 0.00 0.00 0.00 0.00 0.00 0.00 0.00 0.03 0.06 0.05 0.00 0.07 0.00 0.13 0.00 0.05 0.04 0.00 - 7 [ 0.03 .. 0.13]
884-> LEU A 73 HN - LEU A 73 HB* [ 1.80 3.00] 0.00 0.00 0.00 0.00 0.00 0.00 0.00 0.00 0.37 0.00 0.00 0.00 0.00 0.00 0.00 0.00 0.00 0.10 0.00 0.00 - 2 [ 0.10 .. 0.37]
888-> LEU A 73 HA - GLU A 74 HB* [ 1.80 5.34] 0.00 0.00 0.00 0.00 0.00 0.20 0.00 0.00 0.00 0.00 0.00 0.00 0.00 0.00 0.04 0.01 0.00 0.00 0.00 0.00 - 3 [ 0.01 .. 0.20]
894-> LEU A 73 HD* - HIS A 75 HB* [ 1.80 5.73] 0.00 0.00 0.00 0.00 0.00 0.00 0.00 0.00 0.00 0.00 0.05 0.00 0.00 0.04 0.00 0.00 0.02 0.27 0.00 0.00 - 4 [ 0.02 .. 0.27]
895-> GLU A 74 HN - GLU A 74 HB* [ 1.80 3.19] 0.25 0.05 0.00 0.13 0.00 0.00 0.00 0.00 0.11 0.00 0.00 0.00 0.12 0.07 0.00 0.00 0.08 0.07 0.00 0.00 - 8 [ 0.05 .. 0.25]
-------------------------------------------
Number of Violations greater than 0.10 53 49 54 59 43 54 53 48 55 46 51 52 45 49 57 51 52 54 52 47
-------------------------------------------
---- Summary Of Residual Distance Constraint Violations ----
Mod 1 Mod 2 Mod 3 Mod 4 Mod 5 Mod 6 Mod 7 Mod 8 Mod 9 Mod 10 Mod 11 Mod 12 Mod 13 Mod 14 Mod 15 Mod 16 Mod 17 Mod 18 Mod 19 Mod 20 Averages
~~~~~~ ~~~~~~ ~~~~~~ ~~~~~~ ~~~~~~ ~~~~~~ ~~~~~~ ~~~~~~ ~~~~~~ ~~~~~~ ~~~~~~ ~~~~~~ ~~~~~~ ~~~~~~ ~~~~~~ ~~~~~~ ~~~~~~ ~~~~~~ ~~~~~~ ~~~~~~ ~~~~~~~~~~~
0.1 - 0.2 ang: 26 26 28 31 22 25 23 23 30 24 28 23 17 26 31 22 28 31 20 33 25.85
0.2 - 0.5 ang: 27 23 26 28 21 29 30 25 25 22 23 29 28 23 26 29 24 23 32 14 25.35
> 0.5 ang: 0 0 0 0 0 0 0 0 0 0 0 0 0 0 0 0 0 0 0 0 0.00
Total : 100 88 95 103 82 103 93 92 96 82 105 101 93 92 117 94 102 105 99 87 96.45
Minimum Violation : 0.000 0.000 0.000 0.000 0.000 0.000 0.000 0.000 0.000 0.000 0.000 0.000 0.000 0.000 0.000 0.000 0.000 0.000 0.000 0.000 0.000
Maximum Violation : 0.422 0.486 0.414 0.434 0.475 0.488 0.495 0.490 0.478 0.415 0.454 0.491 0.489 0.494 0.494 0.490 0.455 0.490 0.374 0.493 0.495
Max Intra Viol : 0.349 0.361 0.402 0.290 0.280 0.282 0.367 0.397 0.366 0.403 0.327 0.424 0.161 0.230 0.411 0.340 0.274 0.311 0.374 0.236 0.424
Max Seque Viol : 0.370 0.486 0.226 0.336 0.315 0.324 0.316 0.426 0.449 0.270 0.331 0.398 0.415 0.334 0.485 0.373 0.325 0.447 0.245 0.458 0.486
Max Medium Viol : 0.420 0.384 0.404 0.335 0.434 0.488 0.495 0.490 0.460 0.323 0.432 0.491 0.458 0.422 0.494 0.490 0.455 0.419 0.357 0.375 0.495
Max Long Viol : 0.422 0.388 0.414 0.434 0.475 0.469 0.377 0.480 0.478 0.415 0.454 0.466 0.489 0.494 0.450 0.381 0.454 0.490 0.300 0.493 0.494
Average Violation : 0.016 0.014 0.015 0.016 0.012 0.016 0.016 0.015 0.016 0.013 0.015 0.016 0.015 0.014 0.017 0.015 0.015 0.016 0.014 0.012 0.01486
Avge Intra Viol : 0.006 0.007 0.008 0.008 0.005 0.008 0.009 0.005 0.007 0.007 0.008 0.010 0.003 0.005 0.008 0.011 0.006 0.007 0.007 0.005 0.00702
Avge Seque Viol : 0.020 0.016 0.022 0.020 0.012 0.023 0.019 0.017 0.018 0.015 0.017 0.020 0.017 0.015 0.027 0.018 0.017 0.015 0.022 0.015 0.01824
Avge Mediu Viol : 0.012 0.010 0.005 0.009 0.009 0.010 0.010 0.010 0.013 0.008 0.007 0.011 0.012 0.009 0.009 0.010 0.009 0.013 0.006 0.007 0.00943
Avge Long Viol : 0.026 0.025 0.026 0.029 0.026 0.026 0.027 0.029 0.028 0.023 0.030 0.028 0.030 0.032 0.023 0.025 0.033 0.029 0.026 0.026 0.02729
RMS Violation : 0.060 0.058 0.057 0.058 0.052 0.063 0.060 0.059 0.063 0.052 0.056 0.063 0.061 0.057 0.062 0.059 0.061 0.059 0.055 0.051 0.05841
RMS Intra : 0.037 0.042 0.045 0.037 0.028 0.040 0.049 0.037 0.036 0.043 0.043 0.053 0.018 0.026 0.042 0.049 0.030 0.035 0.038 0.028 0.03868
RMS Sequential : 0.071 0.057 0.070 0.063 0.053 0.079 0.067 0.065 0.066 0.055 0.060 0.072 0.065 0.054 0.082 0.065 0.067 0.056 0.071 0.054 0.06509
RMS Medium range : 0.050 0.057 0.030 0.043 0.041 0.042 0.046 0.045 0.060 0.039 0.038 0.052 0.056 0.040 0.047 0.049 0.039 0.053 0.033 0.040 0.04571
RMS Long range : 0.077 0.076 0.076 0.084 0.079 0.082 0.076 0.084 0.083 0.071 0.083 0.075 0.089 0.096 0.069 0.074 0.095 0.085 0.070 0.078 0.08026
Final --global-- Summary for 20 models, 898 NOEs/model, 17960 NOEs total
~~~~~~~~~~~~~~~~~~~~~~~~~~~~~~~~~~~~~~~~~~~~~~~~~~~~~~~~~~~~~~~~~~~~~~~~~~~
Summ of viol : 266.870
Summ sq. viol : 61.280
Maximum viol : 0.495
Average viol : 0.01486
RMSD viol : 0.05841
Std. Dev. viol : 0.05649
RMS Intra : 0.03868
RMS Seque : 0.06509
RMS Medi : 0.04571
RMS Long : 0.08026
table of dihedral angle constraints violations
1-> [VAL A 7] PHI -72.0 -36.0 0.0 0.0 0.0 0.0 0.0 0.0 0.0 0.5 0.0 0.0 0.0 0.0 1.8 0.0 0.0 0.0 0.0 0.0 0.0 0.0 - 2 [ 0.0 .. 1.8]
5-> [PHE A 10] PHI -81.0 -49.0 0.0 0.0 0.0 0.0 0.0 0.0 0.0 0.0 0.0 0.0 0.0 0.0 3.1 0.0 0.2 2.0 0.0 0.0 0.0 0.0 - 3 [ 0.0 .. 3.1]
6-> [PHE A 10] PSI -51.0 -31.0 0.0 0.0 0.0 0.0 0.0 0.0 0.0 0.0 0.0 0.0 0.0 0.0 1.5 0.0 0.0 0.6 0.0 0.0 0.0 0.0 - 2 [ 0.0 .. 1.5]
7-> [ILE A 11] PHI -75.0 -55.0 0.0 0.0 0.0 0.0 0.0 0.0 0.0 0.0 0.0 0.0 0.0 0.0 1.3 0.0 0.0 0.0 0.0 0.0 0.0 0.0 - 1 [ 0.0 .. 1.3]
9-> [GLU A 12] PHI -75.0 -55.0 4.2 3.9 3.1 8.7 5.7 3.6 4.3 3.0 6.8 2.9 3.0 5.6 5.7 7.7 6.6 2.9 5.9 4.4 3.4 6.4 - 20 [ 2.9 .. 8.7]
10-> [GLU A 12] PSI -51.0 -27.0 0.0 0.0 0.0 0.0 1.8 0.7 1.9 0.6 3.0 3.0 1.2 0.9 0.0 0.0 2.9 5.3 2.9 0.0 1.8 3.3 - 13 [ 0.0 .. 5.3]
11-> [ASN A 13] PHI -73.0 -53.0 0.0 0.0 0.0 0.0 2.6 0.0 0.7 0.0 1.5 0.0 0.2 0.0 0.0 1.2 0.0 0.0 1.9 0.0 0.0 1.2 - 7 [ 0.0 .. 2.6]
12-> [ASN A 13] PSI -52.0 -32.0 0.0 0.0 0.0 0.0 0.0 0.0 0.0 1.1 0.0 0.0 0.0 0.0 0.0 1.5 0.0 0.0 2.0 0.0 0.0 0.0 - 3 [ 0.0 .. 2.0]
13-> [LEU A 14] PHI -73.0 -49.0 0.0 0.2 0.0 3.5 0.0 0.0 0.0 0.0 0.0 0.5 0.0 0.0 0.0 0.6 0.0 0.0 0.0 0.0 0.0 0.0 - 4 [ 0.0 .. 3.5]
14-> [LEU A 14] PSI -57.0 -37.0 0.0 0.0 0.0 0.0 0.0 0.0 0.0 0.0 0.0 0.1 0.0 0.0 0.0 0.0 0.0 0.0 0.0 1.8 0.0 0.0 - 2 [ 0.0 .. 1.8]
16-> [LYS A 15] PSI -52.0 -32.0 0.0 0.0 0.0 0.0 0.0 0.0 0.0 0.0 0.0 0.0 0.0 0.0 0.0 0.0 4.2 0.0 0.0 0.0 0.0 0.0 - 1 [ 0.0 .. 4.2]
18-> [GLN A 16] PSI -54.0 -30.0 0.0 0.0 0.0 0.1 0.0 0.0 3.2 0.6 0.0 0.0 0.0 0.0 0.0 2.6 2.5 2.9 0.0 0.0 3.4 0.0 - 7 [ 0.0 .. 3.4]
19-> [HIS A 17] PHI -80.0 -52.0 0.0 0.0 0.0 0.0 0.0 0.0 0.0 0.0 0.0 0.0 0.0 0.0 0.0 0.0 1.9 0.0 0.0 0.0 0.0 0.0 - 1 [ 0.0 .. 1.9]
20-> [HIS A 17] PSI -52.0 -26.0 0.0 0.0 0.0 4.0 0.3 0.7 0.7 1.5 0.0 0.0 0.0 0.0 0.0 0.9 8.7 1.3 0.0 0.0 1.7 0.4 - 10 [ 0.0 .. 8.7]
21-> [ILE A 18] PHI -74.0 -54.0 0.0 0.0 0.0 0.0 0.0 0.0 0.2 0.0 0.0 0.0 0.0 0.4 0.0 0.0 3.1 0.0 0.0 0.0 0.0 0.0 - 3 [ 0.0 .. 3.1]
24-> [GLU A 19] PSI -51.0 -25.0 0.7 2.8 0.9 2.4 0.0 0.0 0.0 4.6 2.9 0.0 4.3 1.2 1.2 0.0 0.0 2.1 3.3 3.0 2.0 3.4 - 14 [ 0.0 .. 4.6]
28-> [HIS A 28] PSI -79.0 17.0 0.0 0.0 0.0 0.0 0.0 0.0 0.0 0.0 0.0 0.0 0.0 0.0 2.4 0.9 0.0 0.0 0.0 0.0 3.0 3.5 - 4 [ 0.0 .. 3.5]
29-> [ALA A 29] PHI -74.0 -50.0 0.0 4.9 2.9 0.4 1.5 1.1 0.4 0.0 4.6 3.9 4.2 1.1 5.7 0.0 0.0 0.8 6.4 0.2 0.0 0.0 - 14 [ 0.0 .. 6.4]
32-> [MET A 30] PSI -53.0 -33.0 0.0 0.0 0.0 0.0 0.0 0.0 0.0 0.0 0.0 0.0 0.0 0.0 0.0 0.0 1.6 0.0 0.0 1.8 0.0 0.1 - 3 [ 0.0 .. 1.8]
33-> [ASN A 31] PHI -78.0 -56.0 2.5 8.7 2.4 0.0 3.6 3.5 3.9 2.3 2.4 5.8 4.4 4.1 4.6 9.5 0.0 0.1 2.9 4.4 4.6 4.3 - 18 [ 0.0 .. 9.5]
35-> [SER A 32] PHI -84.0 -52.0 0.0 0.0 0.0 0.0 0.0 0.0 0.0 0.0 0.0 0.0 0.0 0.0 0.0 5.2 0.0 0.0 0.0 0.0 0.0 0.0 - 1 [ 0.0 .. 5.2]
36-> [SER A 32] PSI -51.0 -23.0 0.0 0.0 3.1 0.4 3.9 4.3 3.2 2.6 2.6 4.4 2.4 3.1 3.3 6.1 0.4 2.4 0.2 3.5 3.5 1.9 - 18 [ 0.0 .. 6.1]
39-> [TYR A 34] PHI -73.0 -49.0 1.8 0.9 0.0 0.0 0.0 0.0 0.0 0.0 0.0 0.0 3.7 0.0 0.0 0.0 0.0 0.0 0.0 0.0 0.0 0.0 - 3 [ 0.0 .. 3.7]
40-> [TYR A 34] PSI -55.0 -31.0 1.7 5.3 0.0 0.0 0.0 0.0 1.8 0.0 0.0 0.0 2.6 0.0 0.0 0.0 0.0 0.0 0.4 0.0 0.0 0.0 - 5 [ 0.0 .. 5.3]
41-> [ARG A 35] PHI -72.0 -48.0 0.0 0.0 3.6 0.0 0.0 0.0 0.0 0.0 0.0 0.0 0.0 0.0 0.0 0.0 2.7 0.0 0.0 5.2 0.3 0.0 - 4 [ 0.0 .. 5.2]
42-> [ARG A 35] PSI -63.0 -27.0 1.7 0.7 6.6 0.0 0.0 0.0 0.0 0.0 0.0 0.0 3.2 0.0 0.0 0.0 0.0 0.0 2.3 0.4 0.0 0.0 - 6 [ 0.0 .. 6.6]
46-> [VAL A 37] PSI -57.0 -33.0 6.0 0.0 0.0 3.4 0.0 6.1 0.0 4.2 0.0 0.0 0.0 0.0 0.0 0.0 0.0 6.1 0.0 0.0 6.8 0.0 - 6 [ 0.0 .. 6.8]
47-> [VAL A 38] PHI -75.0 -51.0 4.7 0.0 0.0 0.0 0.0 1.9 0.0 3.1 1.4 0.0 1.0 0.0 1.0 0.0 2.5 3.8 1.5 4.4 3.8 0.0 - 11 [ 0.0 .. 4.7]
50-> [SER A 39] PSI -59.0 -19.0 0.0 0.6 0.0 0.0 0.0 0.0 0.0 0.0 0.0 0.0 0.0 0.0 0.3 1.3 0.0 0.0 0.0 0.0 0.1 0.0 - 4 [ 0.0 .. 1.3]
51-> [THR A 40] PHI -83.0 -51.0 0.0 4.5 0.0 0.0 0.0 0.0 3.3 0.7 0.0 1.2 0.0 0.0 5.6 0.0 0.0 0.0 0.0 2.8 0.0 0.0 - 6 [ 0.0 .. 5.6]
54-> [LEU A 41] PSI -49.0 -29.0 6.1 0.0 0.0 0.0 0.0 0.0 0.0 0.0 0.0 0.0 0.0 0.0 0.0 1.2 0.0 0.0 0.0 0.0 0.0 0.0 - 2 [ 0.0 .. 6.1]
56-> [VAL A 42] PSI -50.0 22.0 0.0 5.3 0.0 4.1 0.0 7.8 2.9 0.0 0.0 0.0 4.9 0.0 0.0 0.0 3.9 0.0 0.0 0.0 0.0 4.6 - 7 [ 0.0 .. 7.8]
57-> [ASN A 49] PHI -70.0 -50.0 0.7 0.0 0.0 1.4 1.6 0.0 0.0 0.0 0.0 3.3 0.0 0.0 1.2 0.3 0.9 1.0 0.0 0.0 0.0 0.0 - 8 [ 0.0 .. 3.3]
58-> [ASN A 49] PSI -64.0 -12.0 0.0 0.0 0.0 0.0 0.0 0.0 0.0 0.0 0.0 0.0 0.0 1.8 0.0 0.0 0.0 0.0 0.0 0.0 0.0 0.0 - 1 [ 0.0 .. 1.8]
59-> [ALA A 50] PHI -74.0 -54.0 0.0 0.0 0.0 0.0 0.0 0.0 0.0 0.0 0.0 0.0 0.0 0.0 0.0 0.0 0.0 4.4 1.9 0.0 0.0 0.0 - 2 [ 0.0 .. 4.4]
60-> [ALA A 50] PSI -50.0 -24.0 0.0 0.0 0.0 3.2 0.0 0.0 0.0 0.0 0.0 0.0 0.0 1.7 0.2 0.0 0.0 4.8 7.0 0.0 0.0 0.0 - 5 [ 0.0 .. 7.0]
61-> [VAL A 51] PHI -80.0 -52.0 5.1 0.0 0.0 3.8 0.2 0.0 0.0 1.2 0.0 0.0 0.0 0.0 0.0 0.0 0.0 2.0 5.4 2.9 0.8 0.0 - 8 [ 0.0 .. 5.4]
62-> [VAL A 51] PSI -56.0 -34.0 0.0 0.0 9.2 0.0 0.0 0.0 0.0 0.0 8.9 0.0 0.0 0.4 0.0 0.0 0.0 0.0 0.0 0.0 9.1 0.0 - 4 [ 0.0 .. 9.2]
63-> [VAL A 52] PHI -75.0 -55.0 8.2 3.1 6.3 0.8 2.8 1.5 0.0 1.3 1.2 4.3 1.6 3.2 3.7 6.1 0.0 2.5 6.7 0.4 5.9 3.5 - 18 [ 0.0 .. 8.2]
64-> [VAL A 52] PSI -55.0 -35.0 0.4 0.0 2.9 0.0 0.0 0.0 0.0 3.0 0.0 0.0 0.0 1.8 0.0 0.0 0.0 0.0 1.2 0.0 3.1 0.0 - 6 [ 0.0 .. 3.1]
65-> [LEU A 53] PHI -70.0 -50.0 0.0 0.0 0.0 0.0 0.0 0.0 0.0 0.0 0.0 0.0 0.0 4.7 0.0 0.0 0.0 0.0 0.0 0.0 0.0 0.0 - 1 [ 0.0 .. 4.7]
66-> [LEU A 53] PSI -58.0 -18.0 0.0 0.0 0.0 0.0 0.0 0.0 0.0 0.0 0.0 0.0 0.6 0.0 0.0 0.0 0.0 0.0 0.0 0.0 0.0 2.0 - 2 [ 0.0 .. 2.0]
67-> [LYS A 54] PHI -74.0 -54.0 0.0 3.8 0.0 1.2 0.9 0.0 1.2 0.0 1.9 0.0 3.5 2.0 0.0 3.0 0.0 0.0 1.4 0.7 0.0 4.8 - 11 [ 0.0 .. 4.8]
69-> [ARG A 55] PHI -74.0 -54.0 0.0 0.0 0.0 0.0 0.0 0.0 0.0 0.0 0.0 0.0 0.0 1.7 0.0 0.0 0.0 0.0 0.0 0.0 0.0 0.0 - 1 [ 0.0 .. 1.7]
72-> [ILE A 56] PSI -51.0 -31.0 0.0 0.1 0.6 4.6 0.1 0.0 0.1 0.0 1.8 0.0 1.5 0.0 0.0 2.0 0.9 0.0 0.0 0.7 0.0 0.0 - 11 [ 0.0 .. 4.6]
77-> [LEU A 59] PHI -74.0 -54.0 0.0 0.0 0.5 0.0 0.0 0.0 0.0 0.0 1.8 0.4 0.0 0.6 5.2 1.0 0.0 0.5 3.8 2.5 0.0 0.0 - 9 [ 0.0 .. 5.2]
78-> [LEU A 59] PSI -52.0 -24.0 0.0 0.0 1.4 0.0 0.0 0.0 0.0 0.0 0.0 0.0 0.0 0.0 1.4 0.3 0.0 0.0 0.0 0.0 0.0 0.0 - 3 [ 0.0 .. 1.4]
79-> [ASP A 60] PHI -73.0 -53.0 0.4 0.0 0.0 0.2 0.0 2.8 1.8 0.5 1.7 1.2 1.6 2.7 0.0 0.0 5.8 1.4 0.0 0.1 1.6 0.0 - 13 [ 0.0 .. 5.8]
80-> [ASP A 60] PSI -54.0 -30.0 0.1 0.0 0.0 0.0 0.0 0.1 3.1 0.0 0.0 0.8 0.0 3.8 3.3 0.0 4.5 1.6 0.0 1.6 0.0 0.0 - 9 [ 0.0 .. 4.5]
81-> [GLU A 61] PHI -73.0 -53.0 0.0 0.0 0.0 1.6 0.0 0.0 0.0 0.0 0.0 0.0 0.0 0.0 0.0 2.3 0.0 0.0 0.0 0.0 0.0 0.2 - 3 [ 0.0 .. 2.3]
83-> [ALA A 62] PHI -71.0 -51.0 0.0 1.6 2.1 0.0 0.0 2.7 2.1 0.0 0.0 0.3 0.0 2.0 0.0 0.0 0.0 0.0 0.0 0.6 0.0 6.8 - 8 [ 0.0 .. 6.8]
84-> [ALA A 62] PSI -58.0 -34.0 0.1 0.0 0.0 4.3 0.0 0.0 0.0 3.7 0.0 0.0 0.0 0.0 0.0 4.2 0.0 2.1 0.3 0.0 1.7 0.0 - 8 [ 0.0 .. 4.3]
85-> [TYR A 63] PHI -74.0 -50.0 0.0 0.3 1.0 0.0 0.0 0.0 0.0 0.0 0.0 0.0 0.0 0.5 0.0 0.0 3.8 0.0 0.0 1.0 0.0 0.0 - 5 [ 0.0 .. 3.8]
87-> [ASN A 64] PHI -71.0 -51.0 0.0 0.0 0.0 0.0 0.0 0.0 0.0 0.0 0.0 0.0 0.0 0.0 0.0 0.0 0.0 0.0 0.0 0.0 0.0 2.4 - 1 [ 0.0 .. 2.4]
92-> [VAL A 66] PSI -60.0 -28.0 0.0 0.0 0.0 0.0 0.0 0.0 0.0 0.0 0.0 0.0 0.0 0.0 0.0 0.0 0.0 0.0 0.0 2.3 0.0 0.0 - 1 [ 0.0 .. 2.3]
---- ACOSummary Of Residual ACO Constraint Violations ----
Mod 1 Mod 2 Mod 3 Mod 4 Mod 5 Mod 6 Mod 7 Mod 8 Mod 9 Mod 10 Mod 11 Mod 12 Mod 13 Mod 14 Mod 15 Mod 16 Mod 17 Mod 18 Mod 19 Mod 20 Averages
~~~~~~ ~~~~~~ ~~~~~~ ~~~~~~ ~~~~~~ ~~~~~~ ~~~~~~ ~~~~~~ ~~~~~~ ~~~~~~ ~~~~~~ ~~~~~~ ~~~~~~ ~~~~~~ ~~~~~~ ~~~~~~ ~~~~~~ ~~~~~~ ~~~~~~ ~~~~~~ ~~~~~~~~~~~
1 - 10. degrees : 10 10 12 13 8 10 12 12 14 9 15 15 16 15 14 17 16 14 15 13 13.00
> 10. degrees : 0 0 0 0 0 0 0 0 0 0 0 0 0 0 0 0 0 0 0 0 0.00
Total : 16 16 15 20 12 13 17 17 15 14 17 20 19 21 20 21 21 22 19 18 17.65
Minimum Violation : 0.0 0.0 0.0 0.0 0.0 0.0 0.0 0.0 0.0 0.0 0.0 0.0 0.0 0.0 0.0 0.0 0.0 0.0 0.0 0.0 0.00
Maximum Violation : 8.2 8.7 9.2 8.7 5.7 7.8 4.3 4.6 8.9 5.8 4.9 5.6 5.7 9.5 8.7 6.1 7.0 5.2 9.1 6.8 9.45
Max PHI Viol : 8.2 8.7 6.3 8.7 5.7 3.6 4.3 3.1 6.8 5.8 4.4 5.6 5.7 9.5 6.6 4.4 6.7 5.2 5.9 6.8 9.45
Max PSI Viol : 6.1 5.3 9.2 4.6 3.9 7.8 3.2 4.6 8.9 4.4 4.9 3.8 3.3 6.1 8.7 6.1 7.0 3.5 9.1 4.6 9.21
Average Violation : 0.5 0.5 0.5 0.5 0.3 0.4 0.4 0.4 0.4 0.3 0.5 0.5 0.5 0.6 0.6 0.5 0.6 0.5 0.6 0.5 0.473
Avge PHI Viol : 0.758 0.816 0.676 0.672 0.627 0.598 0.612 0.511 0.698 0.705 0.694 0.772 0.902 0.885 0.755 0.669 0.902 0.789 0.665 0.793 0.732
Avge PSI Viol : 0.593 0.556 0.716 0.747 0.357 0.642 0.593 0.676 0.631 0.416 0.657 0.555 0.530 0.661 0.792 0.782 0.641 0.561 0.869 0.632 0.641
RMS Violation : 1.522 1.542 1.550 1.482 0.937 1.312 1.023 1.024 1.421 1.115 1.237 1.192 1.477 1.776 1.665 1.360 1.659 1.231 1.678 1.485 1.405
RMS PHI Viol : 1.725 1.844 1.307 1.510 1.171 0.997 1.070 0.763 1.356 1.368 1.271 1.446 1.916 2.190 1.575 1.124 1.960 1.508 1.339 1.758 1.501
RMS PSI Viol : 1.288 1.165 1.760 1.454 0.621 1.565 0.975 1.230 1.483 0.783 1.202 0.867 0.832 1.230 1.751 1.562 1.290 0.870 1.960 1.150 1.301
Final --global-- Summary for 20 models, 96 ACOs/model, 1920 ACOs total
~~~~~~~~~~~~~~~~~~~~~~~~~~~~~~~~~~~~~~~~~~~~~~~~~~~~~~~~~~~~~~~~~~~~~~~~~~~
Summ. Viol. : 909.03
Summ. Sq. Viol. : 3787.75
Max. Viol. : 9.451
Avg. Viol. : 0.47345
RMS Viol. : 1.40456
Std. Dev. Viol. : 1.32236
JPEG image for inter-residue distance constraints per residue plot
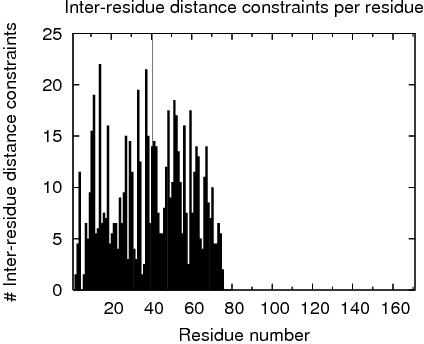
S(phi)|S(psi) V/S Residue number
Text output from PDBStat of phi psi order
# CHAIN .GT. SUM.GT.
# RES ID DIH S(phi) S(psi) S(chi1) S(chi2) S(chi3) S(chi4) S(chi5) 0.90 1.6
# -----------------------------------------------------------------------------------
MET M 1 0.245 0.499 0.406 0.604
SER M 2 0.230 0.163 0.061
GLU M 3 0.340 0.737 0.518 0.527 0.886
LEU M 4 0.125 0.475 0.435 0.607
PHE M 5 0.755 0.735 0.519 0.132
SER M 6 0.686 0.857 0.064
VAL M 7 0.988 0.995 0.918 7 7
PRO M 8 0.993 0.998 0.973 0.926 8 8
TYR M 9 0.997 0.997 0.996 0.605 9 9
PHE M 10 0.991 0.997 0.987 0.911 10 10
ILE M 11 0.997 0.998 0.994 0.991 11 11
GLU M 12 1.000 0.997 0.692 0.520 0.913 12 12
ASN M 13 0.996 0.998 0.566 0.843 13 13
LEU M 14 0.998 0.999 0.987 0.794 14 14
LYS M 15 0.999 0.998 0.651 0.739 0.565 0.344 15 15
GLN M 16 0.998 0.998 0.999 0.999 0.851 16 16
HIS M 17 0.995 0.999 0.924 0.054 17 17
ILE M 18 0.996 0.999 0.992 0.998 18 18
GLU M 19 0.997 0.994 0.818 0.286 0.937 19 19
MET M 20 0.986 0.820 0.717 0.888 0.454 20
ASN M 21 0.720 0.990 0.728 0.449
GLN M 22 0.991 0.843 0.934 0.295 0.362 22
SER M 23 0.796 0.978 0.425
GLU M 24 0.989 0.982 0.973 0.997 0.891 24 24
ASP M 25 0.985 0.980 0.997 0.998 25 25
LYS M 26 0.989 0.997 0.937 0.999 0.728 0.194 26 26
ILE M 27 0.999 0.997 0.660 0.507 27 27
HIS M 28 0.992 0.980 0.747 0.293 28 28
ALA M 29 0.985 0.998 29 29
MET M 30 0.999 0.996 0.823 0.472 0.633 30 30
ASN M 31 0.999 0.996 0.639 0.299 31 31
SER M 32 0.992 1.000 0.188 32 32
TYR M 33 0.999 0.984 0.990 0.503 33 33
TYR M 34 0.990 0.995 0.996 0.234 34 34
ARG M 35 0.996 0.996 0.501 0.666 0.334 0.370 1.000 35 35
SER M 36 0.998 0.999 0.326 36 36
VAL M 37 0.999 0.994 0.830 37 37
VAL M 38 0.998 0.998 0.544 38 38
SER M 39 0.997 0.978 0.263 39 39
THR M 40 0.986 0.997 0.997 40 40
LEU M 41 0.996 0.993 0.763 0.547 41 41
VAL M 42 0.984 0.885 0.836 42
GLN M 43 0.814 0.899 0.711 0.823 0.693 43
ASP M 44 0.617 0.211 0.576 0.828
GLN M 45 0.105 0.416 0.550 0.761 0.421
LEU M 46 0.394 0.960 0.814 0.905
THR M 47 0.908 0.994 0.706 47 47
LYS M 48 0.995 0.995 0.992 0.765 0.730 0.524 48 48
ASN M 49 0.997 0.986 0.646 0.144 49 49
ALA M 50 0.996 0.999 50 50
VAL M 51 0.998 0.989 0.752 51 51
VAL M 52 0.981 0.996 0.929 52 52
LEU M 53 0.997 0.991 0.897 0.294 53 53
LYS M 54 0.997 0.998 0.306 0.858 0.100 0.250 54 54
ARG M 55 0.999 0.999 0.672 0.349 0.163 0.580 1.000 55 55
ILE M 56 0.999 0.999 0.994 0.444 56 56
GLN M 57 0.999 0.998 0.997 0.573 0.410 57 57
HIS M 58 0.998 0.995 0.458 0.339 58 58
LEU M 59 0.996 0.996 0.982 0.618 59 59
ASP M 60 0.998 0.992 0.244 0.810 60 60
GLU M 61 0.998 0.996 0.995 0.999 0.585 61 61
ALA M 62 0.999 0.996 62 62
TYR M 63 0.999 0.999 0.979 0.419 63 63
ASN M 64 0.999 0.995 0.800 0.444 64 64
LYS M 65 0.997 0.998 0.798 0.228 0.865 0.224 65 65
VAL M 66 0.998 0.996 0.950 66 66
LYS M 67 0.999 0.999 0.999 0.938 0.847 0.673 67 67
ARG M 68 0.983 0.987 0.763 0.696 0.459 0.568 1.000 68 68
GLY M 69 0.986 0.957 69 69
GLU M 70 0.890 0.569 0.503 0.338 0.838
SER M 71 0.335 0.510 0.247
LYS M 72 0.440 0.294 0.561 0.707 0.572 0.044
LEU M 73 0.326 0.426 0.481 0.577
GLU M 74 0.367 0.492 0.630 0.786 0.872
HIS M 75 0.143 0.501 0.335 0.156
HIS M 76 0.211 0.375 0.392 0.119
HIS M 77 0.756 0.196 0.468 0.573
HIS M 78 0.390 0.492 0.423 0.043
HIS M 79 0.565 0.443 0.400 0.126
HIS M 80 0.559 0.290 0.399 0.204
MET M 91 0.327 0.375 0.852 0.351 0.674
SER M 92 0.484 0.186 0.223
GLU M 93 0.430 0.710 0.317 0.474 0.893
LEU M 94 0.232 0.478 0.585 0.418
PHE M 95 0.754 0.577 0.542 0.417
SER M 96 0.505 0.850 0.216
VAL M 97 0.987 0.995 0.918 97 97
PRO M 98 0.992 0.996 0.955 0.879 98 98
TYR M 99 0.995 0.997 0.997 0.595 99 99
PHE M 100 0.994 0.999 0.970 0.834 100 100
ILE M 101 0.998 0.999 0.993 0.997 101 101
GLU M 102 0.999 0.998 0.635 0.602 0.917 102 102
ASN M 103 0.997 0.998 0.542 0.177 103 103
LEU M 104 0.998 0.999 0.983 0.672 104 104
LYS M 105 0.999 0.996 0.711 0.838 0.477 0.625 105 105
GLN M 106 0.997 0.998 0.955 0.996 0.710 106 106
HIS M 107 0.996 0.999 0.927 0.135 107 107
ILE M 108 0.996 0.998 0.995 0.999 108 108
GLU M 109 0.995 0.991 0.830 0.417 0.888 109 109
MET M 110 0.967 0.748 0.682 0.523 0.397
ASN M 111 0.536 0.990 0.203 0.518
GLN M 112 0.990 0.985 0.604 0.599 0.599 112 112
SER M 113 0.981 0.923 0.274 113 113
GLU M 114 0.904 0.995 0.997 1.000 0.933 114 114
ASP M 115 0.994 0.962 0.997 0.983 115 115
LYS M 116 0.957 0.999 1.000 0.998 0.417 0.277 116 116
ILE M 117 1.000 0.998 0.645 0.495 117 117
HIS M 118 0.996 0.979 0.872 0.190 118 118
ALA M 119 0.978 0.996 119 119
MET M 120 0.998 0.997 0.883 0.491 0.800 120 120
ASN M 121 0.999 0.996 0.557 0.320 121 121
SER M 122 0.995 0.998 0.481 122 122
TYR M 123 0.995 0.982 1.000 0.311 123 123
TYR M 124 0.991 0.997 0.998 0.603 124 124
ARG M 125 0.998 0.995 0.588 0.705 0.271 0.520 1.000 125 125
SER M 126 0.998 0.998 0.302 126 126
VAL M 127 0.999 0.990 0.834 127 127
VAL M 128 0.998 0.998 0.358 128 128
SER M 129 0.997 0.982 0.213 129 129
THR M 130 0.993 0.999 0.996 130 130
LEU M 131 0.996 0.995 0.786 0.425 131 131
VAL M 132 0.985 0.940 0.943 132 132
GLN M 133 0.865 0.813 0.693 0.702 0.812 133
ASP M 134 0.723 0.413 0.549 0.837
GLN M 135 0.186 0.145 0.406 0.803 0.415
LEU M 136 0.178 0.963 0.735 0.869
THR M 137 0.923 0.992 0.915 137 137
LYS M 138 0.989 0.992 0.993 0.753 0.944 0.598 138 138
ASN M 139 0.996 0.986 0.591 0.231 139 139
ALA M 140 0.992 0.996 140 140
VAL M 141 0.997 0.990 0.750 141 141
VAL M 142 0.988 0.997 0.813 142 142
LEU M 143 0.998 0.986 0.895 0.427 143 143
LYS M 144 0.998 0.998 0.336 0.825 0.272 0.251 144 144
ARG M 145 0.999 0.998 0.836 0.258 0.471 0.605 1.000 145 145
ILE M 146 0.999 0.999 0.992 0.506 146 146
GLN M 147 0.998 0.995 0.997 0.661 0.585 147 147
HIS M 148 0.998 0.994 0.806 0.199 148 148
LEU M 149 0.991 0.995 0.891 0.354 149 149
ASP M 150 0.998 0.991 0.500 0.682 150 150
GLU M 151 0.998 0.998 0.992 0.998 0.540 151 151
ALA M 152 0.999 0.997 152 152
TYR M 153 0.999 0.999 0.994 0.526 153 153
ASN M 154 0.999 0.999 0.824 0.403 154 154
LYS M 155 0.999 0.999 0.759 0.254 0.999 0.236 155 155
VAL M 156 0.999 0.995 0.999 156 156
LYS M 157 0.998 0.993 0.996 0.915 0.991 0.599 157 157
ARG M 158 0.974 0.982 0.689 0.487 0.249 0.523 1.000 158 158
GLY M 159 0.814 0.877 159
GLU M 160 0.928 0.437 0.410 0.239 0.774
SER M 161 0.146 0.270 0.330
LYS M 162 0.151 0.462 0.862 0.807 0.526 0.230
LEU M 163 0.528 0.137 0.336 0.442
GLU M 164 0.295 0.344 0.822 0.642 0.927
HIS M 165 0.248 0.308 0.619 0.341
HIS M 166 0.507 0.359 0.551 0.257
HIS M 167 0.317 0.293 0.403 0.167
HIS M 168 0.390 0.608 0.562 0.565
HIS M 169 0.505 0.024 0.569 0.280
HIS M 170 0.525 0.405 0.361
JPEG image of S(phi)~Residue_number Plot
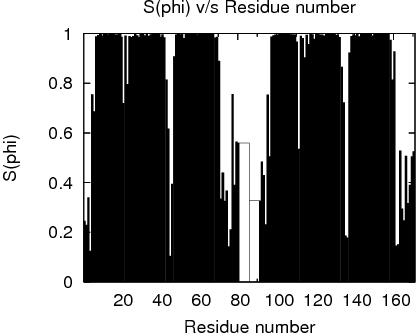
JPEG image of S(psi)~Residue_number Plot
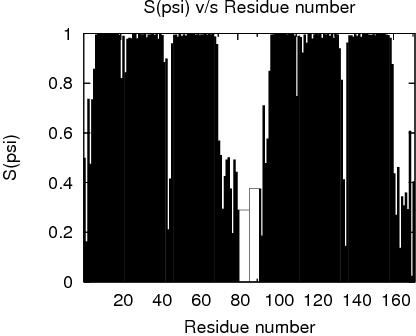
Table of Backbone and Heavy Atom RMSD
Text report of backbone and heavy atom RMSD for ordered regions
>
> Kabsch RMSD data for family `SR478_NMR_em_bcr3.pdb'
>
> Kabsch RMSD of backbone atoms in res. M[7..19],M[24..41],M[47..69],M[97..109],M[112..132],M[137..158],for model 1 is: 0.813
> Kabsch RMSD of backbone atoms in res. M[7..19],M[24..41],M[47..69],M[97..109],M[112..132],M[137..158],for model 2 is: 0.761
> Kabsch RMSD of backbone atoms in res. M[7..19],M[24..41],M[47..69],M[97..109],M[112..132],M[137..158],for model 3 is: 0.825
> Kabsch RMSD of backbone atoms in res. M[7..19],M[24..41],M[47..69],M[97..109],M[112..132],M[137..158],for model 4 is: 0.723
> Kabsch RMSD of backbone atoms in res. M[7..19],M[24..41],M[47..69],M[97..109],M[112..132],M[137..158],for model 5 is: 0.710
> Kabsch RMSD of backbone atoms in res. M[7..19],M[24..41],M[47..69],M[97..109],M[112..132],M[137..158],for model 6 is: 0.352 (*)
> Kabsch RMSD of backbone atoms in res. M[7..19],M[24..41],M[47..69],M[97..109],M[112..132],M[137..158],for model 7 is: 0.442
> Kabsch RMSD of backbone atoms in res. M[7..19],M[24..41],M[47..69],M[97..109],M[112..132],M[137..158],for model 8 is: 0.747
> Kabsch RMSD of backbone atoms in res. M[7..19],M[24..41],M[47..69],M[97..109],M[112..132],M[137..158],for model 9 is: 0.620
> Kabsch RMSD of backbone atoms in res. M[7..19],M[24..41],M[47..69],M[97..109],M[112..132],M[137..158],for model 10 is: 0.531
> Kabsch RMSD of backbone atoms in res. M[7..19],M[24..41],M[47..69],M[97..109],M[112..132],M[137..158],for model 11 is: 0.893
> Kabsch RMSD of backbone atoms in res. M[7..19],M[24..41],M[47..69],M[97..109],M[112..132],M[137..158],for model 12 is: 1.129
> Kabsch RMSD of backbone atoms in res. M[7..19],M[24..41],M[47..69],M[97..109],M[112..132],M[137..158],for model 13 is: 0.987
> Kabsch RMSD of backbone atoms in res. M[7..19],M[24..41],M[47..69],M[97..109],M[112..132],M[137..158],for model 14 is: 0.662
> Kabsch RMSD of backbone atoms in res. M[7..19],M[24..41],M[47..69],M[97..109],M[112..132],M[137..158],for model 15 is: 0.873
> Kabsch RMSD of backbone atoms in res. M[7..19],M[24..41],M[47..69],M[97..109],M[112..132],M[137..158],for model 16 is: 0.892
> Kabsch RMSD of backbone atoms in res. M[7..19],M[24..41],M[47..69],M[97..109],M[112..132],M[137..158],for model 17 is: 0.885
> Kabsch RMSD of backbone atoms in res. M[7..19],M[24..41],M[47..69],M[97..109],M[112..132],M[137..158],for model 18 is: 0.784
> Kabsch RMSD of backbone atoms in res. M[7..19],M[24..41],M[47..69],M[97..109],M[112..132],M[137..158],for model 19 is: 0.577
> Kabsch RMSD of backbone atoms in res. M[7..19],M[24..41],M[47..69],M[97..109],M[112..132],M[137..158],for model 20 is: 0.626
>
> Kabsch RMSD statistics for 20 structures:
> Mean RMSD using as refer. str. `average' for res.[7..19],[24..41],[47..69],[97..109],[112..132],[137..158], is: 0.742
> Range of RMSD values to reference struct. is 0.352 to 1.129
> Kabsch RMSD of heavy atoms in res. M[7..19],M[24..41],M[47..69],M[97..109],M[112..132],M[137..158],for model 1 is: 1.155
> Kabsch RMSD of heavy atoms in res. M[7..19],M[24..41],M[47..69],M[97..109],M[112..132],M[137..158],for model 2 is: 1.276
> Kabsch RMSD of heavy atoms in res. M[7..19],M[24..41],M[47..69],M[97..109],M[112..132],M[137..158],for model 3 is: 1.191
> Kabsch RMSD of heavy atoms in res. M[7..19],M[24..41],M[47..69],M[97..109],M[112..132],M[137..158],for model 4 is: 1.115
> Kabsch RMSD of heavy atoms in res. M[7..19],M[24..41],M[47..69],M[97..109],M[112..132],M[137..158],for model 5 is: 1.141
> Kabsch RMSD of heavy atoms in res. M[7..19],M[24..41],M[47..69],M[97..109],M[112..132],M[137..158],for model 6 is: 0.982 (*)
> Kabsch RMSD of heavy atoms in res. M[7..19],M[24..41],M[47..69],M[97..109],M[112..132],M[137..158],for model 7 is: 1.072
> Kabsch RMSD of heavy atoms in res. M[7..19],M[24..41],M[47..69],M[97..109],M[112..132],M[137..158],for model 8 is: 1.124
> Kabsch RMSD of heavy atoms in res. M[7..19],M[24..41],M[47..69],M[97..109],M[112..132],M[137..158],for model 9 is: 1.035
> Kabsch RMSD of heavy atoms in res. M[7..19],M[24..41],M[47..69],M[97..109],M[112..132],M[137..158],for model 10 is: 1.043
> Kabsch RMSD of heavy atoms in res. M[7..19],M[24..41],M[47..69],M[97..109],M[112..132],M[137..158],for model 11 is: 1.232
> Kabsch RMSD of heavy atoms in res. M[7..19],M[24..41],M[47..69],M[97..109],M[112..132],M[137..158],for model 12 is: 1.402
> Kabsch RMSD of heavy atoms in res. M[7..19],M[24..41],M[47..69],M[97..109],M[112..132],M[137..158],for model 13 is: 1.448
> Kabsch RMSD of heavy atoms in res. M[7..19],M[24..41],M[47..69],M[97..109],M[112..132],M[137..158],for model 14 is: 1.163
> Kabsch RMSD of heavy atoms in res. M[7..19],M[24..41],M[47..69],M[97..109],M[112..132],M[137..158],for model 15 is: 1.305
> Kabsch RMSD of heavy atoms in res. M[7..19],M[24..41],M[47..69],M[97..109],M[112..132],M[137..158],for model 16 is: 1.269
> Kabsch RMSD of heavy atoms in res. M[7..19],M[24..41],M[47..69],M[97..109],M[112..132],M[137..158],for model 17 is: 1.329
> Kabsch RMSD of heavy atoms in res. M[7..19],M[24..41],M[47..69],M[97..109],M[112..132],M[137..158],for model 18 is: 1.170
> Kabsch RMSD of heavy atoms in res. M[7..19],M[24..41],M[47..69],M[97..109],M[112..132],M[137..158],for model 19 is: 1.020
> Kabsch RMSD of heavy atoms in res. M[7..19],M[24..41],M[47..69],M[97..109],M[112..132],M[137..158],for model 20 is: 1.111
>
> Kabsch RMSD statistics for 20 structures:
> Mean RMSD using as refer. str. `average' for res.[7..19],[24..41],[47..69],[97..109],[112..132],[137..158], is: 1.179
> Range of RMSD values to reference struct. is 0.982 to 1.448
Text report of backbone RMSD for entire protein
> Kabsch RMSD of backb atoms in res. *[1..170],for model 1 is: 1.941
> Kabsch RMSD of backb atoms in res. *[1..170],for model 2 is: 1.731
> Kabsch RMSD of backb atoms in res. *[1..170],for model 3 is: 1.803
> Kabsch RMSD of backb atoms in res. *[1..170],for model 4 is: 1.619 (*)
> Kabsch RMSD of backb atoms in res. *[1..170],for model 5 is: 2.150
> Kabsch RMSD of backb atoms in res. *[1..170],for model 6 is: 1.758
> Kabsch RMSD of backb atoms in res. *[1..170],for model 7 is: 2.020
> Kabsch RMSD of backb atoms in res. *[1..170],for model 8 is: 1.638
> Kabsch RMSD of backb atoms in res. *[1..170],for model 9 is: 1.833
> Kabsch RMSD of backb atoms in res. *[1..170],for model 10 is: 1.652
> Kabsch RMSD of backb atoms in res. *[1..170],for model 11 is: 2.303
> Kabsch RMSD of backb atoms in res. *[1..170],for model 12 is: 2.190
> Kabsch RMSD of backb atoms in res. *[1..170],for model 13 is: 2.388
> Kabsch RMSD of backb atoms in res. *[1..170],for model 14 is: 1.932
> Kabsch RMSD of backb atoms in res. *[1..170],for model 15 is: 2.115
> Kabsch RMSD of backb atoms in res. *[1..170],for model 16 is: 3.425
> Kabsch RMSD of backb atoms in res. *[1..170],for model 17 is: 1.940
> Kabsch RMSD of backb atoms in res. *[1..170],for model 18 is: 2.201
> Kabsch RMSD of backb atoms in res. *[1..170],for model 19 is: 2.227
> Kabsch RMSD of backb atoms in res. *[1..170],for model 20 is: 2.190
>
> Kabsch RMSD statistics for 20 structures:
> Mean RMSD using as refer. str. `average' for res.[1..170], is: 2.053
> Range of RMSD values to reference struct. is 1.619 to 3.425
Text report of heavy atom RMSD for entire protein
> Kabsch RMSD of heavy atoms in res. *[1..170],for model 1 is: 2.554
> Kabsch RMSD of heavy atoms in res. *[1..170],for model 2 is: 2.450
> Kabsch RMSD of heavy atoms in res. *[1..170],for model 3 is: 2.375
> Kabsch RMSD of heavy atoms in res. *[1..170],for model 4 is: 2.331
> Kabsch RMSD of heavy atoms in res. *[1..170],for model 5 is: 2.701
> Kabsch RMSD of heavy atoms in res. *[1..170],for model 6 is: 2.398
> Kabsch RMSD of heavy atoms in res. *[1..170],for model 7 is: 2.670
> Kabsch RMSD of heavy atoms in res. *[1..170],for model 8 is: 2.375
> Kabsch RMSD of heavy atoms in res. *[1..170],for model 9 is: 2.498
> Kabsch RMSD of heavy atoms in res. *[1..170],for model 10 is: 2.318 (*)
> Kabsch RMSD of heavy atoms in res. *[1..170],for model 11 is: 2.826
> Kabsch RMSD of heavy atoms in res. *[1..170],for model 12 is: 2.764
> Kabsch RMSD of heavy atoms in res. *[1..170],for model 13 is: 2.951
> Kabsch RMSD of heavy atoms in res. *[1..170],for model 14 is: 2.625
> Kabsch RMSD of heavy atoms in res. *[1..170],for model 15 is: 2.811
> Kabsch RMSD of heavy atoms in res. *[1..170],for model 16 is: 3.995
> Kabsch RMSD of heavy atoms in res. *[1..170],for model 17 is: 2.590
> Kabsch RMSD of heavy atoms in res. *[1..170],for model 18 is: 2.686
> Kabsch RMSD of heavy atoms in res. *[1..170],for model 19 is: 2.871
> Kabsch RMSD of heavy atoms in res. *[1..170],for model 20 is: 2.875
>
> Kabsch RMSD statistics for 20 structures:
> Mean RMSD using as refer. str. `average' for res.[1..170], is: 2.683
> Range of RMSD values to reference struct. is 2.318 to 3.995
Summary of heavy atom and backbone RMSDs over the whole protein and ordered residues
RMSD Values
all residues ordered residues selected residues
All backbone atoms 2.1 0.7 0.7
All heavy atoms 2.7 1.2 1.2
Contact Map (constraints list and 3D Coordinates)
JPEG image of Contact Map for Constraints
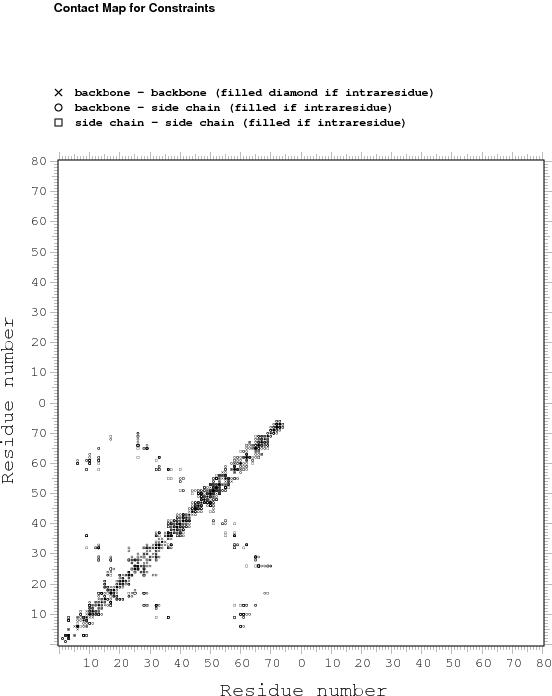
JPEG image of Contact Map for Coordinates
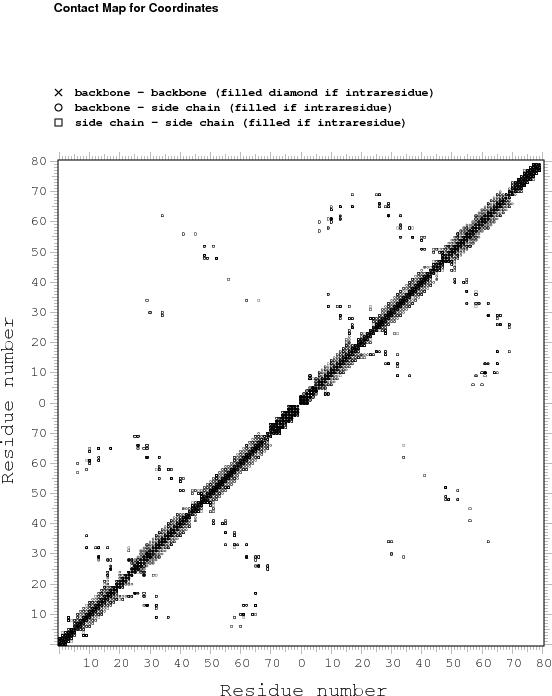
Output from PROCHECK
Ramachandran Plot for all models
Text summary of Ramachandran Plot
+----------<<< P R O C H E C K S U M M A R Y >>>----------+
| |
| SR478_NMR_em_bcr3_020.rin 0.0 2240 residues |
| |
+| Ramachandran plot: 88.9% core 7.2% allow 4.0% gener 0.0% disall |
| |
*| All Ramachandrans: 132 labelled residues (out of2240) |
*| Chi1-chi2 plots: 79 labelled residues (out of1560) |
JPEG image for all model Ramachandran Plot
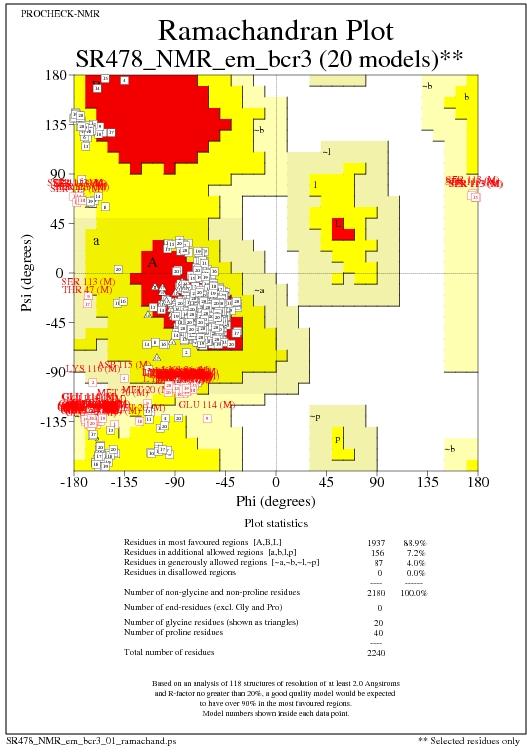
Residue Properties for all models
JPEG for all model Residue Properties - page $num_n
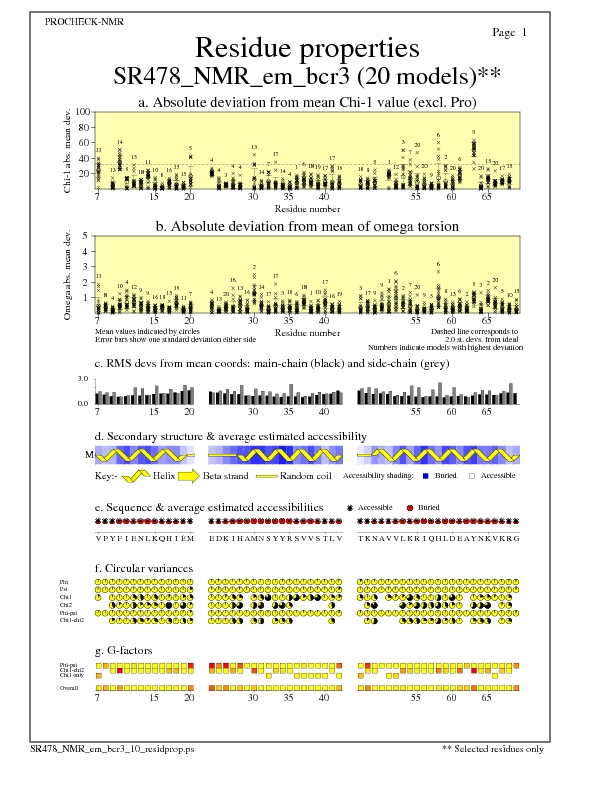
JPEG for all model Residue Properties - page $num_n
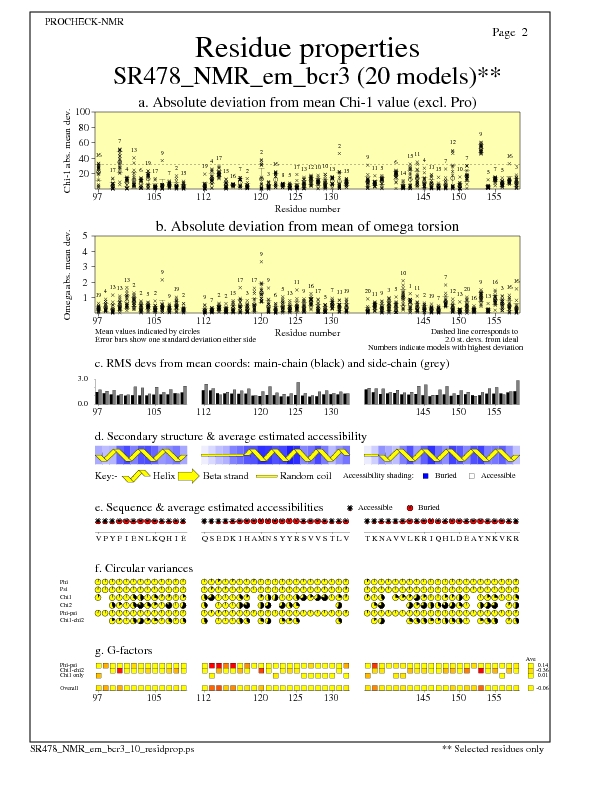
Model Secondary Structures from Procheck
JPEG for Model Secondary Structures - page $num_n

JPEG for Model Secondary Structures - page $num_n

JPEG for Model Secondary Structures - page $num_n
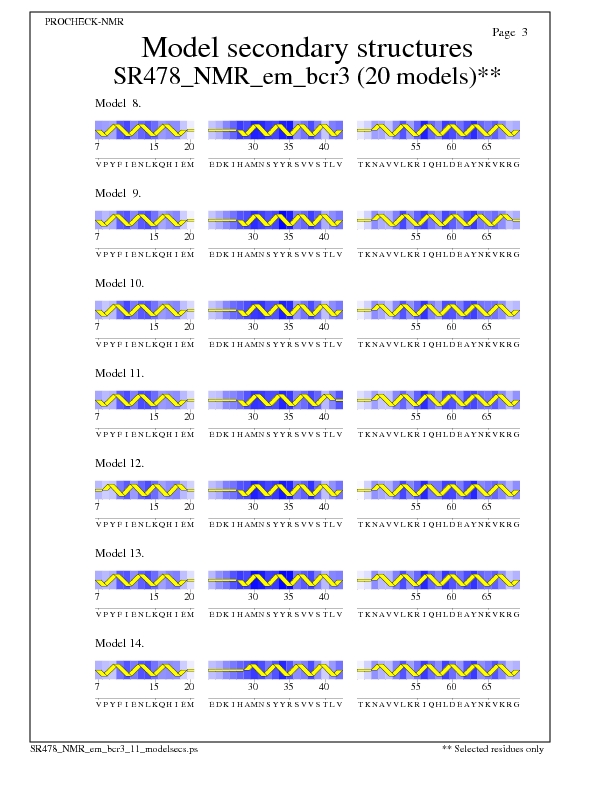
JPEG for Model Secondary Structures - page $num_n

JPEG for Model Secondary Structures - page $num_n

JPEG for Model Secondary Structures - page $num_n

Ramachandran Plots for each residue
JPEG for residue Ramachandran Plots - page $num_n
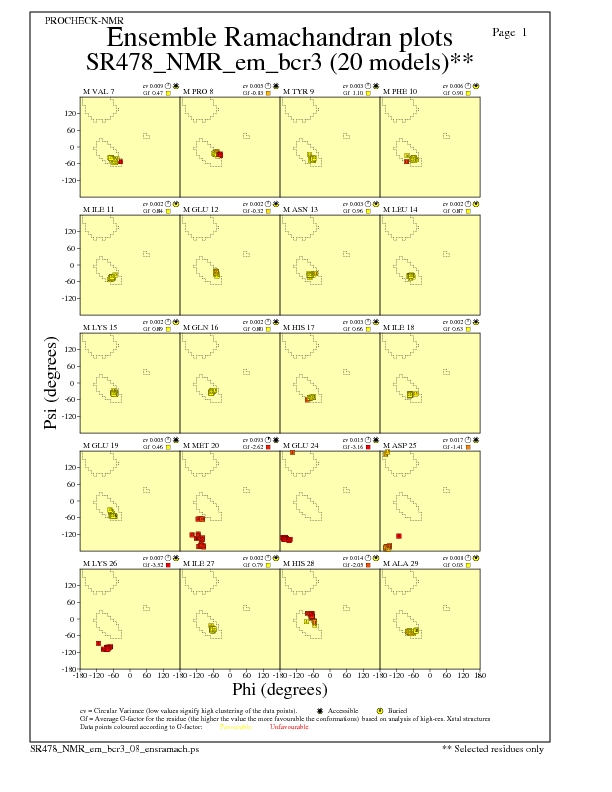
JPEG for residue Ramachandran Plots - page $num_n
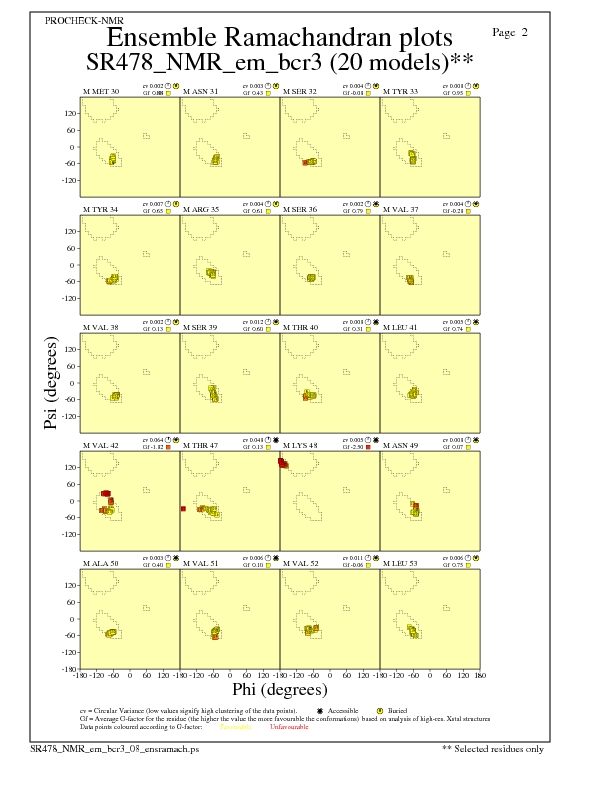
JPEG for residue Ramachandran Plots - page $num_n
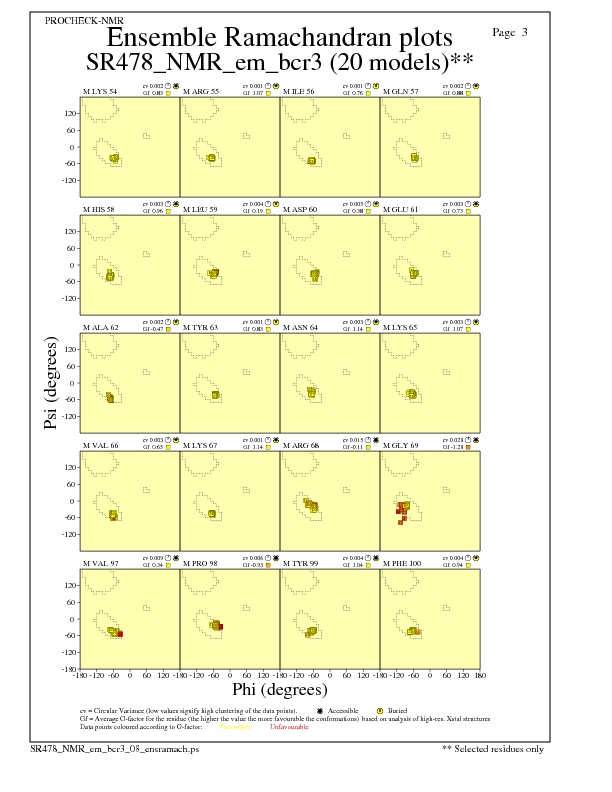
JPEG for residue Ramachandran Plots - page $num_n
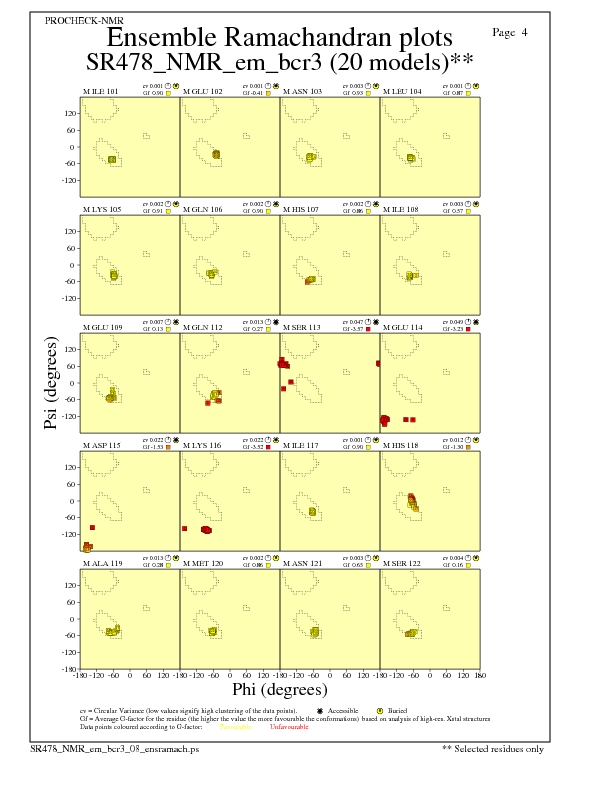
JPEG for residue Ramachandran Plots - page $num_n
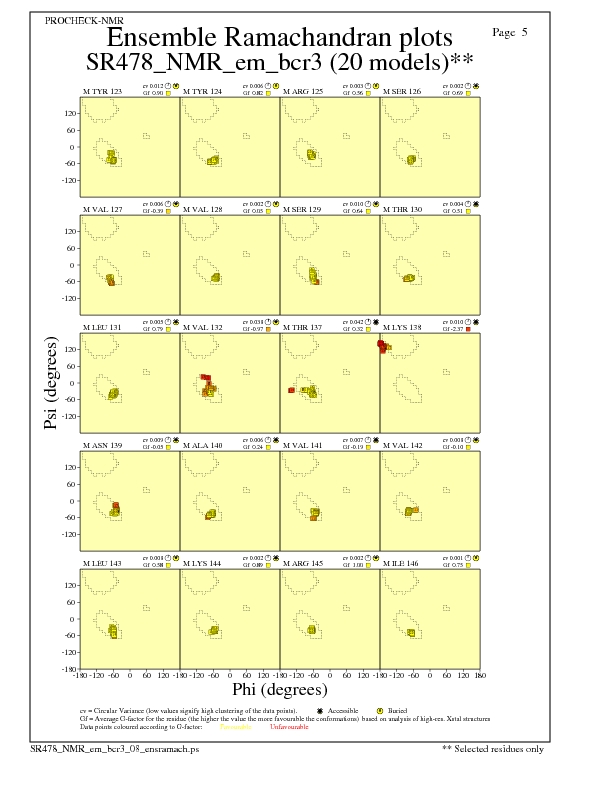
JPEG for residue Ramachandran Plots - page $num_n
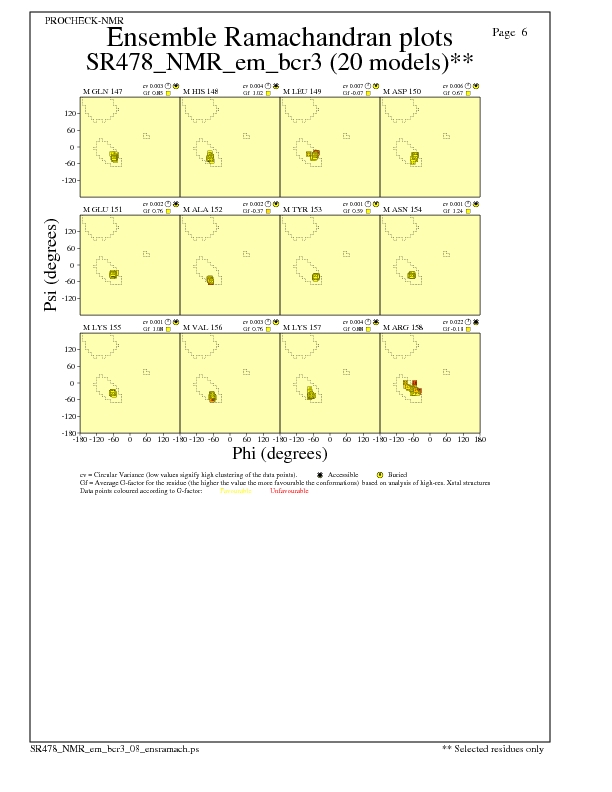
Ramachandran analysis for each residue from Molprobity
Chi1-Chi2 Plots for each residue
JPEG for residue Chi1-Chi2 Plots - page $num_n
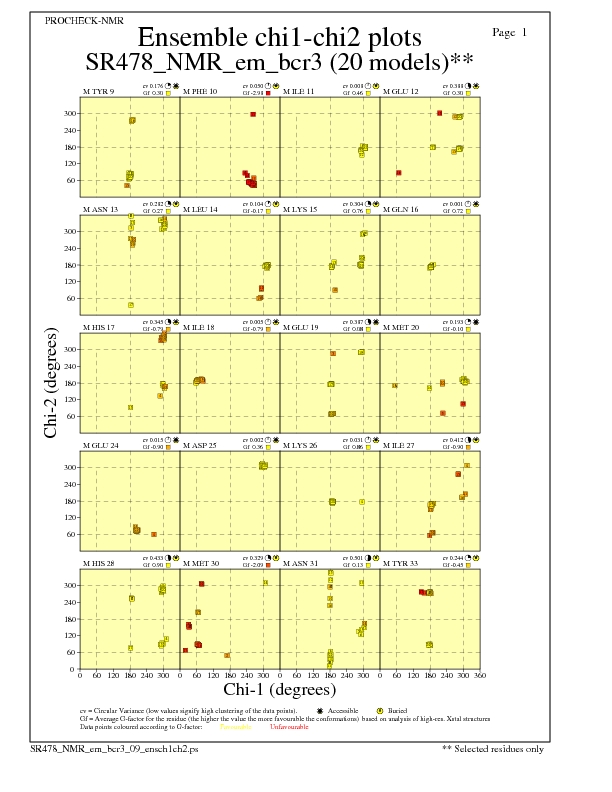
JPEG for residue Chi1-Chi2 Plots - page $num_n
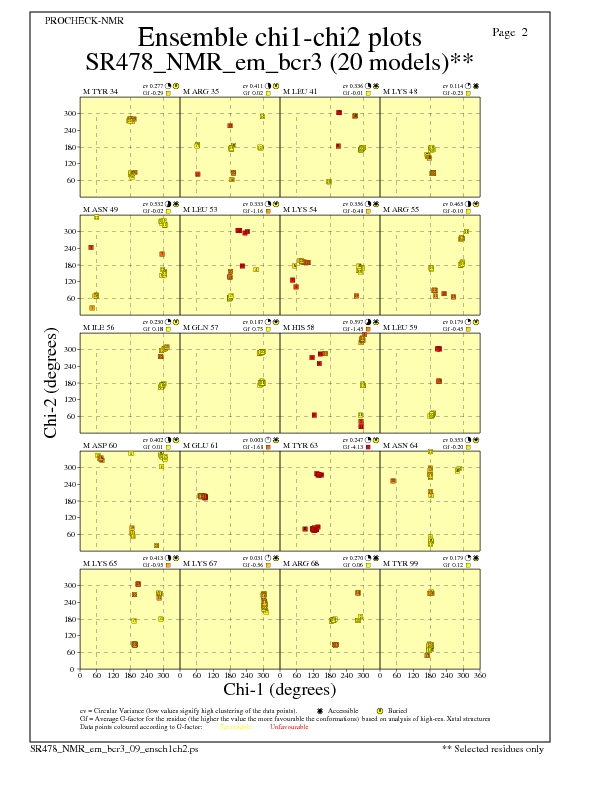
JPEG for residue Chi1-Chi2 Plots - page $num_n
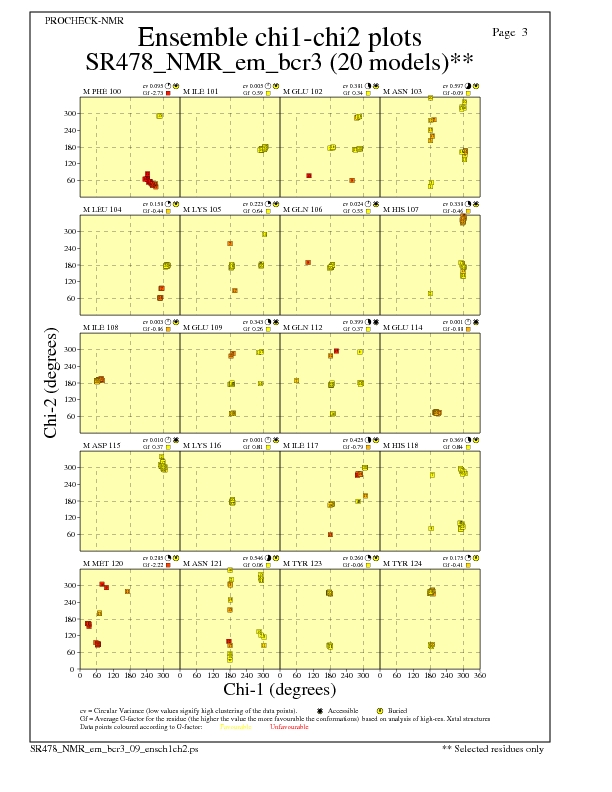
JPEG for residue Chi1-Chi2 Plots - page $num_n
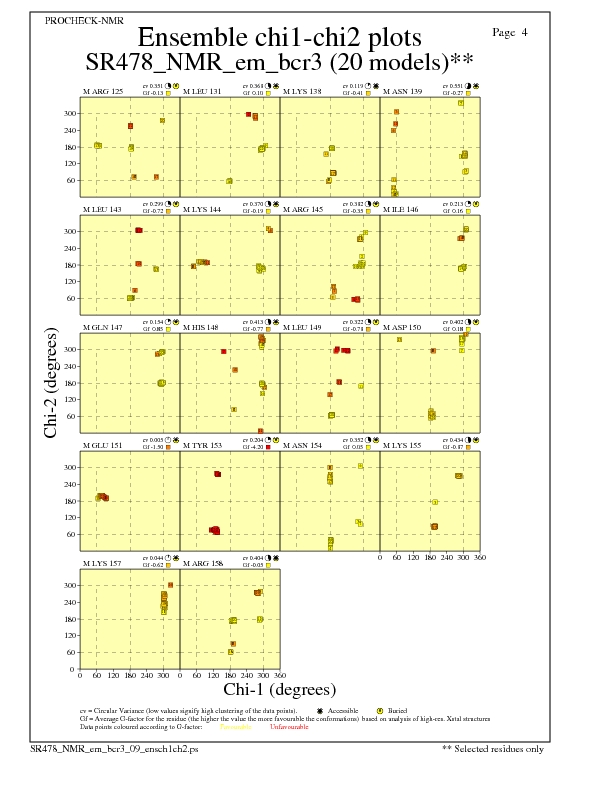
Procheck G-factors for phi-psi for each residue
JPEG image for residue phi-psi G-factors
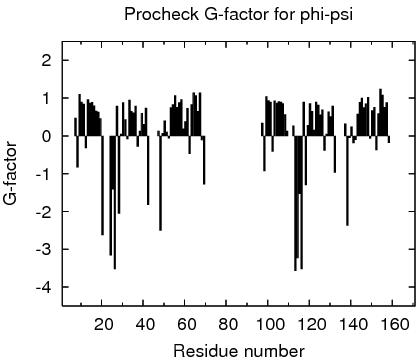
Table of Procheck G-factors for phi-psi for ordered residues
#phipsi_gfactor
#Residue\Model average
7 0.47
8 -0.83
9 1.10
10 0.90
11 0.84
12 -0.32
13 0.96
14 0.87
15 0.89
16 0.80
17 0.66
18 0.63
19 0.46
20 -2.62
24 -3.16
25 -1.41
26 -3.52
27 0.79
28 -2.05
29 0.05
30 0.88
31 0.43
32 -0.08
33 0.95
34 0.65
35 0.61
36 0.79
37 -0.28
38 0.13
39 0.60
40 0.31
41 0.74
42 -1.82
47 0.13
48 -2.50
49 0.07
50 0.40
51 0.10
52 -0.06
53 0.75
54 0.83
55 1.07
56 0.76
57 0.88
58 0.96
59 0.19
60 0.38
61 0.73
62 -0.47
63 0.83
64 1.14
65 1.07
66 0.65
67 1.14
68 -0.11
69 -1.28
97 0.34
98 -0.93
99 1.04
100 0.94
101 0.90
102 -0.41
103 0.93
104 0.87
105 0.91
106 0.90
107 0.86
108 0.57
109 0.13
112 0.27
113 -3.57
114 -3.23
115 -1.53
116 -3.52
117 0.90
118 -1.30
119 0.28
120 0.86
121 0.65
122 0.16
123 0.90
124 0.82
125 0.56
126 0.69
127 -0.39
128 0.05
129 0.64
130 0.51
131 0.79
132 -0.97
137 0.32
138 -2.37
139 -0.05
140 0.24
141 -0.19
142 -0.10
143 0.58
144 0.89
145 1.00
146 0.75
147 0.85
148 1.02
149 -0.07
150 0.67
151 0.76
152 -0.37
153 0.59
154 1.24
155 1.08
156 0.76
157 0.88
158 -0.18
#Reported_Model_Average 0.143
#Overall_Average_Reported 0.143
Procheck G-factors for all dihedral angles for each residue
JPEG image for residue all dihedral G-factors
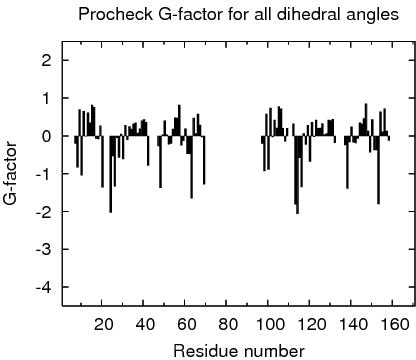
Table of Procheck G-factors for all dihedrals for ordered residues
#alldih_gfactor
#Residue\Model average
7 -0.20
8 -0.83
9 0.70
10 -1.04
11 0.65
12 -0.01
13 0.61
14 0.35
15 0.82
16 0.76
17 -0.07
18 -0.08
19 0.27
20 -1.36
24 -2.03
25 -0.53
26 -1.33
27 -0.05
28 -0.57
29 0.05
30 -0.61
31 0.28
32 -0.10
33 0.25
34 0.18
35 0.32
36 0.35
37 0.08
38 0.19
39 0.40
40 0.43
41 0.36
42 -0.78
47 -0.27
48 -1.37
49 0.03
50 0.40
51 0.03
52 -0.22
53 -0.20
54 0.18
55 0.48
56 0.47
57 0.82
58 -0.25
59 -0.13
60 0.19
61 -0.47
62 -0.47
63 -1.65
64 0.47
65 0.06
66 0.58
67 0.29
68 -0.03
69 -1.28
97 -0.20
98 -0.93
99 0.58
100 -0.89
101 0.74
102 -0.03
103 0.42
104 0.21
105 0.78
106 0.72
107 0.20
108 -0.14
109 0.20
112 0.32
113 -1.81
114 -2.06
115 -0.58
116 -1.35
117 0.06
118 -0.23
119 0.28
120 -0.68
121 0.36
122 -0.02
123 0.42
124 0.20
125 0.21
126 0.33
127 0.04
128 0.06
129 0.42
130 0.41
131 0.44
132 -0.18
137 -0.24
138 -1.39
139 -0.16
140 0.24
141 -0.17
142 -0.19
143 -0.07
144 0.35
145 0.32
146 0.46
147 0.85
148 0.13
149 -0.43
150 0.43
151 -0.37
152 -0.37
153 -1.80
154 0.64
155 0.11
156 0.72
157 0.13
158 -0.12
#Reported_Model_Average -0.067
#Overall_Average_Reported -0.067
Output from Verify3D
Verify3D Score over a window of $winsize_s residues
JPEG image for Verify3D Score
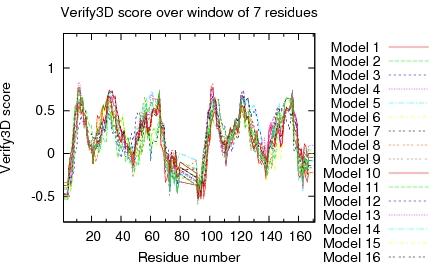
Table of Verify3D scores for ordered residues across all models
#verify3d
#Residue\Model 1 2 3 4 5 6 7 8 9 10 11 12 13 14 15 16 17 18 19 20
7 -0.74 -0.74 -0.74 -0.74 -0.74 -0.74 -0.74 -0.74 -0.74 -0.74 -0.09 -0.74 -0.74 -0.74 -0.74 -0.74 -0.74 -0.74 -0.74 -0.09
8 0.25 0.44 0.25 0.25 0.25 0.25 0.44 0.44 0.44 0.44 0.44 0.44 0.44 0.44 0.25 0.25 0.44 0.44 0.44 0.44
9 -0.43 -0.43 -0.43 -0.43 -0.43 -0.43 1.25 -0.43 -0.43 1.25 1.25 -0.43 1.25 1.25 -0.43 1.25 1.25 1.25 1.25 1.25
10 1.04 0.71 1.04 0.71 1.04 1.04 0.71 1.04 1.04 1.04 0.71 1.04 1.40 0.71 0.71 1.04 0.71 1.04 0.71 0.71
11 0.81 0.81 -0.54 0.81 0.93 0.93 0.93 0.81 0.81 0.81 0.93 0.93 0.81 0.81 0.93 0.81 0.81 0.81 0.81 0.93
12 0.28 0.28 0.28 0.28 0.28 0.28 0.28 0.28 0.28 0.28 0.28 0.28 0.28 0.28 0.28 0.28 0.28 0.28 -0.59 0.28
13 0.51 0.51 0.51 0.51 0.51 0.51 0.51 0.51 0.51 0.51 -0.26 0.51 0.51 -0.26 0.51 0.51 0.51 0.51 0.51 0.51
14 1.06 1.06 1.06 1.06 1.06 1.06 1.06 1.06 1.06 1.06 1.06 1.06 1.06 1.06 1.06 1.06 1.06 1.06 1.06 1.06
15 0.47 0.47 0.47 0.47 0.47 0.47 0.47 0.47 0.47 0.47 0.47 0.47 0.47 0.47 0.47 0.08 0.47 0.47 0.47 0.47
16 0.25 0.25 0.25 0.25 0.25 0.25 0.25 0.25 0.25 0.25 0.25 0.25 0.25 0.25 0.25 0.25 0.25 0.25 0.25 0.25
17 0.54 1.04 0.54 0.54 1.04 0.54 1.04 1.04 0.54 1.04 1.04 1.04 1.04 1.04 1.04 0.54 1.04 1.04 1.04 1.04
18 -0.54 -0.54 -0.54 -0.54 -0.54 -0.54 -0.54 -0.54 -0.54 -0.54 0.81 -0.54 -0.54 -0.54 -0.54 0.81 -0.54 -0.54 -0.54 -0.54
19 0.28 0.28 0.28 0.28 0.28 0.28 0.28 0.28 0.28 0.28 0.28 0.28 0.28 0.28 0.28 0.28 0.28 0.28 0.28 0.28
20 -0.83 -0.83 -0.83 -0.83 -0.83 -0.83 -0.83 -0.83 -0.83 -0.83 -0.83 -0.83 -0.83 -0.83 -0.83 -0.83 -0.83 -0.83 -0.83 -0.83
24 0.28 -0.46 0.28 0.28 0.28 0.28 -0.46 0.28 -0.46 -0.46 -0.46 -0.46 -0.46 -0.46 -0.59 0.28 -0.46 -0.46 -0.46 -0.46
25 0.51 0.51 0.51 0.51 0.51 0.51 0.51 0.51 0.51 0.51 0.51 0.51 0.51 0.51 0.51 0.51 0.51 0.51 0.51 0.51
26 0.47 0.47 0.47 0.47 0.47 0.47 0.47 0.47 0.47 0.47 0.47 0.47 0.47 0.47 0.47 0.47 -0.72 0.47 0.47 0.47
27 0.81 -0.94 -0.94 -0.28 -0.28 0.81 -0.94 -0.28 -0.28 0.81 -0.54 -0.54 -0.28 -0.54 0.81 -0.28 -0.28 -0.28 -0.28 0.81
28 0.20 0.20 0.20 0.20 0.20 0.20 1.04 1.04 0.20 0.20 -0.41 0.20 1.04 0.20 0.20 0.20 1.04 0.20 0.20 0.20
29 0.49 0.49 0.49 0.49 0.49 0.49 0.49 0.49 0.49 0.49 0.49 0.49 0.49 0.49 0.49 0.49 0.49 0.49 0.49 0.49
30 0.91 0.91 0.91 1.00 1.00 1.00 0.91 0.91 1.00 0.91 1.00 0.91 0.91 0.91 1.00 0.91 1.00 0.91 1.00 0.91
31 0.51 0.51 0.51 -0.26 -0.26 -0.26 -0.26 -0.26 -0.26 0.51 -0.26 0.09 -0.26 0.51 0.09 0.51 0.09 -0.26 0.51 0.51
32 0.17 0.17 0.17 0.34 0.17 0.34 0.17 0.17 0.34 0.17 0.17 0.17 0.17 0.17 0.17 0.17 0.17 0.17 0.17 0.34
33 1.14 0.52 1.14 1.14 1.14 1.14 1.14 1.14 1.14 1.14 0.52 1.14 1.14 1.14 1.14 1.14 0.52 1.14 1.14 1.14
34 1.25 1.14 1.14 1.14 1.14 1.14 1.14 1.14 1.14 1.14 1.14 0.52 1.14 1.14 1.14 1.14 1.14 1.14 1.14 0.52
35 0.71 0.71 0.71 0.71 0.24 -0.44 0.71 0.71 -0.44 0.24 0.24 -0.44 0.71 0.71 0.24 0.71 0.24 -0.44 0.24 -0.44
36 0.34 0.34 0.34 0.34 0.34 0.34 0.34 0.34 0.34 0.34 0.17 0.34 0.34 0.34 0.17 0.34 0.34 0.34 0.34 0.34
37 1.00 -0.09 -0.09 -0.09 -0.09 -0.09 -0.09 -0.09 -0.09 1.00 -0.09 -0.09 1.00 -0.09 -0.09 1.00 -0.09 -0.09 -0.09 -0.09
38 -0.09 -0.09 -0.09 -0.09 -0.09 -0.09 -0.09 -0.09 -0.09 -0.09 1.00 -0.09 -0.09 -0.09 1.00 -0.09 -0.09 -0.09 -0.09 -0.09
39 0.34 0.17 0.17 0.34 0.34 0.34 0.17 0.34 0.34 0.17 0.34 0.34 0.17 0.34 0.17 0.34 0.34 0.34 0.34 0.17
40 0.08 0.08 0.08 0.08 0.08 0.08 0.08 0.08 0.08 0.08 0.08 0.08 0.08 0.08 0.08 0.08 0.08 0.08 0.08 0.08
41 0.77 0.29 -0.33 0.29 0.77 0.77 0.29 1.06 0.77 0.77 0.29 0.77 0.77 0.77 0.77 0.77 0.77 0.77 0.77 -0.33
42 -0.09 -0.09 -0.09 -0.09 -0.09 0.66 1.00 -0.09 -0.09 -0.09 1.00 -0.09 -0.09 -0.09 0.66 -0.09 -0.09 -0.09 -0.09 0.66
47 0.08 0.08 0.08 0.08 0.08 0.08 0.08 0.08 0.08 0.08 0.08 0.08 0.08 0.08 0.08 0.08 0.08 0.08 0.08 0.08
48 0.47 -0.10 0.47 0.47 0.47 0.47 0.47 -0.10 0.47 0.47 0.47 0.47 0.47 0.47 -0.10 0.47 0.47 -0.10 0.47 0.47
49 -0.26 0.51 0.51 -0.26 -0.26 -0.26 -0.26 -0.26 -0.26 -0.26 -0.26 -0.26 -0.26 -0.26 -0.26 -0.26 -0.26 -0.26 -0.26 -0.26
50 -0.25 0.14 0.49 0.49 0.49 0.14 0.14 -0.25 -0.25 0.14 0.14 -0.25 0.14 0.14 0.49 0.49 0.14 -0.25 -0.25 0.14
51 -0.74 -0.09 -0.09 -0.74 -0.09 -0.09 -0.09 -0.09 -0.74 -0.09 -0.74 -0.09 -0.74 -0.74 -0.74 -0.09 -0.09 -0.74 -0.09 -0.74
52 0.66 -0.40 0.66 1.00 0.66 0.66 0.66 0.66 -0.09 0.66 0.66 0.66 0.66 0.66 -0.40 -0.09 -0.09 0.66 0.66 -0.40
53 1.06 0.77 0.77 0.77 0.77 0.77 -0.33 0.29 0.77 -0.33 -0.33 0.77 0.77 0.77 1.06 0.29 -0.68 1.06 0.77 -0.33
54 0.47 0.47 0.47 -0.10 -0.10 0.47 0.47 0.47 0.47 0.47 0.47 0.47 0.47 -0.10 0.47 0.47 0.47 0.47 0.47 0.47
55 0.24 0.71 0.24 0.71 0.71 0.71 0.71 0.24 0.24 0.71 -0.44 0.24 0.71 0.24 0.24 0.24 0.24 0.71 0.24 0.24
56 0.81 0.81 0.81 -0.28 0.81 0.81 0.93 0.93 0.81 0.81 0.93 0.93 0.93 0.81 0.93 0.93 0.81 0.81 0.81 0.81
57 -0.57 -0.57 -0.57 0.25 -0.57 -0.57 -0.57 0.25 -0.57 -0.57 -0.57 -0.87 -0.57 0.25 -0.57 -0.57 -0.57 0.25 -0.57 0.25
58 0.20 0.20 0.20 1.04 1.04 1.04 0.20 0.20 0.20 0.20 0.20 0.20 0.20 1.04 0.20 0.20 0.20 0.20 0.20 1.04
59 1.06 1.06 1.06 1.06 1.06 1.06 1.06 1.06 1.06 1.06 1.06 1.06 1.06 1.06 1.06 1.06 0.77 1.06 1.06 1.06
60 0.51 0.51 0.23 0.23 0.51 0.51 0.51 0.51 0.51 0.51 0.51 0.51 0.51 0.51 0.23 0.51 0.51 0.51 0.51 0.51
61 -0.59 -0.59 -0.59 0.28 -0.59 -0.59 -0.59 0.28 -0.59 -0.59 -0.59 -0.59 -0.59 0.28 -0.59 -0.59 -0.59 -0.59 -0.59 -0.59
62 0.49 0.49 0.49 0.49 0.49 0.49 0.49 0.49 0.49 0.49 0.49 0.49 0.49 0.49 0.49 0.49 0.49 0.49 0.49 0.49
63 1.25 -0.43 1.25 1.25 1.14 -1.40 1.14 1.25 1.25 1.14 -1.40 -1.40 1.25 1.25 1.25 1.14 1.14 1.25 1.25 1.25
64 0.51 0.51 0.51 0.51 0.51 0.51 0.51 0.51 0.51 0.51 0.51 0.51 0.51 0.51 0.51 0.51 0.51 0.51 0.51 0.51
65 0.47 0.47 0.47 0.47 0.47 0.47 0.08 0.47 0.47 0.47 0.47 0.47 0.47 0.47 0.47 0.47 0.47 0.47 0.47 0.47
66 1.00 1.00 1.00 1.00 1.00 1.00 1.00 1.00 1.00 1.00 1.00 1.00 1.00 1.00 1.00 1.00 1.00 1.00 1.00 1.00
67 0.08 0.08 0.08 0.47 0.47 0.08 0.47 -0.72 0.47 0.08 0.47 0.47 0.47 0.08 0.47 0.47 0.47 0.47 0.47 0.47
68 0.24 0.24 0.24 0.24 0.24 0.24 0.24 0.24 0.24 -0.41 0.24 0.24 0.24 0.24 0.24 0.24 0.24 0.24 -0.41 0.24
69 1.10 1.10 1.10 1.10 1.10 1.10 1.10 1.10 1.10 1.10 1.10 1.10 1.10 1.10 1.10 1.10 1.10 1.10 1.10 1.10
97 -0.74 -0.74 -0.74 -0.74 -0.74 -0.74 -0.74 -0.74 -0.74 -0.74 -0.74 -0.74 -0.74 -0.74 -0.74 -0.74 -0.74 -0.74 -0.74 -0.74
98 0.25 0.25 0.25 0.25 0.25 0.25 0.25 0.44 0.64 0.44 0.64 0.25 0.25 0.25 0.25 0.25 0.25 0.25 0.25 0.25
99 -1.40 -0.43 -0.43 1.25 -0.43 -0.43 -0.43 1.25 1.25 1.25 1.25 1.25 1.25 1.25 1.25 1.25 1.25 1.25 -0.43 -0.43
100 1.04 1.40 0.71 1.04 1.04 1.04 1.04 0.71 1.04 1.04 1.04 1.04 1.40 1.04 1.04 1.04 1.40 1.04 1.04 1.04
101 0.81 0.81 0.81 0.81 0.93 0.81 0.81 0.81 0.81 0.81 -0.54 0.81 0.81 0.81 -0.28 0.81 0.81 -0.28 0.81 0.81
102 0.28 0.28 0.28 0.28 0.04 0.28 0.28 0.28 0.28 0.28 0.28 0.28 0.28 0.28 0.28 0.28 0.28 0.28 0.28 0.28
103 0.51 -0.26 0.51 -0.26 0.51 -0.26 0.51 -0.26 -0.26 0.51 -0.26 0.51 -0.26 -0.26 0.51 -0.26 0.51 0.51 -0.26 0.51
104 1.06 1.06 1.06 1.06 1.06 1.06 1.06 1.06 1.06 1.06 1.06 1.06 1.06 1.06 1.06 1.06 1.06 1.06 1.06 1.06
105 0.47 0.47 0.47 0.47 0.47 0.47 0.47 0.47 0.47 0.47 0.47 0.47 0.47 0.08 0.47 0.47 0.47 0.47 0.47 0.47
106 0.25 0.25 0.25 0.25 0.25 0.25 0.25 0.25 0.25 0.25 0.25 0.25 0.25 0.25 0.25 0.25 0.25 0.25 -0.03 0.25
107 1.04 1.04 1.04 1.04 1.04 1.04 1.04 1.04 1.04 0.54 1.04 1.04 0.54 1.04 1.04 1.04 1.04 1.04 1.04 1.04
108 -0.54 0.81 0.81 0.81 -0.54 -0.54 -0.54 -0.54 -0.54 -0.54 -0.54 -0.54 -0.28 -0.54 -0.54 -0.54 0.81 -0.54 -0.54 0.81
109 0.28 -0.59 0.28 -0.59 0.28 0.28 0.28 0.04 0.28 0.28 0.28 0.28 0.04 0.28 0.28 0.28 0.28 -0.59 0.28 0.28
112 0.25 0.25 0.25 0.25 0.25 0.25 0.25 -0.03 0.25 -0.03 -0.03 -0.03 0.25 0.25 0.25 -0.03 -0.03 -0.03 0.25 -0.03
113 0.34 0.34 0.34 0.34 0.34 0.34 0.34 0.34 0.34 0.34 0.34 0.34 0.34 0.34 0.34 0.34 0.34 0.34 0.34 0.34
114 0.28 0.28 0.28 0.28 -0.46 -0.59 0.28 -0.46 0.28 -0.46 0.28 -0.46 -0.46 -0.46 -0.46 0.28 0.28 0.28 0.28 0.28
115 0.51 0.51 0.51 0.51 0.51 0.51 0.51 0.51 0.51 0.51 0.51 0.51 0.51 0.51 0.51 0.51 0.51 0.51 0.51 0.51
116 0.47 0.47 0.47 0.47 0.47 0.47 0.47 0.47 0.47 0.47 0.08 0.47 0.47 0.47 0.47 0.47 0.47 0.08 0.47 0.47
117 -0.28 -0.28 -0.28 -0.94 -0.28 -0.28 -0.94 -0.28 -0.28 -0.28 -0.28 -0.28 -0.54 -0.28 -0.54 -0.28 -0.28 -0.28 -0.28 -0.28
118 0.20 0.20 0.20 0.20 0.20 0.20 0.20 0.20 0.20 0.20 0.20 -0.41 1.04 0.20 0.20 0.20 0.20 0.20 0.20 0.20
119 0.49 0.49 0.49 0.49 0.49 0.49 0.49 0.49 0.49 0.49 0.49 0.49 0.49 0.49 0.49 0.49 0.49 0.49 0.49 0.49
120 0.91 0.91 1.00 1.00 0.91 1.00 1.00 1.00 1.00 1.00 0.91 1.00 0.91 0.91 0.91 1.00 0.91 1.00 0.91 1.00
121 0.51 -0.26 0.51 0.51 -0.26 -0.26 -0.26 -0.26 -0.26 -0.26 0.51 0.51 -0.26 0.51 -0.26 0.51 -0.26 0.51 -0.26 -0.26
122 0.34 0.34 0.17 0.17 0.34 0.17 0.17 0.34 0.34 0.17 0.17 0.17 0.17 0.17 0.17 0.17 0.17 0.17 0.17 0.34
123 1.14 1.14 0.52 1.14 1.14 1.14 1.14 1.14 1.14 1.14 0.52 1.14 1.14 1.14 1.14 1.14 1.14 1.14 1.14 1.14
124 1.14 1.14 1.14 1.14 1.14 1.14 1.14 1.14 1.14 1.14 1.14 1.14 1.25 1.14 0.52 1.14 1.14 1.14 1.14 1.14
125 0.24 -0.44 0.71 0.24 0.71 0.24 0.71 -0.44 0.71 0.24 0.71 0.71 -0.44 0.24 0.71 0.24 -0.44 0.71 0.24 -0.44
126 0.34 0.34 0.34 0.34 0.17 0.34 0.34 0.34 0.34 0.34 0.34 0.34 0.34 0.34 0.34 0.34 0.34 0.34 0.34 0.34
127 -0.09 -0.09 -0.09 -0.09 1.00 0.66 1.00 -0.09 1.00 -0.09 -0.09 -0.09 1.00 -0.09 -0.09 -0.09 1.00 -0.09 -0.09 -0.09
128 -0.09 -0.09 -0.09 -0.09 -0.09 -0.09 -0.09 -0.09 -0.09 -0.09 -0.09 1.00 -0.09 -0.09 -0.09 -0.09 -0.09 0.66 -0.09 1.00
129 0.34 0.34 0.34 0.34 0.17 0.34 0.34 0.17 0.34 0.34 0.34 0.34 0.17 0.17 0.34 0.34 0.34 0.34 0.34 0.34
130 0.08 0.08 0.08 0.08 0.55 0.08 0.08 0.08 0.08 0.08 0.08 0.08 0.08 0.08 0.08 0.08 0.08 0.08 0.08 0.08
131 0.77 0.29 0.77 0.77 1.06 0.29 0.77 0.29 -0.33 0.77 0.77 0.77 0.77 0.29 0.29 0.77 1.06 0.77 0.29 0.29
132 -0.74 -0.09 -0.74 -0.09 1.00 -0.74 -0.09 -0.09 -0.09 -0.09 -0.09 1.00 -0.09 -0.09 -0.09 -0.09 -0.09 -0.74 -0.09 -0.09
137 0.08 0.08 0.08 0.08 0.08 0.08 0.08 0.08 0.08 0.08 0.08 0.08 0.08 0.08 0.08 0.08 0.08 0.08 0.08 0.08
138 0.47 0.47 0.47 0.47 0.47 0.47 0.47 0.47 -0.10 -0.10 0.47 -0.10 0.47 -0.10 0.47 0.47 0.47 0.47 0.47 0.47
139 -0.26 -0.26 -0.26 0.51 -0.26 -0.26 -0.26 -0.26 -0.26 -0.26 -0.26 -0.26 -0.26 -0.26 -0.26 -0.26 -0.26 -0.26 -0.26 -0.26
140 0.49 -0.25 -0.25 0.14 0.14 -0.25 0.14 0.14 -0.25 0.14 0.14 0.14 0.14 0.14 -0.25 -0.25 -0.25 -0.25 -0.25 0.14
141 -0.74 -0.74 -0.74 -0.74 -0.74 -0.74 -0.74 -0.74 -0.74 -0.09 -0.74 -0.74 -0.09 -0.74 -0.74 -0.74 -0.74 -0.09 -0.74 -0.74
142 0.66 0.66 0.66 0.66 0.66 -0.40 0.66 0.66 0.66 0.66 -0.09 0.66 0.66 0.66 0.66 0.66 0.66 0.66 0.66 -0.40
143 1.06 0.29 0.29 0.77 -0.33 0.77 0.29 0.29 -0.33 0.77 0.77 0.77 1.06 0.29 0.77 0.29 -0.68 0.77 0.77 -0.33
144 0.47 0.47 0.47 0.47 0.47 0.47 0.47 0.47 0.47 0.47 0.47 0.47 0.47 0.47 -0.10 0.47 0.47 0.47 0.47 0.47
145 0.24 0.24 0.24 0.71 0.71 -0.44 0.24 0.24 0.24 0.71 0.71 0.24 -0.88 0.71 0.24 0.24 -0.44 0.24 0.24 0.24
146 -0.28 0.81 0.81 0.93 0.81 -0.54 0.93 0.81 0.81 -0.54 0.81 0.81 0.93 0.81 0.81 -0.28 0.81 0.81 -0.28 0.81
147 -0.57 0.25 -0.57 -0.57 0.25 -0.57 0.25 0.25 -0.57 0.25 -0.57 0.25 -0.57 -0.57 0.25 0.25 0.25 0.25 -0.57 -0.57
148 0.20 0.20 0.20 0.20 0.20 0.20 0.20 0.20 0.20 0.20 0.20 0.20 0.20 0.20 0.20 0.20 0.20 0.20 0.20 0.20
149 1.06 1.06 1.06 1.06 1.06 1.06 1.06 1.06 1.06 1.06 1.06 1.06 1.06 1.06 1.06 1.06 1.06 1.06 1.06 1.06
150 0.51 0.51 0.23 0.51 0.51 0.51 0.51 0.51 0.51 0.51 0.51 0.51 0.51 0.51 0.51 0.51 0.51 0.51 0.51 0.51
151 -0.59 -0.59 -0.59 0.28 -0.59 -0.59 0.28 -0.59 0.28 -0.59 0.28 0.28 -0.59 -0.59 0.28 -0.59 -0.59 -0.59 -0.59 0.28
152 0.49 0.49 0.49 0.49 0.49 0.49 0.49 0.49 0.49 0.49 0.49 0.49 0.49 0.49 0.49 0.49 0.49 0.49 0.49 0.49
153 1.25 1.25 1.25 1.25 1.25 1.25 1.25 1.25 1.25 1.14 1.25 1.25 1.25 1.25 1.25 1.25 1.25 1.25 1.25 1.25
154 0.51 0.51 0.51 0.51 0.51 0.51 -0.26 0.51 0.51 0.51 0.51 0.51 0.51 0.51 0.51 0.51 0.51 0.51 0.51 0.51
155 0.47 0.47 0.47 0.47 0.47 0.47 0.47 0.47 0.47 0.47 0.47 0.47 0.47 0.47 0.47 -0.72 0.47 0.47 0.47 0.47
156 1.00 1.00 1.00 1.00 1.00 1.00 1.00 1.00 1.00 1.00 1.00 1.00 1.00 1.00 1.00 1.00 1.00 1.00 1.00 1.00
157 0.47 0.47 0.47 0.47 0.08 0.47 0.47 0.47 0.47 0.47 0.47 0.47 0.47 0.47 0.47 0.08 0.47 0.47 0.47 0.47
158 -0.41 0.24 0.24 0.24 -0.41 0.24 -0.41 -0.41 0.24 0.24 -0.41 0.24 0.24 0.24 -0.41 0.24 0.24 -0.41 -0.41 -0.41
#Reported_Model_Average 0.340 0.294 0.320 0.378 0.363 0.285 0.356 0.331 0.317 0.361 0.314 0.337 0.380 0.350 0.332 0.360 0.337 0.351 0.304 0.339
#Overall_Average_Reported 0.337
Output from ProsaII
ProsaII Score over a window of $winsize_s residues
JPEG image for ProsaII Score
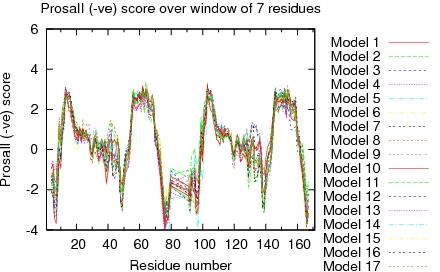
Table of Verify3D scores for ordered residues across all models
#verify3d
#Residue\Model 1 2 3 4 5 6 7 8 9 10 11 12 13 14 15 16 17 18 19 20
7 -0.74 -0.74 -0.74 -0.74 -0.74 -0.74 -0.74 -0.74 -0.74 -0.74 -0.09 -0.74 -0.74 -0.74 -0.74 -0.74 -0.74 -0.74 -0.74 -0.09
8 0.25 0.44 0.25 0.25 0.25 0.25 0.44 0.44 0.44 0.44 0.44 0.44 0.44 0.44 0.25 0.25 0.44 0.44 0.44 0.44
9 -0.43 -0.43 -0.43 -0.43 -0.43 -0.43 1.25 -0.43 -0.43 1.25 1.25 -0.43 1.25 1.25 -0.43 1.25 1.25 1.25 1.25 1.25
10 1.04 0.71 1.04 0.71 1.04 1.04 0.71 1.04 1.04 1.04 0.71 1.04 1.40 0.71 0.71 1.04 0.71 1.04 0.71 0.71
11 0.81 0.81 -0.54 0.81 0.93 0.93 0.93 0.81 0.81 0.81 0.93 0.93 0.81 0.81 0.93 0.81 0.81 0.81 0.81 0.93
12 0.28 0.28 0.28 0.28 0.28 0.28 0.28 0.28 0.28 0.28 0.28 0.28 0.28 0.28 0.28 0.28 0.28 0.28 -0.59 0.28
13 0.51 0.51 0.51 0.51 0.51 0.51 0.51 0.51 0.51 0.51 -0.26 0.51 0.51 -0.26 0.51 0.51 0.51 0.51 0.51 0.51
14 1.06 1.06 1.06 1.06 1.06 1.06 1.06 1.06 1.06 1.06 1.06 1.06 1.06 1.06 1.06 1.06 1.06 1.06 1.06 1.06
15 0.47 0.47 0.47 0.47 0.47 0.47 0.47 0.47 0.47 0.47 0.47 0.47 0.47 0.47 0.47 0.08 0.47 0.47 0.47 0.47
16 0.25 0.25 0.25 0.25 0.25 0.25 0.25 0.25 0.25 0.25 0.25 0.25 0.25 0.25 0.25 0.25 0.25 0.25 0.25 0.25
17 0.54 1.04 0.54 0.54 1.04 0.54 1.04 1.04 0.54 1.04 1.04 1.04 1.04 1.04 1.04 0.54 1.04 1.04 1.04 1.04
18 -0.54 -0.54 -0.54 -0.54 -0.54 -0.54 -0.54 -0.54 -0.54 -0.54 0.81 -0.54 -0.54 -0.54 -0.54 0.81 -0.54 -0.54 -0.54 -0.54
19 0.28 0.28 0.28 0.28 0.28 0.28 0.28 0.28 0.28 0.28 0.28 0.28 0.28 0.28 0.28 0.28 0.28 0.28 0.28 0.28
20 -0.83 -0.83 -0.83 -0.83 -0.83 -0.83 -0.83 -0.83 -0.83 -0.83 -0.83 -0.83 -0.83 -0.83 -0.83 -0.83 -0.83 -0.83 -0.83 -0.83
24 0.28 -0.46 0.28 0.28 0.28 0.28 -0.46 0.28 -0.46 -0.46 -0.46 -0.46 -0.46 -0.46 -0.59 0.28 -0.46 -0.46 -0.46 -0.46
25 0.51 0.51 0.51 0.51 0.51 0.51 0.51 0.51 0.51 0.51 0.51 0.51 0.51 0.51 0.51 0.51 0.51 0.51 0.51 0.51
26 0.47 0.47 0.47 0.47 0.47 0.47 0.47 0.47 0.47 0.47 0.47 0.47 0.47 0.47 0.47 0.47 -0.72 0.47 0.47 0.47
27 0.81 -0.94 -0.94 -0.28 -0.28 0.81 -0.94 -0.28 -0.28 0.81 -0.54 -0.54 -0.28 -0.54 0.81 -0.28 -0.28 -0.28 -0.28 0.81
28 0.20 0.20 0.20 0.20 0.20 0.20 1.04 1.04 0.20 0.20 -0.41 0.20 1.04 0.20 0.20 0.20 1.04 0.20 0.20 0.20
29 0.49 0.49 0.49 0.49 0.49 0.49 0.49 0.49 0.49 0.49 0.49 0.49 0.49 0.49 0.49 0.49 0.49 0.49 0.49 0.49
30 0.91 0.91 0.91 1.00 1.00 1.00 0.91 0.91 1.00 0.91 1.00 0.91 0.91 0.91 1.00 0.91 1.00 0.91 1.00 0.91
31 0.51 0.51 0.51 -0.26 -0.26 -0.26 -0.26 -0.26 -0.26 0.51 -0.26 0.09 -0.26 0.51 0.09 0.51 0.09 -0.26 0.51 0.51
32 0.17 0.17 0.17 0.34 0.17 0.34 0.17 0.17 0.34 0.17 0.17 0.17 0.17 0.17 0.17 0.17 0.17 0.17 0.17 0.34
33 1.14 0.52 1.14 1.14 1.14 1.14 1.14 1.14 1.14 1.14 0.52 1.14 1.14 1.14 1.14 1.14 0.52 1.14 1.14 1.14
34 1.25 1.14 1.14 1.14 1.14 1.14 1.14 1.14 1.14 1.14 1.14 0.52 1.14 1.14 1.14 1.14 1.14 1.14 1.14 0.52
35 0.71 0.71 0.71 0.71 0.24 -0.44 0.71 0.71 -0.44 0.24 0.24 -0.44 0.71 0.71 0.24 0.71 0.24 -0.44 0.24 -0.44
36 0.34 0.34 0.34 0.34 0.34 0.34 0.34 0.34 0.34 0.34 0.17 0.34 0.34 0.34 0.17 0.34 0.34 0.34 0.34 0.34
37 1.00 -0.09 -0.09 -0.09 -0.09 -0.09 -0.09 -0.09 -0.09 1.00 -0.09 -0.09 1.00 -0.09 -0.09 1.00 -0.09 -0.09 -0.09 -0.09
38 -0.09 -0.09 -0.09 -0.09 -0.09 -0.09 -0.09 -0.09 -0.09 -0.09 1.00 -0.09 -0.09 -0.09 1.00 -0.09 -0.09 -0.09 -0.09 -0.09
39 0.34 0.17 0.17 0.34 0.34 0.34 0.17 0.34 0.34 0.17 0.34 0.34 0.17 0.34 0.17 0.34 0.34 0.34 0.34 0.17
40 0.08 0.08 0.08 0.08 0.08 0.08 0.08 0.08 0.08 0.08 0.08 0.08 0.08 0.08 0.08 0.08 0.08 0.08 0.08 0.08
41 0.77 0.29 -0.33 0.29 0.77 0.77 0.29 1.06 0.77 0.77 0.29 0.77 0.77 0.77 0.77 0.77 0.77 0.77 0.77 -0.33
42 -0.09 -0.09 -0.09 -0.09 -0.09 0.66 1.00 -0.09 -0.09 -0.09 1.00 -0.09 -0.09 -0.09 0.66 -0.09 -0.09 -0.09 -0.09 0.66
47 0.08 0.08 0.08 0.08 0.08 0.08 0.08 0.08 0.08 0.08 0.08 0.08 0.08 0.08 0.08 0.08 0.08 0.08 0.08 0.08
48 0.47 -0.10 0.47 0.47 0.47 0.47 0.47 -0.10 0.47 0.47 0.47 0.47 0.47 0.47 -0.10 0.47 0.47 -0.10 0.47 0.47
49 -0.26 0.51 0.51 -0.26 -0.26 -0.26 -0.26 -0.26 -0.26 -0.26 -0.26 -0.26 -0.26 -0.26 -0.26 -0.26 -0.26 -0.26 -0.26 -0.26
50 -0.25 0.14 0.49 0.49 0.49 0.14 0.14 -0.25 -0.25 0.14 0.14 -0.25 0.14 0.14 0.49 0.49 0.14 -0.25 -0.25 0.14
51 -0.74 -0.09 -0.09 -0.74 -0.09 -0.09 -0.09 -0.09 -0.74 -0.09 -0.74 -0.09 -0.74 -0.74 -0.74 -0.09 -0.09 -0.74 -0.09 -0.74
52 0.66 -0.40 0.66 1.00 0.66 0.66 0.66 0.66 -0.09 0.66 0.66 0.66 0.66 0.66 -0.40 -0.09 -0.09 0.66 0.66 -0.40
53 1.06 0.77 0.77 0.77 0.77 0.77 -0.33 0.29 0.77 -0.33 -0.33 0.77 0.77 0.77 1.06 0.29 -0.68 1.06 0.77 -0.33
54 0.47 0.47 0.47 -0.10 -0.10 0.47 0.47 0.47 0.47 0.47 0.47 0.47 0.47 -0.10 0.47 0.47 0.47 0.47 0.47 0.47
55 0.24 0.71 0.24 0.71 0.71 0.71 0.71 0.24 0.24 0.71 -0.44 0.24 0.71 0.24 0.24 0.24 0.24 0.71 0.24 0.24
56 0.81 0.81 0.81 -0.28 0.81 0.81 0.93 0.93 0.81 0.81 0.93 0.93 0.93 0.81 0.93 0.93 0.81 0.81 0.81 0.81
57 -0.57 -0.57 -0.57 0.25 -0.57 -0.57 -0.57 0.25 -0.57 -0.57 -0.57 -0.87 -0.57 0.25 -0.57 -0.57 -0.57 0.25 -0.57 0.25
58 0.20 0.20 0.20 1.04 1.04 1.04 0.20 0.20 0.20 0.20 0.20 0.20 0.20 1.04 0.20 0.20 0.20 0.20 0.20 1.04
59 1.06 1.06 1.06 1.06 1.06 1.06 1.06 1.06 1.06 1.06 1.06 1.06 1.06 1.06 1.06 1.06 0.77 1.06 1.06 1.06
60 0.51 0.51 0.23 0.23 0.51 0.51 0.51 0.51 0.51 0.51 0.51 0.51 0.51 0.51 0.23 0.51 0.51 0.51 0.51 0.51
61 -0.59 -0.59 -0.59 0.28 -0.59 -0.59 -0.59 0.28 -0.59 -0.59 -0.59 -0.59 -0.59 0.28 -0.59 -0.59 -0.59 -0.59 -0.59 -0.59
62 0.49 0.49 0.49 0.49 0.49 0.49 0.49 0.49 0.49 0.49 0.49 0.49 0.49 0.49 0.49 0.49 0.49 0.49 0.49 0.49
63 1.25 -0.43 1.25 1.25 1.14 -1.40 1.14 1.25 1.25 1.14 -1.40 -1.40 1.25 1.25 1.25 1.14 1.14 1.25 1.25 1.25
64 0.51 0.51 0.51 0.51 0.51 0.51 0.51 0.51 0.51 0.51 0.51 0.51 0.51 0.51 0.51 0.51 0.51 0.51 0.51 0.51
65 0.47 0.47 0.47 0.47 0.47 0.47 0.08 0.47 0.47 0.47 0.47 0.47 0.47 0.47 0.47 0.47 0.47 0.47 0.47 0.47
66 1.00 1.00 1.00 1.00 1.00 1.00 1.00 1.00 1.00 1.00 1.00 1.00 1.00 1.00 1.00 1.00 1.00 1.00 1.00 1.00
67 0.08 0.08 0.08 0.47 0.47 0.08 0.47 -0.72 0.47 0.08 0.47 0.47 0.47 0.08 0.47 0.47 0.47 0.47 0.47 0.47
68 0.24 0.24 0.24 0.24 0.24 0.24 0.24 0.24 0.24 -0.41 0.24 0.24 0.24 0.24 0.24 0.24 0.24 0.24 -0.41 0.24
69 1.10 1.10 1.10 1.10 1.10 1.10 1.10 1.10 1.10 1.10 1.10 1.10 1.10 1.10 1.10 1.10 1.10 1.10 1.10 1.10
97 -0.74 -0.74 -0.74 -0.74 -0.74 -0.74 -0.74 -0.74 -0.74 -0.74 -0.74 -0.74 -0.74 -0.74 -0.74 -0.74 -0.74 -0.74 -0.74 -0.74
98 0.25 0.25 0.25 0.25 0.25 0.25 0.25 0.44 0.64 0.44 0.64 0.25 0.25 0.25 0.25 0.25 0.25 0.25 0.25 0.25
99 -1.40 -0.43 -0.43 1.25 -0.43 -0.43 -0.43 1.25 1.25 1.25 1.25 1.25 1.25 1.25 1.25 1.25 1.25 1.25 -0.43 -0.43
100 1.04 1.40 0.71 1.04 1.04 1.04 1.04 0.71 1.04 1.04 1.04 1.04 1.40 1.04 1.04 1.04 1.40 1.04 1.04 1.04
101 0.81 0.81 0.81 0.81 0.93 0.81 0.81 0.81 0.81 0.81 -0.54 0.81 0.81 0.81 -0.28 0.81 0.81 -0.28 0.81 0.81
102 0.28 0.28 0.28 0.28 0.04 0.28 0.28 0.28 0.28 0.28 0.28 0.28 0.28 0.28 0.28 0.28 0.28 0.28 0.28 0.28
103 0.51 -0.26 0.51 -0.26 0.51 -0.26 0.51 -0.26 -0.26 0.51 -0.26 0.51 -0.26 -0.26 0.51 -0.26 0.51 0.51 -0.26 0.51
104 1.06 1.06 1.06 1.06 1.06 1.06 1.06 1.06 1.06 1.06 1.06 1.06 1.06 1.06 1.06 1.06 1.06 1.06 1.06 1.06
105 0.47 0.47 0.47 0.47 0.47 0.47 0.47 0.47 0.47 0.47 0.47 0.47 0.47 0.08 0.47 0.47 0.47 0.47 0.47 0.47
106 0.25 0.25 0.25 0.25 0.25 0.25 0.25 0.25 0.25 0.25 0.25 0.25 0.25 0.25 0.25 0.25 0.25 0.25 -0.03 0.25
107 1.04 1.04 1.04 1.04 1.04 1.04 1.04 1.04 1.04 0.54 1.04 1.04 0.54 1.04 1.04 1.04 1.04 1.04 1.04 1.04
108 -0.54 0.81 0.81 0.81 -0.54 -0.54 -0.54 -0.54 -0.54 -0.54 -0.54 -0.54 -0.28 -0.54 -0.54 -0.54 0.81 -0.54 -0.54 0.81
109 0.28 -0.59 0.28 -0.59 0.28 0.28 0.28 0.04 0.28 0.28 0.28 0.28 0.04 0.28 0.28 0.28 0.28 -0.59 0.28 0.28
112 0.25 0.25 0.25 0.25 0.25 0.25 0.25 -0.03 0.25 -0.03 -0.03 -0.03 0.25 0.25 0.25 -0.03 -0.03 -0.03 0.25 -0.03
113 0.34 0.34 0.34 0.34 0.34 0.34 0.34 0.34 0.34 0.34 0.34 0.34 0.34 0.34 0.34 0.34 0.34 0.34 0.34 0.34
114 0.28 0.28 0.28 0.28 -0.46 -0.59 0.28 -0.46 0.28 -0.46 0.28 -0.46 -0.46 -0.46 -0.46 0.28 0.28 0.28 0.28 0.28
115 0.51 0.51 0.51 0.51 0.51 0.51 0.51 0.51 0.51 0.51 0.51 0.51 0.51 0.51 0.51 0.51 0.51 0.51 0.51 0.51
116 0.47 0.47 0.47 0.47 0.47 0.47 0.47 0.47 0.47 0.47 0.08 0.47 0.47 0.47 0.47 0.47 0.47 0.08 0.47 0.47
117 -0.28 -0.28 -0.28 -0.94 -0.28 -0.28 -0.94 -0.28 -0.28 -0.28 -0.28 -0.28 -0.54 -0.28 -0.54 -0.28 -0.28 -0.28 -0.28 -0.28
118 0.20 0.20 0.20 0.20 0.20 0.20 0.20 0.20 0.20 0.20 0.20 -0.41 1.04 0.20 0.20 0.20 0.20 0.20 0.20 0.20
119 0.49 0.49 0.49 0.49 0.49 0.49 0.49 0.49 0.49 0.49 0.49 0.49 0.49 0.49 0.49 0.49 0.49 0.49 0.49 0.49
120 0.91 0.91 1.00 1.00 0.91 1.00 1.00 1.00 1.00 1.00 0.91 1.00 0.91 0.91 0.91 1.00 0.91 1.00 0.91 1.00
121 0.51 -0.26 0.51 0.51 -0.26 -0.26 -0.26 -0.26 -0.26 -0.26 0.51 0.51 -0.26 0.51 -0.26 0.51 -0.26 0.51 -0.26 -0.26
122 0.34 0.34 0.17 0.17 0.34 0.17 0.17 0.34 0.34 0.17 0.17 0.17 0.17 0.17 0.17 0.17 0.17 0.17 0.17 0.34
123 1.14 1.14 0.52 1.14 1.14 1.14 1.14 1.14 1.14 1.14 0.52 1.14 1.14 1.14 1.14 1.14 1.14 1.14 1.14 1.14
124 1.14 1.14 1.14 1.14 1.14 1.14 1.14 1.14 1.14 1.14 1.14 1.14 1.25 1.14 0.52 1.14 1.14 1.14 1.14 1.14
125 0.24 -0.44 0.71 0.24 0.71 0.24 0.71 -0.44 0.71 0.24 0.71 0.71 -0.44 0.24 0.71 0.24 -0.44 0.71 0.24 -0.44
126 0.34 0.34 0.34 0.34 0.17 0.34 0.34 0.34 0.34 0.34 0.34 0.34 0.34 0.34 0.34 0.34 0.34 0.34 0.34 0.34
127 -0.09 -0.09 -0.09 -0.09 1.00 0.66 1.00 -0.09 1.00 -0.09 -0.09 -0.09 1.00 -0.09 -0.09 -0.09 1.00 -0.09 -0.09 -0.09
128 -0.09 -0.09 -0.09 -0.09 -0.09 -0.09 -0.09 -0.09 -0.09 -0.09 -0.09 1.00 -0.09 -0.09 -0.09 -0.09 -0.09 0.66 -0.09 1.00
129 0.34 0.34 0.34 0.34 0.17 0.34 0.34 0.17 0.34 0.34 0.34 0.34 0.17 0.17 0.34 0.34 0.34 0.34 0.34 0.34
130 0.08 0.08 0.08 0.08 0.55 0.08 0.08 0.08 0.08 0.08 0.08 0.08 0.08 0.08 0.08 0.08 0.08 0.08 0.08 0.08
131 0.77 0.29 0.77 0.77 1.06 0.29 0.77 0.29 -0.33 0.77 0.77 0.77 0.77 0.29 0.29 0.77 1.06 0.77 0.29 0.29
132 -0.74 -0.09 -0.74 -0.09 1.00 -0.74 -0.09 -0.09 -0.09 -0.09 -0.09 1.00 -0.09 -0.09 -0.09 -0.09 -0.09 -0.74 -0.09 -0.09
137 0.08 0.08 0.08 0.08 0.08 0.08 0.08 0.08 0.08 0.08 0.08 0.08 0.08 0.08 0.08 0.08 0.08 0.08 0.08 0.08
138 0.47 0.47 0.47 0.47 0.47 0.47 0.47 0.47 -0.10 -0.10 0.47 -0.10 0.47 -0.10 0.47 0.47 0.47 0.47 0.47 0.47
139 -0.26 -0.26 -0.26 0.51 -0.26 -0.26 -0.26 -0.26 -0.26 -0.26 -0.26 -0.26 -0.26 -0.26 -0.26 -0.26 -0.26 -0.26 -0.26 -0.26
140 0.49 -0.25 -0.25 0.14 0.14 -0.25 0.14 0.14 -0.25 0.14 0.14 0.14 0.14 0.14 -0.25 -0.25 -0.25 -0.25 -0.25 0.14
141 -0.74 -0.74 -0.74 -0.74 -0.74 -0.74 -0.74 -0.74 -0.74 -0.09 -0.74 -0.74 -0.09 -0.74 -0.74 -0.74 -0.74 -0.09 -0.74 -0.74
142 0.66 0.66 0.66 0.66 0.66 -0.40 0.66 0.66 0.66 0.66 -0.09 0.66 0.66 0.66 0.66 0.66 0.66 0.66 0.66 -0.40
143 1.06 0.29 0.29 0.77 -0.33 0.77 0.29 0.29 -0.33 0.77 0.77 0.77 1.06 0.29 0.77 0.29 -0.68 0.77 0.77 -0.33
144 0.47 0.47 0.47 0.47 0.47 0.47 0.47 0.47 0.47 0.47 0.47 0.47 0.47 0.47 -0.10 0.47 0.47 0.47 0.47 0.47
145 0.24 0.24 0.24 0.71 0.71 -0.44 0.24 0.24 0.24 0.71 0.71 0.24 -0.88 0.71 0.24 0.24 -0.44 0.24 0.24 0.24
146 -0.28 0.81 0.81 0.93 0.81 -0.54 0.93 0.81 0.81 -0.54 0.81 0.81 0.93 0.81 0.81 -0.28 0.81 0.81 -0.28 0.81
147 -0.57 0.25 -0.57 -0.57 0.25 -0.57 0.25 0.25 -0.57 0.25 -0.57 0.25 -0.57 -0.57 0.25 0.25 0.25 0.25 -0.57 -0.57
148 0.20 0.20 0.20 0.20 0.20 0.20 0.20 0.20 0.20 0.20 0.20 0.20 0.20 0.20 0.20 0.20 0.20 0.20 0.20 0.20
149 1.06 1.06 1.06 1.06 1.06 1.06 1.06 1.06 1.06 1.06 1.06 1.06 1.06 1.06 1.06 1.06 1.06 1.06 1.06 1.06
150 0.51 0.51 0.23 0.51 0.51 0.51 0.51 0.51 0.51 0.51 0.51 0.51 0.51 0.51 0.51 0.51 0.51 0.51 0.51 0.51
151 -0.59 -0.59 -0.59 0.28 -0.59 -0.59 0.28 -0.59 0.28 -0.59 0.28 0.28 -0.59 -0.59 0.28 -0.59 -0.59 -0.59 -0.59 0.28
152 0.49 0.49 0.49 0.49 0.49 0.49 0.49 0.49 0.49 0.49 0.49 0.49 0.49 0.49 0.49 0.49 0.49 0.49 0.49 0.49
153 1.25 1.25 1.25 1.25 1.25 1.25 1.25 1.25 1.25 1.14 1.25 1.25 1.25 1.25 1.25 1.25 1.25 1.25 1.25 1.25
154 0.51 0.51 0.51 0.51 0.51 0.51 -0.26 0.51 0.51 0.51 0.51 0.51 0.51 0.51 0.51 0.51 0.51 0.51 0.51 0.51
155 0.47 0.47 0.47 0.47 0.47 0.47 0.47 0.47 0.47 0.47 0.47 0.47 0.47 0.47 0.47 -0.72 0.47 0.47 0.47 0.47
156 1.00 1.00 1.00 1.00 1.00 1.00 1.00 1.00 1.00 1.00 1.00 1.00 1.00 1.00 1.00 1.00 1.00 1.00 1.00 1.00
157 0.47 0.47 0.47 0.47 0.08 0.47 0.47 0.47 0.47 0.47 0.47 0.47 0.47 0.47 0.47 0.08 0.47 0.47 0.47 0.47
158 -0.41 0.24 0.24 0.24 -0.41 0.24 -0.41 -0.41 0.24 0.24 -0.41 0.24 0.24 0.24 -0.41 0.24 0.24 -0.41 -0.41 -0.41
#Reported_Model_Average 0.340 0.294 0.320 0.378 0.363 0.285 0.356 0.331 0.317 0.361 0.314 0.337 0.380 0.350 0.332 0.360 0.337 0.351 0.304 0.339
#Overall_Average_Reported 0.337
Output from MolProbity
VdW violations from MAGE
JPEG image for MAGE VdW violation
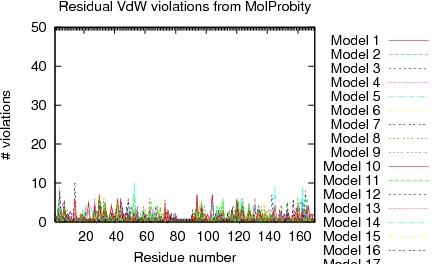
Table of MAGE VdW violations for ordered residues across all models
#mage_clash
#Residue\Model 1 2 3 4 5 6 7 8 9 10 11 12 13 14 15 16 17 18 19 20
7.000 2 1 1 5 2 3 5 1 1 2 3 1 2 2 2 3 3 2 4 4
8.000 2 1 1 2 2 2 2 1 1 2 1 1 2 2 1 1 1 2 1 2
9.000 0 0 0 0 0 1 1 3 0 0 0 2 2 0 0 0 0 0 2 2
10.000 1 1 0 0 1 1 2 0 1 1 1 1 0 1 1 2 0 1 1 0
11.000 0 0 0 0 0 0 0 0 0 0 0 3 0 0 1 0 1 0 0 0
12.000 0 0 0 0 0 0 0 0 0 0 0 0 1 0 0 2 0 0 0 0
13.000 0 0 0 0 0 0 0 0 0 0 0 0 2 0 0 0 0 0 0 0
14.000 1 0 0 0 2 0 1 1 0 0 0 1 1 0 1 10 2 2 6 0
15.000 0 0 1 1 1 1 0 0 0 1 0 1 0 1 0 0 1 0 0 0
16.000 0 0 0 0 1 1 1 0 1 1 0 0 0 0 0 0 0 0 0 0
17.000 0 1 0 0 0 0 0 1 0 0 0 0 2 1 2 1 0 0 0 0
18.000 1 0 0 2 2 1 1 0 0 1 2 1 0 0 3 1 1 1 2 2
19.000 1 0 1 2 3 2 1 0 0 2 2 2 0 1 1 1 2 1 1 2
20.000 0 1 0 0 1 1 1 1 0 1 0 0 0 1 0 1 0 1 0 0
24.000 1 0 1 0 0 0 0 1 0 0 0 0 0 2 1 0 1 0 1 0
25.000 0 1 0 0 0 0 0 1 0 0 0 0 0 4 1 0 0 0 0 0
26.000 0 0 0 1 1 0 0 0 1 1 1 0 0 1 0 1 0 0 1 2
27.000 2 4 4 0 3 4 2 3 1 3 3 4 6 4 2 0 2 5 4 2
28.000 0 0 3 0 0 0 2 1 0 0 0 0 2 2 0 0 2 2 1 1
29.000 2 2 2 0 2 2 2 1 1 2 2 2 3 3 3 0 2 2 1 1
30.000 4 1 2 4 6 7 4 2 2 4 4 4 4 3 4 6 3 4 7 6
31.000 2 1 1 1 4 5 4 1 0 2 3 5 1 0 4 2 2 1 2 4
32.000 1 0 1 0 0 0 0 0 0 0 0 0 0 0 0 0 1 0 1 0
33.000 2 5 2 0 4 1 2 1 1 1 3 1 1 3 2 3 6 3 3 0
34.000 2 3 2 2 4 5 4 4 2 4 2 3 2 2 2 2 2 5 2 5
35.000 0 1 0 1 0 1 2 0 0 0 2 1 0 0 0 1 0 0 0 0
36.000 0 0 0 0 0 0 0 0 0 0 0 0 0 0 0 0 0 0 0 1
37.000 0 0 2 0 0 0 0 0 0 0 0 0 0 2 0 0 0 2 2 0
38.000 2 1 3 3 2 3 2 2 0 2 0 1 0 0 1 2 0 5 4 1
39.000 2 2 0 3 0 2 0 1 0 0 0 0 0 1 0 2 0 0 2 0
40.000 0 0 0 0 0 1 0 0 0 0 1 1 1 0 0 0 1 1 0 1
41.000 2 2 3 0 2 1 2 2 0 1 2 0 0 2 0 1 4 1 2 0
42.000 0 4 1 4 2 3 4 2 0 0 4 0 0 0 4 0 0 1 6 5
47.000 0 0 0 0 0 0 0 0 0 2 1 1 0 0 0 1 2 1 0 0
48.000 0 1 1 0 1 0 1 0 2 0 0 3 1 0 0 4 2 1 1 1
49.000 2 0 0 1 3 1 6 0 0 2 2 0 0 1 0 1 0 0 0 1
50.000 1 1 2 1 4 0 1 0 2 0 0 1 1 0 0 3 1 3 1 0
51.000 0 0 0 0 0 0 0 0 0 0 0 4 1 0 0 2 4 0 0 0
52.000 0 0 0 1 2 1 0 0 1 1 2 3 0 2 1 0 1 0 0 4
53.000 3 0 0 0 1 3 0 2 2 2 3 1 0 10 1 0 0 0 1 7
54.000 0 2 1 0 0 0 0 2 1 1 3 1 3 1 1 1 1 1 0 1
55.000 1 2 4 0 2 1 1 1 0 2 1 0 2 1 0 1 3 0 0 1
56.000 0 0 0 0 1 1 1 0 0 0 0 1 0 1 0 0 0 0 0 1
57.000 0 1 1 0 1 0 0 0 0 0 0 0 0 2 0 1 0 2 0 0
58.000 0 1 1 1 1 1 0 1 1 0 2 1 1 3 1 1 0 1 1 1
59.000 0 2 0 1 0 2 3 2 0 0 0 2 0 1 2 0 4 1 0 1
60.000 0 0 2 0 0 0 0 1 0 0 0 0 1 0 1 2 0 3 0 0
61.000 0 1 1 0 0 1 0 0 0 0 1 1 2 1 3 1 1 2 1 1
62.000 0 1 0 1 0 0 0 1 0 0 0 1 1 1 1 0 1 2 0 0
63.000 1 3 3 0 0 0 0 6 4 0 0 0 1 2 2 0 1 3 0 2
64.000 0 5 1 0 0 0 0 1 0 0 0 0 0 3 0 0 1 1 0 2
65.000 0 1 0 3 0 0 0 0 0 0 2 2 0 0 0 0 1 0 0 0
66.000 0 0 0 0 1 0 0 1 0 0 0 0 0 1 0 1 1 3 0 2
67.000 0 1 1 0 0 0 0 1 0 1 0 1 0 0 0 4 2 2 1 1
68.000 0 4 0 3 0 0 0 0 0 0 0 0 0 1 0 0 0 0 0 0
69.000 0 1 0 0 0 0 0 1 0 0 1 0 0 0 1 0 2 2 0 0
97.000 2 2 1 1 2 1 2 4 5 4 5 1 1 1 1 3 2 2 2 2
98.000 2 2 1 1 1 1 1 2 2 2 2 1 1 1 1 1 1 2 2 2
99.000 0 0 0 0 1 0 1 2 4 2 2 0 0 0 0 0 2 0 0 0
100.000 1 0 1 0 1 1 1 1 1 1 0 1 0 1 1 1 1 1 1 0
101.000 0 0 0 1 0 0 0 0 0 2 0 0 0 0 1 0 0 0 0 0
102.000 0 0 0 0 0 0 0 0 0 0 0 0 0 0 0 0 0 0 0 0
103.000 0 0 0 0 1 0 0 1 1 2 0 0 2 0 0 0 3 0 0 0
104.000 0 1 0 7 0 0 0 0 1 7 0 5 0 5 1 0 1 2 0 1
105.000 1 0 1 1 2 0 0 0 1 1 1 1 1 0 0 1 0 1 0 1
106.000 0 0 1 0 0 2 2 1 1 1 1 0 1 0 0 2 1 2 0 0
107.000 0 4 0 0 1 1 0 2 0 1 0 0 2 1 0 1 3 0 1 3
108.000 1 3 1 2 3 2 2 0 1 1 0 1 1 1 0 3 1 2 0 0
109.000 2 1 2 2 4 1 2 0 2 2 1 2 2 1 0 1 1 3 0 1
112.000 1 0 1 0 0 1 0 0 0 0 0 1 0 0 0 0 0 0 0 1
113.000 1 0 0 0 0 0 0 0 0 0 0 1 0 0 0 0 0 0 0 0
114.000 1 1 2 0 0 0 0 0 0 0 2 0 0 0 0 2 0 1 0 0
115.000 0 3 0 0 0 0 0 0 1 0 3 0 0 0 0 1 0 1 1 0
116.000 0 0 0 0 1 1 1 0 0 1 0 1 0 0 0 1 0 1 0 0
117.000 1 1 2 2 1 3 3 2 3 1 0 3 3 2 1 4 0 3 3 2
118.000 0 0 0 0 0 1 2 0 4 0 0 0 1 0 0 0 0 0 1 0
119.000 1 0 2 2 1 0 2 1 2 1 0 2 0 2 0 1 0 2 1 1
120.000 2 4 3 4 4 6 4 2 5 5 4 3 6 5 6 6 4 2 4 4
121.000 0 1 1 2 2 2 5 0 2 3 0 4 3 1 5 1 5 0 2 5
122.000 1 0 2 0 0 1 0 0 0 0 1 0 0 0 0 1 0 0 0 0
123.000 3 1 3 4 6 1 1 3 2 3 4 2 0 3 1 1 2 3 3 5
124.000 4 4 2 4 2 2 1 2 6 3 4 4 3 3 3 3 1 1 3 4
125.000 1 1 1 0 0 1 0 2 2 0 1 2 1 0 0 0 2 0 0 2
126.000 0 0 0 0 1 0 0 0 0 0 0 0 0 0 0 0 0 0 0 0
127.000 0 2 0 0 0 0 0 2 1 0 2 0 2 1 0 2 0 1 0 2
128.000 1 3 0 3 0 2 0 5 4 1 4 3 0 3 1 2 2 3 3 4
129.000 0 0 0 1 0 2 0 2 2 0 0 2 0 2 0 2 3 2 2 0
130.000 1 0 0 0 0 0 0 0 2 0 0 1 0 0 0 0 0 1 0 1
131.000 0 4 0 1 1 1 0 2 0 2 0 0 2 2 3 2 1 0 1 1
132.000 0 1 0 0 5 0 0 6 1 0 0 4 4 0 1 0 0 0 0 0
137.000 2 2 0 1 0 0 0 0 2 2 0 0 0 2 0 0 4 0 0 0
138.000 0 0 0 2 3 0 0 0 0 1 0 0 0 1 5 0 4 1 1 2
139.000 0 1 1 0 0 2 0 0 1 2 1 1 0 0 0 0 0 3 1 0
140.000 0 1 0 1 2 2 0 0 1 1 2 0 0 0 3 1 1 2 1 1
141.000 0 0 1 0 3 0 0 1 0 0 0 0 0 0 2 0 3 1 0 0
142.000 0 0 0 2 1 0 0 0 0 1 0 0 0 0 0 0 0 0 1 2
143.000 0 0 2 1 1 0 2 0 0 0 0 7 0 0 2 0 0 0 1 1
144.000 1 1 2 0 3 2 1 1 2 1 1 0 2 1 0 1 0 2 3 1
145.000 0 1 0 0 9 1 1 0 0 4 2 0 2 3 3 0 0 1 0 1
146.000 0 1 0 2 0 0 0 0 0 0 0 1 0 0 0 0 0 1 0 0
147.000 0 0 0 0 0 1 0 0 0 0 0 0 0 0 0 0 0 0 0 1
148.000 1 1 1 0 0 1 3 2 2 1 2 0 3 0 0 1 0 0 1 1
149.000 1 0 1 0 0 0 4 0 1 0 0 3 0 0 3 3 0 2 0 0
150.000 0 0 0 0 0 1 1 0 0 0 1 0 2 0 0 0 0 0 0 0
151.000 0 0 0 0 0 0 1 2 1 2 1 0 2 0 0 0 0 1 0 0
152.000 0 0 1 0 0 0 0 0 0 1 1 0 1 0 0 0 0 1 0 0
153.000 1 4 2 3 0 0 2 0 2 0 2 2 0 0 0 5 0 3 0 0
154.000 2 1 0 1 0 1 1 0 0 1 1 1 0 0 0 1 0 2 0 0
155.000 0 1 1 0 0 1 0 0 0 0 0 0 0 0 0 0 1 0 0 1
156.000 0 0 0 1 0 0 0 0 0 0 0 0 0 0 0 0 0 1 0 1
157.000 0 0 0 4 0 1 0 0 0 0 0 0 0 0 1 6 0 0 2 0
158.000 1 1 0 1 0 1 0 0 0 1 1 1 0 0 0 0 0 1 0 0
#Reported_Model_Average 0.661 1.009 0.786 0.893 1.107 0.946 0.929 0.875 0.830 0.946 0.946 1.062 0.839 0.991 0.848 1.116 1.018 1.143 0.920 1.080
#Overall_Average_Reported 0.947
List of bad contacts calculated by MAGE for model $num_n
/farm/software/bin/probe
: 2637:M 98 PRO 1HD :M 97 VAL N : -0.626: 0
: 2637:M 97 VAL N :M 98 PRO CD : -0.433: 0
: 2637:M 8 PRO CD :M 7 VAL N : -0.612: 0
: 2637:M 8 PRO 1HD :M 7 VAL N : -0.421: 0
: 2637:M 124 TYR O :M 128 VAL 3HG2 : -0.609: 0
: 2637:M 124 TYR CB :M 120 MET O : -0.466: 0
: 2637:M 120 MET O :M 124 TYR N : -0.419: 0
: 2637:M 124 TYR CE1 :M 149 LEU 3HD2 : -0.409: 0
: 2637:M 30 MET O :M 34 TYR N : -0.600: 0
: 2637:M 30 MET SD :M 31 ASN N : -0.587: 0
: 2637:M 30 MET CG :M 31 ASN N : -0.470: 0
: 2637:M 34 TYR CB :M 30 MET O : -0.464: 0
: 2637:M 135 GLN O :M 136 LEU C : -0.536: 0
: 2637:M 137 THR N :M 135 GLN O : -0.510: 0
: 2637:M 137 THR N :M 135 GLN C : -0.400: 0
: 2637:M 167 HIS N :M 166 HIS CG : -0.533: 0
: 2637:M 167 HIS O :M 167 HIS CG : -0.432: 0
: 2637:M 166 HIS CD2 :M 167 HIS N : -0.412: 0
: 2637:M 133 GLN O :M 134 ASP C : -0.530: 0
: 2637:M 130 THR O :M 134 ASP N : -0.430: 0
: 2637:M 125 ARG NH2 :M 63 TYR OH : -0.529: 0
: 2637:M 91 MET SD :M 91 MET N : -0.521: 0
: 2637:M 96 SER H :M 94 LEU C : -0.519: 0
: 2637:M 21 ASN C :M 23 SER H : -0.516: 0
: 2637:M 22 GLN O :M 23 SER CB : -0.427: 0
: 2637:M 23 SER OG :M 23 SER O : -0.418: 0
: 2637:M 22 GLN O :M 23 SER OG : -0.402: 0
: 2637:M 154 ASN 1HD2 :M 158 ARG NH2 : -0.505: 0
: 2637:M 153 TYR CD1 :M 154 ASN N : -0.452: 0
: 2637:M 117 ILE C :M 119 ALA H : -0.502: 0
: 2637:M 55 ARG CZ :M 41 LEU 2HD2 : -0.502: 0
: 2637:M 44 ASP N :M 44 ASP OD1 : -0.451: 0
: 2637:M 41 LEU O :M 44 ASP OD1 : -0.446: 0
: 2637:M 44 ASP OD1 :M 45 GLN N : -0.438: 0
: 2637:M 44 ASP CG :M 45 GLN N : -0.426: 0
: 2637:M 45 GLN H :M 44 ASP CG : -0.424: 0
: 2637:M 39 SER N :M 38 VAL 3HG2 : -0.490: 0
: 2637:M 39 SER N :M 38 VAL CG2 : -0.421: 0
: 2637:M 27 ILE C :M 29 ALA H : -0.471: 0
: 2637:M 29 ALA N :M 27 ILE C : -0.449: 0
: 2637:M 10 PHE CD2 :M 33 TYR OH : -0.467: 0
: 2637:M 14 LEU 1HD2 :M 33 TYR CD1 : -0.441: 0
: 2637:M 53 LEU 3HD1 :M 53 LEU O : -0.461: 0
: 2637:M 49 ASN O :M 53 LEU 1HB : -0.448: 0
: 2637:M 49 ASN OD1 :M 50 ALA N : -0.429: 0
: 2637:M 77 HIS O :M 77 HIS CG : -0.457: 0
: 2637:M 5 PHE H :M 3 GLU C : -0.444: 0
: 2637:M 5 PHE CE2 :M 6 SER OG : -0.437: 0
: 2637:M 5 PHE N :M 3 GLU C : -0.433: 0
: 2637:M 93 GLU CD :M 93 GLU N : -0.429: 0
: 2637:M 123 TYR CD1 :M 123 TYR C : -0.427: 0
: 2637:M 100 PHE CD2 :M 123 TYR OH : -0.423: 0
: 2637:M 114 GLU OE2 :M 122 SER CB : -0.422: 0
: 2637:M 112 GLN O :M 113 SER OG : -0.421: 0
: 2637:M 24 GLU OE2 :M 32 SER OG : -0.410: 0
: 2637:M 109 GLU CB :M 105 LYS O : -0.409: 0
: 2637:M 18 ILE 2HG1 :M 19 GLU N : -0.409: 0
: 2637:M 109 GLU N :M 108 ILE 2HG1 : -0.401: 0
: 2637:M 144 LYS O :M 148 HIS CD2 : -0.405: 0
#sum2 ::22.37 clashscore : 22.37 clashscore B<40
#summary::2637 atoms:2637 atoms B<40:296604 potential dots:18540.0 A^2:59 bumps:59 bumps B<40:627.5 score
List of bad contacts calculated by MAGE for model $num_n
/farm/software/bin/probe
: 2637:M 124 TYR O :M 128 VAL 3HG2 : -0.819: 0
: 2637:M 120 MET O :M 124 TYR N : -0.558: 0
: 2637:M 153 TYR CD1 :M 154 ASN N : -0.483: 0
: 2637:M 124 TYR CB :M 120 MET O : -0.458: 0
: 2637:M 128 VAL 2HG2 :M 146 ILE 2HD1 : -0.443: 0
: 2637:M 117 ILE O :M 120 MET CG : -0.441: 0
: 2637:M 153 TYR CD1 :M 153 TYR C : -0.436: 0
: 2637:M 128 VAL 1HG2 :M 124 TYR CE1 : -0.433: 0
: 2637:M 120 MET SD :M 153 TYR O : -0.428: 0
: 2637:M 4 LEU N :M 4 LEU 2HD1 : -0.662: 0
: 2637:M 4 LEU O :M 6 SER N : -0.587: 0
: 2637:M 4 LEU N :M 4 LEU CD1 : -0.543: 0
: 2637:M 4 LEU H :M 4 LEU 2HD1 : -0.469: 0
: 2637:M 8 PRO CD :M 7 VAL N : -0.652: 0
: 2637:M 42 VAL O :M 42 VAL 2HG1 : -0.651: 0
: 2637:M 42 VAL O :M 42 VAL CG1 : -0.406: 0
: 2637:M 30 MET O :M 34 TYR N : -0.647: 0
: 2637:M 34 TYR CG :M 125 ARG NH2 : -0.521: 0
: 2637:M 38 VAL 1HG2 :M 34 TYR CE1 : -0.464: 0
: 2637:M 27 ILE C :M 29 ALA H : -0.590: 0
: 2637:M 67 LYS 2HD :M 27 ILE 1HD1 : -0.474: 0
: 2637:M 70 GLU OE2 :M 27 ILE 2HD1 : -0.457: 0
: 2637:M 27 ILE C :M 29 ALA N : -0.425: 0
: 2637:M 115 ASP O :M 107 HIS NE2 : -0.573: 0
: 2637:M 115 ASP O :M 107 HIS CD2 : -0.461: 0
: 2637:M 115 ASP N :M 107 HIS NE2 : -0.450: 0
: 2637:M 114 GLU N :M 107 HIS CE1 : -0.441: 0
: 2637:M 64 ASN ND2 :M 68 ARG NE : -0.572: 0
: 2637:M 64 ASN 1HD2 :M 68 ARG CD : -0.543: 0
: 2637:M 68 ARG CZ :M 64 ASN OD1 : -0.463: 0
: 2637:M 63 TYR CD1 :M 64 ASN N : -0.462: 0
: 2637:M 63 TYR CD1 :M 63 TYR C : -0.412: 0
: 2637:M 68 ARG NE :M 64 ASN 1HD2 : -0.407: 0
: 2637:M 127 VAL CG1 :M 131 LEU 3HD2 : -0.554: 0
: 2637:M 131 LEU 3HD2 :M 127 VAL 2HG1 : -0.551: 0
: 2637:M 145 ARG 1HD :M 131 LEU 1HD1 : -0.494: 0
: 2637:M 134 ASP OD1 :M 134 ASP C : -0.482: 0
: 2637:M 131 LEU O :M 134 ASP OD1 : -0.458: 0
: 2637:M 59 LEU 3HD1 :M 59 LEU O : -0.551: 0
: 2637:M 98 PRO CD :M 97 VAL N : -0.542: 0
: 2637:M 98 PRO 1HD :M 97 VAL N : -0.410: 0
: 2637:M 121 ASN ND2 :M 31 ASN OD1 : -0.495: 0
: 2637:M 163 LEU N :M 161 SER O : -0.494: 0
: 2637:M 163 LEU C :M 165 HIS H : -0.462: 0
: 2637:M 164 GLU O :M 165 HIS O : -0.456: 0
: 2637:M 165 HIS N :M 163 LEU C : -0.441: 0
: 2637:M 65 LYS O :M 69 GLY N : -0.492: 0
: 2637:M 139 ASN OD1 :M 140 ALA N : -0.491: 0
: 2637:M 21 ASN OD1 :M 25 ASP N : -0.489: 0
: 2637:M 20 MET O :M 21 ASN CB : -0.447: 0
: 2637:M 21 ASN CA :M 17 HIS O : -0.416: 0
: 2637:M 144 LYS O :M 148 HIS CD2 : -0.485: 0
: 2637:M 41 LEU C :M 43 GLN H : -0.460: 0
: 2637:M 55 ARG HE :M 41 LEU CD1 : -0.459: 0
: 2637:M 35 ARG O :M 39 SER CB : -0.438: 0
: 2637:M 54 LYS CG :M 55 ARG N : -0.426: 0
: 2637:M 39 SER O :M 43 GLN CD : -0.410: 0
: 2637:M 54 LYS O :M 58 HIS ND1 : -0.404: 0
: 2637:M 10 PHE CD2 :M 33 TYR OH : -0.455: 0
: 2637:M 33 TYR O :M 33 TYR CD1 : -0.453: 0
: 2637:M 33 TYR CD1 :M 33 TYR C : -0.409: 0
: 2637:M 158 ARG 1HH2 :M 155 LYS CE : -0.451: 0
: 2637:M 111 ASN N :M 111 ASN OD1 : -0.448: 0
: 2637:M 44 ASP O :M 46 LEU N : -0.441: 0
: 2637:M 45 GLN O :M 46 LEU C : -0.432: 0
: 2637:M 44 ASP O :M 45 GLN 2HB : -0.414: 0
: 2637:M 80 HIS CG :M 79 HIS O : -0.437: 0
: 2637:M 79 HIS C :M 80 HIS CG : -0.416: 0
: 2637:M 109 GLU N :M 108 ILE CG1 : -0.427: 0
: 2637:M 108 ILE H :M 108 ILE 3HG2 : -0.403: 0
: 2637:M 62 ALA N :M 61 GLU CG : -0.426: 0
: 2637:M 57 GLN 1HE2 :M 132 VAL 1HG1 : -0.425: 0
: 2637:M 137 THR 2HG2 :M 137 THR H : -0.422: 0
: 2637:M 48 LYS 1HG :M 50 ALA 3HB : -0.401: 0
: 2637:M 104 LEU 1HD2 :M 123 TYR CD1 : -0.400: 0
#sum2 ::28.44 clashscore : 28.44 clashscore B<40
#summary::2637 atoms:2637 atoms B<40:296534 potential dots:18530.0 A^2:75 bumps:75 bumps B<40:515.2 score
List of bad contacts calculated by MAGE for model $num_n
/farm/software/bin/probe
: 2637:M 98 PRO CD :M 97 VAL N : -0.640: 0
: 2637:M 41 LEU 2HD2 :M 55 ARG 1HH1 : -0.626: 0
: 2637:M 44 ASP O :M 45 GLN CB : -0.491: 0
: 2637:M 45 GLN 1HE2 :M 55 ARG CZ : -0.489: 0
: 2637:M 44 ASP O :M 45 GLN 2HB : -0.476: 0
: 2637:M 41 LEU 2HD2 :M 55 ARG NH1 : -0.472: 0
: 2637:M 55 ARG NH2 :M 45 GLN 1HE2 : -0.470: 0
: 2637:M 41 LEU O :M 44 ASP O : -0.433: 0
: 2637:M 44 ASP OD1 :M 44 ASP C : -0.420: 0
: 2637:M 27 ILE 1HD1 :M 67 LYS 1HD : -0.623: 0
: 2637:M 27 ILE C :M 29 ALA H : -0.580: 0
: 2637:M 27 ILE C :M 29 ALA N : -0.434: 0
: 2637:M 28 HIS O :M 28 HIS CG : -0.417: 0
: 2637:M 27 ILE 3HG2 :M 28 HIS N : -0.414: 0
: 2637:M 95 PHE H :M 93 GLU C : -0.597: 0
: 2637:M 95 PHE N :M 93 GLU C : -0.463: 0
: 2637:M 163 LEU O :M 165 HIS N : -0.574: 0
: 2637:M 165 HIS ND1 :M 165 HIS N : -0.543: 0
: 2637:M 165 HIS CD2 :M 163 LEU 1HD1 : -0.523: 0
: 2637:M 165 HIS N :M 163 LEU C : -0.499: 0
: 2637:M 165 HIS ND1 :M 166 HIS N : -0.439: 0
: 2637:M 163 LEU C :M 165 HIS H : -0.433: 0
: 2637:M 163 LEU O :M 164 GLU C : -0.421: 0
: 2637:M 165 HIS CG :M 166 HIS N : -0.412: 0
: 2637:M 8 PRO CD :M 7 VAL N : -0.547: 0
: 2637:M 117 ILE C :M 119 ALA H : -0.527: 0
: 2637:M 117 ILE C :M 119 ALA N : -0.409: 0
: 2637:M 120 MET CG :M 121 ASN N : -0.526: 0
: 2637:M 124 TYR CB :M 120 MET O : -0.468: 0
: 2637:M 120 MET O :M 124 TYR N : -0.405: 0
: 2637:M 48 LYS 1HG :M 50 ALA H : -0.516: 0
: 2637:M 139 ASN 2HD2 :M 50 ALA N : -0.441: 0
: 2637:M 38 VAL N :M 37 VAL 3HG2 : -0.490: 0
: 2637:M 38 VAL N :M 37 VAL CG2 : -0.489: 0
: 2637:M 42 VAL 3HG2 :M 38 VAL O : -0.418: 0
: 2637:M 60 ASP OD1 :M 61 GLU N : -0.486: 0
: 2637:M 57 GLN O :M 60 ASP OD1 : -0.438: 0
: 2637:M 125 ARG NH2 :M 31 ASN OD1 : -0.476: 0
: 2637:M 77 HIS ND1 :M 79 HIS O : -0.476: 0
: 2637:M 77 HIS CE1 :M 79 HIS O : -0.408: 0
: 2637:M 112 GLN NE2 :M 106 GLN CD : -0.473: 0
: 2637:M 155 LYS O :M 159 GLY N : -0.467: 0
: 2637:M 34 TYR CB :M 30 MET O : -0.466: 0
: 2637:M 30 MET O :M 34 TYR N : -0.414: 0
: 2637:M 114 GLU OE2 :M 122 SER CB : -0.465: 0
: 2637:M 122 SER OG :M 114 GLU OE2 : -0.402: 0
: 2637:M 100 PHE CD2 :M 123 TYR OH : -0.460: 0
: 2637:M 123 TYR CD1 :M 123 TYR C : -0.430: 0
: 2637:M 144 LYS O :M 148 HIS CD2 : -0.456: 0
: 2637:M 144 LYS CG :M 141 VAL O : -0.410: 0
: 2637:M 63 TYR CD1 :M 64 ASN N : -0.450: 0
: 2637:M 63 TYR CD1 :M 63 TYR C : -0.402: 0
: 2637:M 108 ILE CG1 :M 109 GLU N : -0.434: 0
: 2637:M 109 GLU CB :M 105 LYS O : -0.414: 0
: 2637:M 33 TYR CD1 :M 33 TYR C : -0.431: 0
: 2637:M 152 ALA 3HB :M 149 LEU 3HD1 : -0.426: 0
: 2637:M 24 GLU OE2 :M 32 SER CB : -0.414: 0
: 2637:M 153 TYR CD1 :M 153 TYR C : -0.411: 0
: 2637:M 160 GLU OE2 :M 161 SER O : -0.408: 0
: 2637:M 143 LEU C :M 143 LEU CD1 : -0.407: 0
: 2637:M 15 LYS O :M 19 GLU CB : -0.406: 0
: 2637:M 54 LYS O :M 58 HIS CD2 : -0.402: 0
#sum2 ::23.51 clashscore : 23.51 clashscore B<40
#summary::2637 atoms:2637 atoms B<40:296633 potential dots:18540.0 A^2:62 bumps:62 bumps B<40:598.8 score
List of bad contacts calculated by MAGE for model $num_n
/farm/software/bin/probe
: 2637:M 124 TYR O :M 128 VAL 3HG2 : -0.778: 0
: 2637:M 157 LYS O :M 159 GLY N : -0.481: 0
: 2637:M 159 GLY N :M 157 LYS C : -0.473: 0
: 2637:M 128 VAL 1HG2 :M 124 TYR CE1 : -0.459: 0
: 2637:M 121 ASN OD1 :M 35 ARG NH1 : -0.457: 0
: 2637:M 124 TYR CB :M 120 MET O : -0.457: 0
: 2637:M 120 MET SD :M 157 LYS CG : -0.450: 0
: 2637:M 157 LYS O :M 158 ARG C : -0.440: 0
: 2637:M 142 VAL O :M 146 ILE 1HG1 : -0.435: 0
: 2637:M 120 MET O :M 124 TYR N : -0.427: 0
: 2637:M 143 LEU N :M 142 VAL 3HG2 : -0.424: 0
: 2637:M 128 VAL 2HG2 :M 146 ILE 2HD1 : -0.414: 0
: 2637:M 120 MET 2HG :M 121 ASN H : -0.405: 0
: 2637:M 42 VAL O :M 42 VAL 2HG1 : -0.696: 0
: 2637:M 42 VAL O :M 42 VAL CG1 : -0.500: 0
: 2637:M 104 LEU N :M 104 LEU 2HD1 : -0.643: 0
: 2637:M 104 LEU 1HD1 :M 123 TYR CE1 : -0.532: 0
: 2637:M 104 LEU 1HD1 :M 123 TYR CD1 : -0.531: 0
: 2637:M 104 LEU N :M 104 LEU CD1 : -0.525: 0
: 2637:M 123 TYR CD1 :M 123 TYR C : -0.415: 0
: 2637:M 101 ILE O :M 104 LEU N : -0.414: 0
: 2637:M 68 ARG NH2 :M 65 LYS NZ : -0.559: 0
: 2637:M 65 LYS NZ :M 68 ARG 1HH2 : -0.536: 0
: 2637:M 68 ARG 1HH2 :M 65 LYS CE : -0.499: 0
: 2637:M 98 PRO CD :M 97 VAL N : -0.557: 0
: 2637:M 30 MET SD :M 31 ASN N : -0.556: 0
: 2637:M 39 SER N :M 38 VAL 3HG1 : -0.506: 0
: 2637:M 39 SER N :M 38 VAL CG1 : -0.482: 0
: 2637:M 34 TYR CB :M 30 MET O : -0.455: 0
: 2637:M 30 MET C :M 30 MET SD : -0.424: 0
: 2637:M 39 SER O :M 43 GLN CB : -0.407: 0
: 2637:M 34 TYR O :M 38 VAL 2HG1 : -0.403: 0
: 2637:M 7 VAL CG1 :M 58 HIS NE2 : -0.515: 0
: 2637:M 8 PRO CD :M 7 VAL N : -0.499: 0
: 2637:M 8 PRO 1HD :M 7 VAL N : -0.451: 0
: 2637:M 7 VAL 3HG2 :M 7 VAL H : -0.439: 0
: 2637:M 62 ALA 3HB :M 59 LEU O : -0.497: 0
: 2637:M 75 HIS CD2 :M 73 LEU O : -0.494: 0
: 2637:M 73 LEU N :M 73 LEU 2HD2 : -0.458: 0
: 2637:M 73 LEU N :M 73 LEU CD2 : -0.446: 0
: 2637:M 156 VAL O :M 160 GLU CG : -0.488: 0
: 2637:M 108 ILE 2HD1 :M 160 GLU OE1 : -0.451: 0
: 2637:M 108 ILE CG1 :M 109 GLU N : -0.438: 0
: 2637:M 109 GLU CB :M 105 LYS O : -0.409: 0
: 2637:M 26 LYS HA :M 18 ILE 2HG2 : -0.479: 0
: 2637:M 18 ILE CG1 :M 19 GLU N : -0.472: 0
: 2637:M 19 GLU CB :M 15 LYS O : -0.416: 0
: 2637:M 153 TYR CD1 :M 153 TYR C : -0.474: 0
: 2637:M 154 ASN N :M 153 TYR CG : -0.469: 0
: 2637:M 169 HIS CG :M 168 HIS O : -0.469: 0
: 2637:M 5 PHE N :M 3 GLU C : -0.462: 0
: 2637:M 5 PHE H :M 3 GLU C : -0.445: 0
: 2637:M 131 LEU O :M 134 ASP O : -0.455: 0
: 2637:M 170 HIS N :M 170 HIS ND1 : -0.453: 0
: 2637:M 129 SER O :M 133 GLN CB : -0.452: 0
: 2637:M 163 LEU O :M 164 GLU O : -0.445: 0
: 2637:M 162 LYS O :M 164 GLU N : -0.403: 0
: 2637:M 49 ASN OD1 :M 50 ALA N : -0.439: 0
: 2637:M 140 ALA H :M 138 LYS 2HG : -0.433: 0
: 2637:M 138 LYS N :M 137 THR 3HG2 : -0.426: 0
: 2637:M 45 GLN O :M 52 VAL CG1 : -0.429: 0
: 2637:M 166 HIS CE1 :M 165 HIS CE1 : -0.417: 0
: 2637:M 166 HIS ND1 :M 166 HIS N : -0.401: 0
: 2637:M 117 ILE C :M 119 ALA H : -0.414: 0
: 2637:M 117 ILE C :M 119 ALA N : -0.405: 0
: 2637:M 76 HIS O :M 76 HIS CG : -0.401: 0
#sum2 ::25.03 clashscore : 25.03 clashscore B<40
#summary::2637 atoms:2637 atoms B<40:296904 potential dots:18560.0 A^2:66 bumps:66 bumps B<40:582.5 score
List of bad contacts calculated by MAGE for model $num_n
/farm/software/bin/probe
: 2637:M 34 TYR O :M 38 VAL 3HG2 : -0.802: 0
: 2637:M 30 MET SD :M 31 ASN N : -0.527: 0
: 2637:M 31 ASN ND2 :M 30 MET CE : -0.498: 0
: 2637:M 30 MET O :M 34 TYR N : -0.473: 0
: 2637:M 34 TYR CB :M 30 MET O : -0.469: 0
: 2637:M 38 VAL 1HG2 :M 34 TYR CE2 : -0.451: 0
: 2637:M 31 ASN N :M 30 MET CG : -0.421: 0
: 2637:M 31 ASN 1HD2 :M 30 MET 1HE : -0.417: 0
: 2637:M 141 VAL 2HG2 :M 138 LYS O : -0.690: 0
: 2637:M 142 VAL N :M 141 VAL 3HG2 : -0.433: 0
: 2637:M 140 ALA H :M 138 LYS 2HG : -0.424: 0
: 2637:M 143 LEU N :M 140 ALA O : -0.420: 0
: 2637:M 138 LYS 2HB :M 141 VAL 3HG1 : -0.404: 0
: 2637:M 132 VAL O :M 132 VAL 2HG1 : -0.667: 0
: 2637:M 57 GLN NE2 :M 132 VAL 1HG1 : -0.605: 0
: 2637:M 132 VAL O :M 132 VAL CG1 : -0.477: 0
: 2637:M 49 ASN 2HD2 :M 50 ALA N : -0.651: 0
: 2637:M 48 LYS 1HG :M 50 ALA H : -0.485: 0
: 2637:M 50 ALA N :M 49 ASN ND2 : -0.444: 0
: 2637:M 50 ALA H :M 49 ASN ND2 : -0.409: 0
: 2637:M 124 TYR N :M 120 MET O : -0.631: 0
: 2637:M 120 MET SD :M 121 ASN N : -0.555: 0
: 2637:M 124 TYR 1HB :M 120 MET O : -0.460: 0
: 2637:M 120 MET CG :M 121 ASN N : -0.440: 0
: 2637:M 96 SER CB :M 95 PHE O : -0.603: 0
: 2637:M 94 LEU O :M 96 SER N : -0.560: 0
: 2637:M 94 LEU N :M 94 LEU 2HD2 : -0.542: 0
: 2637:M 94 LEU N :M 94 LEU CD2 : -0.513: 0
: 2637:M 98 PRO 1HD :M 97 VAL N : -0.498: 0
: 2637:M 99 TYR 2HB :M 96 SER O : -0.460: 0
: 2637:M 96 SER O :M 97 VAL C : -0.452: 0
: 2637:M 8 PRO CD :M 7 VAL N : -0.597: 0
: 2637:M 8 PRO 1HD :M 7 VAL N : -0.419: 0
: 2637:M 27 ILE C :M 29 ALA H : -0.567: 0
: 2637:M 27 ILE 1HG1 :M 26 LYS 3HZ : -0.480: 0
: 2637:M 29 ALA N :M 27 ILE C : -0.435: 0
: 2637:M 145 ARG HE :M 145 ARG N : -0.554: 0
: 2637:M 145 ARG N :M 145 ARG NE : -0.518: 0
: 2637:M 145 ARG HE :M 144 LYS C : -0.495: 0
: 2637:M 145 ARG CA :M 145 ARG NE : -0.483: 0
: 2637:M 145 ARG CD :M 145 ARG N : -0.472: 0
: 2637:M 144 LYS 1HG :M 144 LYS H : -0.403: 0
: 2637:M 163 LEU 2HD1 :M 163 LEU N : -0.505: 0
: 2637:M 163 LEU O :M 165 HIS N : -0.495: 0
: 2637:M 165 HIS CG :M 166 HIS N : -0.494: 0
: 2637:M 163 LEU H :M 163 LEU 2HD1 : -0.486: 0
: 2637:M 164 GLU C :M 163 LEU O : -0.455: 0
: 2637:M 163 LEU CD1 :M 163 LEU N : -0.430: 0
: 2637:M 165 HIS ND1 :M 166 HIS N : -0.411: 0
: 2637:M 162 LYS O :M 163 LEU C : -0.407: 0
: 2637:M 117 ILE C :M 119 ALA H : -0.496: 0
: 2637:M 52 VAL O :M 56 ILE 1HG1 : -0.491: 0
: 2637:M 53 LEU N :M 52 VAL 3HG2 : -0.439: 0
: 2637:M 20 MET CB :M 16 GLN O : -0.476: 0
: 2637:M 170 HIS N :M 168 HIS O : -0.474: 0
: 2637:M 170 HIS ND1 :M 168 HIS O : -0.400: 0
: 2637:M 105 LYS NZ :M 109 GLU OE1 : -0.450: 0
: 2637:M 108 ILE CG1 :M 109 GLU N : -0.436: 0
: 2637:M 109 GLU CB :M 105 LYS O : -0.428: 0
: 2637:M 116 LYS O :M 108 ILE 2HG2 : -0.410: 0
: 2637:M 108 ILE 2HG1 :M 109 GLU N : -0.401: 0
: 2637:M 41 LEU O :M 44 ASP O : -0.447: 0
: 2637:M 42 VAL C :M 44 ASP N : -0.443: 0
: 2637:M 55 ARG 2HD :M 41 LEU 2HD2 : -0.436: 0
: 2637:M 55 ARG O :M 58 HIS ND1 : -0.435: 0
: 2637:M 44 ASP CG :M 43 GLN O : -0.429: 0
: 2637:M 44 ASP H :M 42 VAL C : -0.416: 0
: 2637:M 131 LEU C :M 133 GLN H : -0.446: 0
: 2637:M 14 LEU 3HD1 :M 66 VAL CG2 : -0.442: 0
: 2637:M 10 PHE CD2 :M 33 TYR OH : -0.414: 0
: 2637:M 33 TYR CD1 :M 14 LEU 1HD2 : -0.409: 0
: 2637:M 33 TYR CD1 :M 33 TYR C : -0.404: 0
: 2637:M 18 ILE CG1 :M 19 GLU N : -0.441: 0
: 2637:M 19 GLU CB :M 15 LYS O : -0.413: 0
: 2637:M 18 ILE 2HG1 :M 19 GLU N : -0.405: 0
: 2637:M 100 PHE CD2 :M 123 TYR OH : -0.438: 0
: 2637:M 123 TYR O :M 123 TYR CD1 : -0.427: 0
: 2637:M 123 TYR CD1 :M 123 TYR C : -0.405: 0
: 2637:M 126 SER HG :M 123 TYR C : -0.403: 0
: 2637:M 103 ASN ND2 :M 107 HIS NE2 : -0.431: 0
: 2637:M 3 GLU O :M 4 LEU CB : -0.417: 0
#sum2 ::30.72 clashscore : 30.72 clashscore B<40
#summary::2637 atoms:2637 atoms B<40:296644 potential dots:18540.0 A^2:81 bumps:81 bumps B<40:556.4 score
List of bad contacts calculated by MAGE for model $num_n
/farm/software/bin/probe
: 2637:M 157 LYS 1HD :M 117 ILE 1HD1 : -0.675: 0
: 2637:M 35 ARG 2HH1 :M 120 MET CE : -0.563: 0
: 2637:M 120 MET O :M 124 TYR N : -0.545: 0
: 2637:M 120 MET SD :M 121 ASN N : -0.537: 0
: 2637:M 124 TYR CB :M 120 MET O : -0.493: 0
: 2637:M 120 MET CG :M 121 ASN N : -0.451: 0
: 2637:M 117 ILE 3HG2 :M 118 HIS N : -0.417: 0
: 2637:M 120 MET 2HG :M 117 ILE O : -0.402: 0
: 2637:M 71 SER H :M 70 GLU CG : -0.639: 0
: 2637:M 139 ASN 2HD2 :M 140 ALA N : -0.639: 0
: 2637:M 70 GLU OE2 :M 27 ILE 3HD1 : -0.564: 0
: 2637:M 27 ILE C :M 29 ALA H : -0.558: 0
: 2637:M 74 GLU C :M 71 SER HG : -0.539: 0
: 2637:M 140 ALA N :M 139 ASN ND2 : -0.510: 0
: 2637:M 71 SER H :M 70 GLU CD : -0.489: 0
: 2637:M 70 GLU OE2 :M 27 ILE CD1 : -0.446: 0
: 2637:M 70 GLU CG :M 71 SER N : -0.437: 0
: 2637:M 29 ALA N :M 27 ILE C : -0.436: 0
: 2637:M 59 LEU 3HD1 :M 59 LEU O : -0.626: 0
: 2637:M 98 PRO CD :M 97 VAL N : -0.609: 0
: 2637:M 42 VAL O :M 42 VAL 2HG1 : -0.600: 0
: 2637:M 147 GLN NE2 :M 42 VAL 1HG1 : -0.442: 0
: 2637:M 8 PRO 1HD :M 7 VAL N : -0.586: 0
: 2637:M 9 TYR N :M 7 VAL C : -0.413: 0
: 2637:M 7 VAL N :M 8 PRO CD : -0.401: 0
: 2637:M 39 SER N :M 38 VAL 3HG2 : -0.567: 0
: 2637:M 31 ASN ND2 :M 30 MET CE : -0.553: 0
: 2637:M 30 MET SD :M 31 ASN N : -0.551: 0
: 2637:M 38 VAL CG2 :M 39 SER N : -0.487: 0
: 2637:M 34 TYR CB :M 30 MET O : -0.475: 0
: 2637:M 30 MET CE :M 31 ASN 1HD2 : -0.440: 0
: 2637:M 34 TYR O :M 38 VAL 2HG2 : -0.439: 0
: 2637:M 30 MET CG :M 31 ASN N : -0.436: 0
: 2637:M 34 TYR C :M 34 TYR CD1 : -0.421: 0
: 2637:M 31 ASN 1HD2 :M 30 MET 1HE : -0.410: 0
: 2637:M 30 MET O :M 34 TYR N : -0.400: 0
: 2637:M 73 LEU O :M 75 HIS N : -0.547: 0
: 2637:M 41 LEU 2HD2 :M 55 ARG 1HH1 : -0.522: 0
: 2637:M 129 SER N :M 128 VAL 3HG2 : -0.506: 0
: 2637:M 129 SER N :M 128 VAL CG2 : -0.452: 0
: 2637:M 44 ASP N :M 40 THR O : -0.485: 0
: 2637:M 58 HIS ND1 :M 61 GLU OE2 : -0.483: 0
: 2637:M 150 ASP O :M 154 ASN ND2 : -0.468: 0
: 2637:M 20 MET CB :M 16 GLN O : -0.462: 0
: 2637:M 100 PHE CD2 :M 123 TYR OH : -0.462: 0
: 2637:M 110 MET CB :M 106 GLN O : -0.457: 0
: 2637:M 112 GLN 2HE2 :M 110 MET 2HG : -0.440: 0
: 2637:M 111 ASN CA :M 107 HIS O : -0.420: 0
: 2637:M 106 GLN O :M 110 MET N : -0.412: 0
: 2637:M 110 MET O :M 111 ASN CB : -0.410: 0
: 2637:M 49 ASN O :M 53 LEU 1HB : -0.445: 0
: 2637:M 53 LEU 3HD1 :M 53 LEU O : -0.430: 0
: 2637:M 19 GLU N :M 18 ILE CG1 : -0.442: 0
: 2637:M 19 GLU CB :M 15 LYS O : -0.413: 0
: 2637:M 116 LYS O :M 108 ILE 2HG2 : -0.440: 0
: 2637:M 108 ILE CG1 :M 109 GLU N : -0.420: 0
: 2637:M 10 PHE CD2 :M 33 TYR OH : -0.431: 0
: 2637:M 131 LEU O :M 134 ASP O : -0.431: 0
: 2637:M 52 VAL O :M 56 ILE 2HG1 : -0.431: 0
: 2637:M 134 ASP O :M 135 GLN 2HB : -0.404: 0
: 2637:M 148 HIS CG :M 144 LYS O : -0.430: 0
: 2637:M 145 ARG N :M 144 LYS CG : -0.415: 0
: 2637:M 125 ARG 1HG :M 122 SER O : -0.422: 0
: 2637:M 168 HIS CD2 :M 170 HIS C : -0.405: 0
: 2637:M 155 LYS 1HD :M 158 ARG 1HH2 : -0.405: 0
#sum2 ::24.65 clashscore : 24.65 clashscore B<40
#summary::2637 atoms:2637 atoms B<40:296685 potential dots:18540.0 A^2:65 bumps:65 bumps B<40:560.8 score
List of bad contacts calculated by MAGE for model $num_n
/farm/software/bin/probe
: 2637:M 38 VAL 1HG2 :M 34 TYR CE1 : -0.746: 0
: 2637:M 59 LEU 3HD1 :M 59 LEU O : -0.607: 0
: 2637:M 30 MET SD :M 31 ASN N : -0.590: 0
: 2637:M 121 ASN ND2 :M 120 MET CE : -0.535: 0
: 2637:M 34 TYR CD2 :M 59 LEU 2HD1 : -0.493: 0
: 2637:M 124 TYR CB :M 120 MET O : -0.488: 0
: 2637:M 120 MET SD :M 121 ASN N : -0.485: 0
: 2637:M 34 TYR CB :M 30 MET O : -0.472: 0
: 2637:M 38 VAL 2HG2 :M 56 ILE 2HD1 : -0.471: 0
: 2637:M 121 ASN CG :M 31 ASN 1HD2 : -0.457: 0
: 2637:M 30 MET O :M 34 TYR N : -0.449: 0
: 2637:M 30 MET CG :M 31 ASN N : -0.433: 0
: 2637:M 31 ASN 1HD2 :M 121 ASN ND2 : -0.414: 0
: 2637:M 120 MET CE :M 121 ASN 1HD2 : -0.409: 0
: 2637:M 49 ASN H :M 49 ASN 2HD2 : -0.686: 0
: 2637:M 49 ASN N :M 49 ASN 2HD2 : -0.664: 0
: 2637:M 49 ASN N :M 49 ASN ND2 : -0.435: 0
: 2637:M 149 LEU 3HD1 :M 149 LEU O : -0.680: 0
: 2637:M 149 LEU C :M 149 LEU 3HD1 : -0.446: 0
: 2637:M 163 LEU 3HD2 :M 162 LYS C : -0.633: 0
: 2637:M 163 LEU O :M 165 HIS N : -0.590: 0
: 2637:M 165 HIS NE2 :M 166 HIS ND1 : -0.558: 0
: 2637:M 164 GLU N :M 162 LYS O : -0.529: 0
: 2637:M 163 LEU O :M 164 GLU C : -0.496: 0
: 2637:M 165 HIS CG :M 166 HIS N : -0.483: 0
: 2637:M 164 GLU N :M 162 LYS C : -0.457: 0
: 2637:M 163 LEU C :M 162 LYS O : -0.436: 0
: 2637:M 162 LYS O :M 163 LEU HG : -0.416: 0
: 2637:M 42 VAL O :M 42 VAL 2HG1 : -0.622: 0
: 2637:M 42 VAL O :M 42 VAL CG1 : -0.433: 0
: 2637:M 153 TYR CE2 :M 35 ARG NH2 : -0.608: 0
: 2637:M 153 TYR CZ :M 35 ARG NH2 : -0.586: 0
: 2637:M 98 PRO CD :M 97 VAL N : -0.576: 0
: 2637:M 97 VAL C :M 99 TYR N : -0.402: 0
: 2637:M 27 ILE C :M 29 ALA H : -0.557: 0
: 2637:M 27 ILE C :M 29 ALA N : -0.415: 0
: 2637:M 117 ILE C :M 119 ALA H : -0.517: 0
: 2637:M 117 ILE 1HD1 :M 116 LYS NZ : -0.451: 0
: 2637:M 119 ALA N :M 117 ILE C : -0.417: 0
: 2637:M 148 HIS ND1 :M 148 HIS C : -0.492: 0
: 2637:M 41 LEU 2HD2 :M 55 ARG NH1 : -0.492: 0
: 2637:M 151 GLU OE1 :M 148 HIS CD2 : -0.443: 0
: 2637:M 41 LEU C :M 43 GLN H : -0.400: 0
: 2637:M 94 LEU N :M 94 LEU 2HD1 : -0.486: 0
: 2637:M 94 LEU N :M 94 LEU CD1 : -0.423: 0
: 2637:M 93 GLU O :M 92 SER O : -0.485: 0
: 2637:M 95 PHE N :M 93 GLU C : -0.460: 0
: 2637:M 95 PHE H :M 93 GLU C : -0.443: 0
: 2637:M 6 SER OG :M 7 VAL N : -0.480: 0
: 2637:M 9 TYR N :M 7 VAL O : -0.465: 0
: 2637:M 8 PRO 1HD :M 7 VAL N : -0.440: 0
: 2637:M 10 PHE CD2 :M 33 TYR OH : -0.427: 0
: 2637:M 7 VAL O :M 8 PRO C : -0.419: 0
: 2637:M 7 VAL O :M 10 PHE N : -0.412: 0
: 2637:M 14 LEU 1HD2 :M 33 TYR CD1 : -0.410: 0
: 2637:M 18 ILE CG1 :M 19 GLU N : -0.479: 0
: 2637:M 161 SER N :M 160 GLU OE1 : -0.477: 0
: 2637:M 160 GLU O :M 161 SER CB : -0.419: 0
: 2637:M 160 GLU CD :M 161 SER N : -0.405: 0
: 2637:M 143 LEU 3HD2 :M 143 LEU O : -0.469: 0
: 2637:M 110 MET CB :M 106 GLN O : -0.453: 0
: 2637:M 106 GLN O :M 110 MET N : -0.414: 0
: 2637:M 118 HIS O :M 118 HIS ND1 : -0.451: 0
: 2637:M 167 HIS O :M 168 HIS CG : -0.448: 0
: 2637:M 168 HIS CG :M 167 HIS C : -0.418: 0
: 2637:M 44 ASP O :M 46 LEU N : -0.440: 0
: 2637:M 45 GLN O :M 46 LEU C : -0.440: 0
: 2637:M 20 MET CB :M 16 GLN O : -0.437: 0
: 2637:M 100 PHE CD2 :M 123 TYR OH : -0.427: 0
: 2637:M 145 ARG N :M 144 LYS CG : -0.423: 0
: 2637:M 48 LYS 1HG :M 50 ALA H : -0.418: 0
: 2637:M 79 HIS CG :M 78 HIS O : -0.417: 0
: 2637:M 136 LEU N :M 135 GLN CG : -0.414: 0
: 2637:M 109 GLU N :M 108 ILE CG1 : -0.406: 0
: 2637:M 109 GLU N :M 108 ILE 2HG1 : -0.405: 0
: 2637:M 28 HIS O :M 28 HIS CG : -0.404: 0
: 2637:M 150 ASP O :M 154 ASN OD1 : -0.403: 0
#sum2 ::29.20 clashscore : 29.20 clashscore B<40
#summary::2637 atoms:2637 atoms B<40:296894 potential dots:18560.0 A^2:77 bumps:77 bumps B<40:564.5 score
List of bad contacts calculated by MAGE for model $num_n
/farm/software/bin/probe
: 2637:M 8 PRO CD :M 7 VAL N : -0.653: 0
: 2637:M 125 ARG NH2 :M 63 TYR CD1 : -0.605: 0
: 2637:M 125 ARG NH2 :M 63 TYR CG : -0.548: 0
: 2637:M 63 TYR C :M 63 TYR CD1 : -0.530: 0
: 2637:M 60 ASP O :M 63 TYR CD2 : -0.471: 0
: 2637:M 63 TYR CG :M 64 ASN N : -0.468: 0
: 2637:M 129 SER N :M 128 VAL 3HG2 : -0.598: 0
: 2637:M 132 VAL 3HG2 :M 128 VAL O : -0.564: 0
: 2637:M 132 VAL O :M 132 VAL 2HG1 : -0.526: 0
: 2637:M 117 ILE C :M 119 ALA H : -0.491: 0
: 2637:M 129 SER N :M 128 VAL CG2 : -0.471: 0
: 2637:M 128 VAL O :M 132 VAL CG2 : -0.467: 0
: 2637:M 132 VAL O :M 132 VAL CG1 : -0.461: 0
: 2637:M 124 TYR CB :M 120 MET O : -0.457: 0
: 2637:M 128 VAL 2HG2 :M 124 TYR O : -0.423: 0
: 2637:M 120 MET CG :M 117 ILE O : -0.407: 0
: 2637:M 161 SER O :M 163 LEU N : -0.587: 0
: 2637:M 163 LEU H :M 161 SER C : -0.583: 0
: 2637:M 161 SER C :M 163 LEU N : -0.440: 0
: 2637:M 73 LEU N :M 71 SER O : -0.567: 0
: 2637:M 27 ILE 1HD1 :M 67 LYS 1HD : -0.556: 0
: 2637:M 70 GLU N :M 66 VAL O : -0.482: 0
: 2637:M 27 ILE C :M 29 ALA H : -0.439: 0
: 2637:M 70 GLU 2HG :M 27 ILE 2HD1 : -0.436: 0
: 2637:M 70 GLU O :M 69 GLY O : -0.428: 0
: 2637:M 4 LEU 2HD2 :M 4 LEU N : -0.532: 0
: 2637:M 4 LEU CD2 :M 4 LEU N : -0.520: 0
: 2637:M 4 LEU 1HD1 :M 9 TYR CE1 : -0.463: 0
: 2637:M 9 TYR CD1 :M 4 LEU CD1 : -0.451: 0
: 2637:M 4 LEU CD1 :M 9 TYR CE1 : -0.444: 0
: 2637:M 4 LEU H :M 4 LEU 2HD2 : -0.419: 0
: 2637:M 97 VAL O :M 99 TYR N : -0.528: 0
: 2637:M 98 PRO 1HD :M 97 VAL N : -0.481: 0
: 2637:M 98 PRO C :M 97 VAL O : -0.415: 0
: 2637:M 99 TYR N :M 97 VAL C : -0.406: 0
: 2637:M 41 LEU 3HD2 :M 41 LEU O : -0.517: 0
: 2637:M 39 SER N :M 38 VAL 3HG1 : -0.495: 0
: 2637:M 34 TYR CB :M 30 MET O : -0.457: 0
: 2637:M 62 ALA 3HB :M 59 LEU O : -0.443: 0
: 2637:M 59 LEU 3HD2 :M 34 TYR CE2 : -0.440: 0
: 2637:M 34 TYR O :M 38 VAL 2HG1 : -0.416: 0
: 2637:M 30 MET O :M 34 TYR N : -0.412: 0
: 2637:M 94 LEU C :M 94 LEU 3HD2 : -0.494: 0
: 2637:M 42 VAL O :M 42 VAL 2HG1 : -0.483: 0
: 2637:M 148 HIS CE1 :M 151 GLU OE1 : -0.479: 0
: 2637:M 148 HIS NE2 :M 151 GLU OE1 : -0.415: 0
: 2637:M 5 PHE H :M 3 GLU C : -0.471: 0
: 2637:M 5 PHE N :M 3 GLU C : -0.454: 0
: 2637:M 127 VAL O :M 131 LEU N : -0.470: 0
: 2637:M 131 LEU CB :M 127 VAL O : -0.449: 0
: 2637:M 111 ASN ND2 :M 107 HIS O : -0.469: 0
: 2637:M 103 ASN O :M 107 HIS CD2 : -0.414: 0
: 2637:M 20 MET O :M 21 ASN CB : -0.467: 0
: 2637:M 33 TYR CD2 :M 14 LEU 1HD2 : -0.467: 0
: 2637:M 21 ASN 1HD2 :M 24 GLU C : -0.463: 0
: 2637:M 21 ASN OD1 :M 25 ASP N : -0.443: 0
: 2637:M 21 ASN CA :M 17 HIS O : -0.434: 0
: 2637:M 53 LEU 3HD1 :M 53 LEU O : -0.442: 0
: 2637:M 100 PHE CD2 :M 123 TYR OH : -0.438: 0
: 2637:M 123 TYR C :M 123 TYR CD1 : -0.424: 0
: 2637:M 58 HIS CE1 :M 54 LYS O : -0.435: 0
: 2637:M 166 HIS O :M 168 HIS N : -0.435: 0
: 2637:M 54 LYS 2HG :M 55 ARG N : -0.409: 0
: 2637:M 164 GLU O :M 166 HIS N : -0.403: 0
: 2637:M 144 LYS CG :M 141 VAL O : -0.434: 0
: 2637:M 110 MET CB :M 106 GLN O : -0.422: 0
: 2637:M 28 HIS NE2 :M 31 ASN ND2 : -0.417: 0
: 2637:M 91 MET C :M 93 GLU N : -0.408: 0
: 2637:M 170 HIS ND1 :M 170 HIS N : -0.400: 0
#sum2 ::26.17 clashscore : 26.17 clashscore B<40
#summary::2637 atoms:2637 atoms B<40:296465 potential dots:18530.0 A^2:69 bumps:69 bumps B<40:543.2 score
List of bad contacts calculated by MAGE for model $num_n
/farm/software/bin/probe
: 2637:M 8 PRO CD :M 7 VAL N : -0.635: 0
: 2637:M 144 LYS 2HZ :M 140 ALA C : -0.627: 0
: 2637:M 144 LYS O :M 148 HIS CD2 : -0.464: 0
: 2637:M 148 HIS O :M 151 GLU 1HG : -0.417: 0
: 2637:M 129 SER N :M 128 VAL 3HG2 : -0.601: 0
: 2637:M 117 ILE C :M 119 ALA H : -0.594: 0
: 2637:M 120 MET SD :M 121 ASN N : -0.565: 0
: 2637:M 117 ILE O :M 120 MET SD : -0.511: 0
: 2637:M 120 MET O :M 124 TYR N : -0.507: 0
: 2637:M 124 TYR CB :M 120 MET O : -0.487: 0
: 2637:M 128 VAL CG2 :M 129 SER N : -0.480: 0
: 2637:M 120 MET CE :M 121 ASN OD1 : -0.443: 0
: 2637:M 128 VAL 2HG2 :M 124 TYR O : -0.430: 0
: 2637:M 119 ALA N :M 117 ILE C : -0.429: 0
: 2637:M 132 VAL 3HG2 :M 128 VAL O : -0.420: 0
: 2637:M 124 TYR CD2 :M 149 LEU 2HD1 : -0.406: 0
: 2637:M 124 TYR CD1 :M 124 TYR C : -0.401: 0
: 2637:M 50 ALA 3HB :M 48 LYS 2HE : -0.581: 0
: 2637:M 48 LYS 1HG :M 50 ALA H : -0.462: 0
: 2637:M 16 GLN 2HE2 :M 22 GLN 2HE2 : -0.534: 0
: 2637:M 104 LEU 1HD2 :M 123 TYR CD1 : -0.525: 0
: 2637:M 100 PHE CD2 :M 123 TYR OH : -0.461: 0
: 2637:M 71 SER OG :M 72 LYS N : -0.509: 0
: 2637:M 70 GLU OE1 :M 26 LYS NZ : -0.481: 0
: 2637:M 70 GLU O :M 72 LYS N : -0.437: 0
: 2637:M 70 GLU O :M 71 SER OG : -0.400: 0
: 2637:M 44 ASP CB :M 43 GLN O : -0.508: 0
: 2637:M 44 ASP O :M 45 GLN 2HB : -0.435: 0
: 2637:M 30 MET O :M 34 TYR N : -0.507: 0
: 2637:M 34 TYR CB :M 30 MET O : -0.487: 0
: 2637:M 27 ILE C :M 29 ALA H : -0.501: 0
: 2637:M 94 LEU CD1 :M 99 TYR CD1 : -0.498: 0
: 2637:M 98 PRO 1HD :M 97 VAL N : -0.490: 0
: 2637:M 97 VAL O :M 99 TYR N : -0.472: 0
: 2637:M 96 SER OG :M 97 VAL N : -0.457: 0
: 2637:M 103 ASN ND2 :M 99 TYR O : -0.451: 0
: 2637:M 94 LEU N :M 94 LEU CD2 : -0.440: 0
: 2637:M 94 LEU N :M 94 LEU 2HD2 : -0.430: 0
: 2637:M 97 VAL O :M 98 PRO C : -0.415: 0
: 2637:M 94 LEU H :M 94 LEU CD2 : -0.410: 0
: 2637:M 99 TYR N :M 97 VAL C : -0.404: 0
: 2637:M 163 LEU O :M 165 HIS N : -0.488: 0
: 2637:M 78 HIS O :M 76 HIS CD2 : -0.474: 0
: 2637:M 118 HIS CD2 :M 118 HIS O : -0.472: 0
: 2637:M 118 HIS O :M 118 HIS CG : -0.449: 0
: 2637:M 125 ARG NH2 :M 63 TYR CE2 : -0.461: 0
: 2637:M 125 ARG NH2 :M 63 TYR CD2 : -0.438: 0
: 2637:M 63 TYR CD1 :M 63 TYR C : -0.410: 0
: 2637:M 130 THR O :M 134 ASP N : -0.458: 0
: 2637:M 127 VAL O :M 130 THR N : -0.443: 0
: 2637:M 53 LEU N :M 52 VAL 3HG2 : -0.437: 0
: 2637:M 53 LEU CD2 :M 139 ASN ND2 : -0.408: 0
: 2637:M 10 PHE CD2 :M 33 TYR OH : -0.432: 0
: 2637:M 106 GLN O :M 110 MET N : -0.428: 0
: 2637:M 109 GLU CB :M 105 LYS O : -0.422: 0
: 2637:M 109 GLU N :M 108 ILE CG1 : -0.411: 0
: 2637:M 111 ASN OD1 :M 115 ASP N : -0.416: 0
: 2637:M 137 THR 2HG2 :M 137 THR H : -0.415: 0
: 2637:M 80 HIS N :M 79 HIS CG : -0.414: 0
: 2637:M 2 SER OG :M 3 GLU N : -0.411: 0
: 2637:M 23 SER H :M 21 ASN C : -0.408: 0
: 2637:M 153 TYR CD1 :M 153 TYR C : -0.404: 0
: 2637:M 58 HIS CG :M 54 LYS O : -0.400: 0
#sum2 ::23.89 clashscore : 23.89 clashscore B<40
#summary::2637 atoms:2637 atoms B<40:295863 potential dots:18490.0 A^2:63 bumps:63 bumps B<40:606.5 score
List of bad contacts calculated by MAGE for model $num_n
/farm/software/bin/probe
: 2637:M 34 TYR O :M 38 VAL 3HG2 : -0.660: 0
: 2637:M 30 MET SD :M 31 ASN N : -0.572: 0
: 2637:M 30 MET O :M 34 TYR N : -0.531: 0
: 2637:M 34 TYR CB :M 30 MET O : -0.465: 0
: 2637:M 34 TYR O :M 38 VAL CG2 : -0.448: 0
: 2637:M 31 ASN N :M 30 MET CG : -0.414: 0
: 2637:M 8 PRO 1HD :M 7 VAL N : -0.659: 0
: 2637:M 7 VAL N :M 8 PRO CD : -0.458: 0
: 2637:M 124 TYR O :M 128 VAL 3HG2 : -0.650: 0
: 2637:M 120 MET SD :M 121 ASN N : -0.543: 0
: 2637:M 121 ASN ND2 :M 120 MET CE : -0.505: 0
: 2637:M 120 MET O :M 124 TYR N : -0.502: 0
: 2637:M 124 TYR CB :M 120 MET O : -0.476: 0
: 2637:M 121 ASN N :M 120 MET CG : -0.433: 0
: 2637:M 98 PRO 1HD :M 97 VAL N : -0.631: 0
: 2637:M 97 VAL O :M 99 TYR N : -0.437: 0
: 2637:M 97 VAL N :M 98 PRO CD : -0.433: 0
: 2637:M 99 TYR N :M 97 VAL C : -0.428: 0
: 2637:M 145 ARG 2HH1 :M 135 GLN NE2 : -0.629: 0
: 2637:M 145 ARG NH2 :M 131 LEU CD2 : -0.491: 0
: 2637:M 144 LYS CG :M 145 ARG N : -0.487: 0
: 2637:M 145 ARG CZ :M 131 LEU CD2 : -0.484: 0
: 2637:M 135 GLN O :M 136 LEU C : -0.432: 0
: 2637:M 104 LEU N :M 104 LEU 2HD1 : -0.617: 0
: 2637:M 104 LEU N :M 104 LEU CD1 : -0.519: 0
: 2637:M 100 PHE CD2 :M 123 TYR OH : -0.465: 0
: 2637:M 101 ILE C :M 103 ASN N : -0.432: 0
: 2637:M 104 LEU 1HD1 :M 123 TYR CE1 : -0.429: 0
: 2637:M 103 ASN O :M 107 HIS CD2 : -0.416: 0
: 2637:M 101 ILE O :M 104 LEU N : -0.409: 0
: 2637:M 104 LEU 1HD1 :M 123 TYR CD1 : -0.403: 0
: 2637:M 27 ILE 1HD1 :M 67 LYS 1HD : -0.568: 0
: 2637:M 27 ILE C :M 29 ALA H : -0.551: 0
: 2637:M 27 ILE C :M 29 ALA N : -0.415: 0
: 2637:M 43 GLN O :M 44 ASP C : -0.559: 0
: 2637:M 140 ALA N :M 139 ASN ND2 : -0.536: 0
: 2637:M 49 ASN CG :M 139 ASN ND2 : -0.448: 0
: 2637:M 49 ASN O :M 53 LEU CB : -0.422: 0
: 2637:M 53 LEU N :M 52 VAL 3HG2 : -0.401: 0
: 2637:M 45 GLN O :M 46 LEU C : -0.525: 0
: 2637:M 47 THR N :M 45 GLN O : -0.514: 0
: 2637:M 47 THR N :M 45 GLN C : -0.417: 0
: 2637:M 73 LEU 2HD1 :M 73 LEU N : -0.524: 0
: 2637:M 73 LEU CD1 :M 73 LEU N : -0.473: 0
: 2637:M 158 ARG CZ :M 154 ASN ND2 : -0.522: 0
: 2637:M 91 MET SD :M 91 MET N : -0.519: 0
: 2637:M 54 LYS CG :M 55 ARG N : -0.516: 0
: 2637:M 55 ARG 1HG :M 41 LEU 2HD2 : -0.485: 0
: 2637:M 148 HIS CE1 :M 151 GLU OE2 : -0.496: 0
: 2637:M 152 ALA N :M 151 GLU CG : -0.407: 0
: 2637:M 166 HIS ND1 :M 166 HIS N : -0.491: 0
: 2637:M 164 GLU C :M 166 HIS N : -0.437: 0
: 2637:M 164 GLU C :M 166 HIS H : -0.413: 0
: 2637:M 77 HIS N :M 76 HIS CG : -0.487: 0
: 2637:M 70 GLU OE1 :M 26 LYS NZ : -0.486: 0
: 2637:M 160 GLU OE1 :M 116 LYS NZ : -0.481: 0
: 2637:M 160 GLU CD :M 161 SER N : -0.451: 0
: 2637:M 159 GLY O :M 162 LYS N : -0.443: 0
: 2637:M 160 GLU C :M 159 GLY O : -0.421: 0
: 2637:M 161 SER N :M 160 GLU CG : -0.407: 0
: 2637:M 20 MET CB :M 16 GLN O : -0.472: 0
: 2637:M 93 GLU CD :M 93 GLU N : -0.461: 0
: 2637:M 10 PHE CD2 :M 33 TYR OH : -0.459: 0
: 2637:M 80 HIS CG :M 79 HIS O : -0.457: 0
: 2637:M 79 HIS C :M 80 HIS CG : -0.410: 0
: 2637:M 165 HIS CG :M 165 HIS O : -0.455: 0
: 2637:M 18 ILE CG1 :M 19 GLU N : -0.451: 0
: 2637:M 19 GLU CB :M 15 LYS O : -0.411: 0
: 2637:M 117 ILE C :M 119 ALA H : -0.434: 0
: 2637:M 110 MET CB :M 106 GLN O : -0.428: 0
: 2637:M 138 LYS O :M 142 VAL 3HG2 : -0.414: 0
: 2637:M 109 GLU CB :M 105 LYS O : -0.412: 0
: 2637:M 109 GLU N :M 108 ILE CG1 : -0.410: 0
: 2637:M 4 LEU 3HD2 :M 4 LEU O : -0.405: 0
: 2637:M 137 THR 2HG2 :M 137 THR H : -0.404: 0
#sum2 ::28.44 clashscore : 28.44 clashscore B<40
#summary::2637 atoms:2637 atoms B<40:296623 potential dots:18540.0 A^2:75 bumps:75 bumps B<40:561.7 score
List of bad contacts calculated by MAGE for model $num_n
/farm/software/bin/probe
: 2637:M 134 ASP O :M 135 GLN O : -0.901: 0
: 2637:M 135 GLN 1HE2 :M 145 ARG HE : -0.613: 0
: 2637:M 145 ARG HE :M 135 GLN NE2 : -0.589: 0
: 2637:M 111 ASN 1HD2 :M 115 ASP N : -0.745: 0
: 2637:M 111 ASN 1HD2 :M 115 ASP CA : -0.678: 0
: 2637:M 110 MET SD :M 110 MET N : -0.530: 0
: 2637:M 114 GLU OE2 :M 122 SER CB : -0.493: 0
: 2637:M 111 ASN 1HD2 :M 114 GLU C : -0.465: 0
: 2637:M 111 ASN ND2 :M 115 ASP N : -0.450: 0
: 2637:M 106 GLN O :M 110 MET N : -0.424: 0
: 2637:M 110 MET O :M 111 ASN CB : -0.409: 0
: 2637:M 124 TYR O :M 128 VAL 3HG2 : -0.726: 0
: 2637:M 120 MET O :M 124 TYR N : -0.533: 0
: 2637:M 120 MET C :M 120 MET SD : -0.528: 0
: 2637:M 128 VAL N :M 127 VAL 3HG2 : -0.516: 0
: 2637:M 124 TYR 1HB :M 120 MET O : -0.435: 0
: 2637:M 128 VAL N :M 127 VAL CG2 : -0.428: 0
: 2637:M 124 TYR O :M 128 VAL CG2 : -0.425: 0
: 2637:M 8 PRO CD :M 7 VAL N : -0.627: 0
: 2637:M 7 VAL 3HG2 :M 7 VAL H : -0.423: 0
: 2637:M 42 VAL O :M 42 VAL 2HG1 : -0.618: 0
: 2637:M 42 VAL O :M 42 VAL CG1 : -0.433: 0
: 2637:M 140 ALA 2HB :M 49 ASN ND2 : -0.574: 0
: 2637:M 53 LEU N :M 52 VAL 3HG2 : -0.562: 0
: 2637:M 49 ASN O :M 53 LEU CB : -0.471: 0
: 2637:M 139 ASN OD1 :M 140 ALA N : -0.468: 0
: 2637:M 53 LEU N :M 52 VAL CG2 : -0.467: 0
: 2637:M 30 MET SD :M 31 ASN N : -0.573: 0
: 2637:M 125 ARG 2HH2 :M 31 ASN CG : -0.567: 0
: 2637:M 30 MET O :M 34 TYR N : -0.536: 0
: 2637:M 30 MET CG :M 31 ASN N : -0.459: 0
: 2637:M 34 TYR CB :M 30 MET O : -0.408: 0
: 2637:M 27 ILE C :M 29 ALA H : -0.566: 0
: 2637:M 27 ILE C :M 29 ALA N : -0.444: 0
: 2637:M 70 GLU OE2 :M 27 ILE 1HD1 : -0.442: 0
: 2637:M 70 GLU OE1 :M 70 GLU C : -0.438: 0
: 2637:M 70 GLU OE1 :M 26 LYS NZ : -0.419: 0
: 2637:M 41 LEU 3HD1 :M 41 LEU O : -0.549: 0
: 2637:M 97 VAL O :M 99 TYR N : -0.533: 0
: 2637:M 98 PRO 1HD :M 97 VAL N : -0.520: 0
: 2637:M 96 SER OG :M 97 VAL N : -0.452: 0
: 2637:M 98 PRO C :M 97 VAL O : -0.431: 0
: 2637:M 99 TYR N :M 97 VAL C : -0.425: 0
: 2637:M 71 SER OG :M 72 LYS N : -0.480: 0
: 2637:M 33 TYR CD1 :M 33 TYR C : -0.479: 0
: 2637:M 10 PHE CD1 :M 33 TYR OH : -0.448: 0
: 2637:M 148 HIS ND1 :M 144 LYS O : -0.474: 0
: 2637:M 150 ASP N :M 148 HIS C : -0.444: 0
: 2637:M 58 HIS ND1 :M 54 LYS O : -0.472: 0
: 2637:M 58 HIS CG :M 54 LYS O : -0.469: 0
: 2637:M 55 ARG N :M 54 LYS CG : -0.406: 0
: 2637:M 166 HIS O :M 168 HIS CE1 : -0.461: 0
: 2637:M 3 GLU CD :M 3 GLU N : -0.455: 0
: 2637:M 3 GLU OE1 :M 3 GLU N : -0.443: 0
: 2637:M 151 GLU CG :M 152 ALA N : -0.451: 0
: 2637:M 75 HIS O :M 76 HIS O : -0.446: 0
: 2637:M 75 HIS O :M 75 HIS ND1 : -0.410: 0
: 2637:M 35 ARG C :M 35 ARG CD : -0.445: 0
: 2637:M 94 LEU C :M 94 LEU 3HD2 : -0.444: 0
: 2637:M 94 LEU C :M 94 LEU CD2 : -0.409: 0
: 2637:M 65 LYS O :M 69 GLY N : -0.443: 0
: 2637:M 65 LYS CG :M 61 GLU O : -0.402: 0
: 2637:M 123 TYR C :M 123 TYR CD1 : -0.441: 0
: 2637:M 123 TYR O :M 123 TYR CD1 : -0.409: 0
: 2637:M 1 MET SD :M 1 MET C : -0.438: 0
: 2637:M 154 ASN O :M 158 ARG CB : -0.434: 0
: 2637:M 44 ASP CB :M 40 THR O : -0.422: 0
: 2637:M 165 HIS O :M 167 HIS N : -0.418: 0
: 2637:M 153 TYR CD1 :M 153 TYR C : -0.415: 0
: 2637:M 18 ILE 2HG1 :M 19 GLU N : -0.405: 0
: 2637:M 19 GLU N :M 18 ILE CG1 : -0.401: 0
: 2637:M 45 GLN 1HG :M 47 THR H : -0.403: 0
: 2637:M 105 LYS O :M 109 GLU CB : -0.400: 0
#sum2 ::27.68 clashscore : 27.68 clashscore B<40
#summary::2637 atoms:2637 atoms B<40:296465 potential dots:18530.0 A^2:73 bumps:73 bumps B<40:510 score
List of bad contacts calculated by MAGE for model $num_n
/farm/software/bin/probe
: 2637:M 44 ASP O :M 45 GLN O : -1.129: 0
: 2637:M 45 GLN O :M 44 ASP C : -0.534: 0
: 2637:M 44 ASP O :M 45 GLN C : -0.529: 0
: 2637:M 40 THR O :M 44 ASP N : -0.435: 0
: 2637:M 4 LEU H :M 4 LEU 2HD2 : -0.903: 0
: 2637:M 4 LEU 2HD2 :M 4 LEU N : -0.711: 0
: 2637:M 4 LEU H :M 4 LEU CD2 : -0.620: 0
: 2637:M 4 LEU 2HD1 :M 9 TYR CD2 : -0.546: 0
: 2637:M 9 TYR CE2 :M 4 LEU 2HD1 : -0.449: 0
: 2637:M 132 VAL O :M 132 VAL 2HG1 : -0.677: 0
: 2637:M 132 VAL O :M 132 VAL CG1 : -0.483: 0
: 2637:M 52 VAL N :M 51 VAL 3HG2 : -0.663: 0
: 2637:M 48 LYS 1HG :M 50 ALA H : -0.584: 0
: 2637:M 51 VAL 2HG2 :M 48 LYS O : -0.571: 0
: 2637:M 52 VAL N :M 51 VAL CG2 : -0.567: 0
: 2637:M 52 VAL H :M 51 VAL CG2 : -0.493: 0
: 2637:M 48 LYS N :M 47 THR 3HG2 : -0.439: 0
: 2637:M 31 ASN ND2 :M 125 ARG HE : -0.657: 0
: 2637:M 121 ASN 1HD2 :M 31 ASN ND2 : -0.590: 0
: 2637:M 120 MET SD :M 121 ASN N : -0.576: 0
: 2637:M 30 MET SD :M 31 ASN N : -0.574: 0
: 2637:M 129 SER N :M 128 VAL 3HG2 : -0.559: 0
: 2637:M 128 VAL CG2 :M 129 SER N : -0.490: 0
: 2637:M 124 TYR CB :M 120 MET O : -0.487: 0
: 2637:M 34 TYR O :M 38 VAL 3HG2 : -0.481: 0
: 2637:M 120 MET CG :M 121 ASN N : -0.480: 0
: 2637:M 125 ARG HE :M 31 ASN 2HD2 : -0.479: 0
: 2637:M 121 ASN OD1 :M 35 ARG NH1 : -0.450: 0
: 2637:M 34 TYR CB :M 30 MET O : -0.448: 0
: 2637:M 31 ASN N :M 30 MET CG : -0.439: 0
: 2637:M 128 VAL 2HG2 :M 124 TYR O : -0.438: 0
: 2637:M 124 TYR CD1 :M 124 TYR C : -0.417: 0
: 2637:M 30 MET O :M 34 TYR N : -0.413: 0
: 2637:M 104 LEU N :M 104 LEU 2HD1 : -0.644: 0
: 2637:M 123 TYR CE2 :M 104 LEU 1HD1 : -0.619: 0
: 2637:M 104 LEU N :M 104 LEU CD1 : -0.507: 0
: 2637:M 146 ILE O :M 149 LEU N : -0.425: 0
: 2637:M 123 TYR HE2 :M 149 LEU 1HD1 : -0.422: 0
: 2637:M 149 LEU 1HD2 :M 100 PHE CD2 : -0.414: 0
: 2637:M 98 PRO CD :M 97 VAL N : -0.640: 0
: 2637:M 8 PRO CD :M 7 VAL N : -0.637: 0
: 2637:M 27 ILE C :M 29 ALA H : -0.578: 0
: 2637:M 27 ILE 1HD1 :M 67 LYS 1HD : -0.562: 0
: 2637:M 70 GLU OE2 :M 27 ILE 2HD1 : -0.535: 0
: 2637:M 27 ILE C :M 29 ALA N : -0.445: 0
: 2637:M 117 ILE C :M 119 ALA H : -0.553: 0
: 2637:M 117 ILE C :M 119 ALA N : -0.455: 0
: 2637:M 117 ILE 1HD1 :M 116 LYS NZ : -0.429: 0
: 2637:M 59 LEU O :M 59 LEU 3HD2 : -0.519: 0
: 2637:M 91 MET SD :M 91 MET N : -0.517: 0
: 2637:M 92 SER C :M 94 LEU H : -0.453: 0
: 2637:M 92 SER O :M 91 MET O : -0.423: 0
: 2637:M 153 TYR CD1 :M 153 TYR C : -0.480: 0
: 2637:M 78 HIS N :M 77 HIS CG : -0.476: 0
: 2637:M 11 ILE 3HD1 :M 65 LYS 2HD : -0.470: 0
: 2637:M 11 ILE 3HD1 :M 65 LYS CD : -0.419: 0
: 2637:M 11 ILE O :M 14 LEU N : -0.402: 0
: 2637:M 139 ASN O :M 143 LEU 1HB : -0.464: 0
: 2637:M 143 LEU C :M 143 LEU CD1 : -0.405: 0
: 2637:M 143 LEU O :M 143 LEU 3HD1 : -0.402: 0
: 2637:M 143 LEU C :M 143 LEU 3HD1 : -0.400: 0
: 2637:M 10 PHE CD2 :M 33 TYR OH : -0.454: 0
: 2637:M 54 LYS O :M 58 HIS CD2 : -0.454: 0
: 2637:M 79 HIS ND1 :M 80 HIS O : -0.451: 0
: 2637:M 53 LEU HA :M 56 ILE 2HD1 : -0.440: 0
: 2637:M 93 GLU C :M 95 PHE N : -0.435: 0
: 2637:M 109 GLU N :M 108 ILE CG1 : -0.430: 0
: 2637:M 109 GLU CB :M 105 LYS O : -0.423: 0
: 2637:M 154 ASN O :M 158 ARG CG : -0.429: 0
: 2637:M 62 ALA N :M 61 GLU CG : -0.421: 0
: 2637:M 18 ILE 2HG1 :M 19 GLU N : -0.414: 0
: 2637:M 19 GLU CB :M 15 LYS O : -0.402: 0
: 2637:M 165 HIS H :M 163 LEU 3HD2 : -0.412: 0
: 2637:M 163 LEU CD2 :M 165 HIS H : -0.406: 0
: 2637:M 130 THR O :M 134 ASP CB : -0.405: 0
: 2637:M 112 GLN O :M 113 SER OG : -0.404: 0
#sum2 ::28.82 clashscore : 28.82 clashscore B<40
#summary::2637 atoms:2637 atoms B<40:296718 potential dots:18540.0 A^2:76 bumps:76 bumps B<40:502.3 score
List of bad contacts calculated by MAGE for model $num_n
/farm/software/bin/probe
: 2637:M 132 VAL O :M 132 VAL 2HG1 : -0.689: 0
: 2637:M 132 VAL O :M 132 VAL CG1 : -0.501: 0
: 2637:M 27 ILE C :M 29 ALA H : -0.673: 0
: 2637:M 27 ILE O :M 30 MET SD : -0.557: 0
: 2637:M 30 MET O :M 34 TYR N : -0.551: 0
: 2637:M 30 MET SD :M 31 ASN N : -0.478: 0
: 2637:M 27 ILE 3HG2 :M 28 HIS N : -0.471: 0
: 2637:M 34 TYR CB :M 30 MET O : -0.471: 0
: 2637:M 27 ILE C :M 29 ALA N : -0.466: 0
: 2637:M 28 HIS N :M 27 ILE CG2 : -0.448: 0
: 2637:M 27 ILE O :M 29 ALA N : -0.418: 0
: 2637:M 161 SER H :M 160 GLU CG : -0.670: 0
: 2637:M 161 SER H :M 160 GLU CD : -0.570: 0
: 2637:M 117 ILE 3HD1 :M 160 GLU OE1 : -0.543: 0
: 2637:M 117 ILE 3HD1 :M 160 GLU CD : -0.463: 0
: 2637:M 117 ILE 3HG2 :M 118 HIS N : -0.457: 0
: 2637:M 160 GLU CG :M 161 SER N : -0.411: 0
: 2637:M 125 ARG NH2 :M 63 TYR CE2 : -0.665: 0
: 2637:M 73 LEU O :M 75 HIS N : -0.620: 0
: 2637:M 120 MET CG :M 121 ASN N : -0.570: 0
: 2637:M 121 ASN H :M 120 MET 1HG : -0.490: 0
: 2637:M 124 TYR CB :M 120 MET O : -0.475: 0
: 2637:M 121 ASN H :M 120 MET CG : -0.438: 0
: 2637:M 120 MET O :M 124 TYR N : -0.434: 0
: 2637:M 124 TYR 1HB :M 120 MET O : -0.411: 0
: 2637:M 14 LEU 1HD2 :M 33 TYR CD1 : -0.562: 0
: 2637:M 98 PRO CD :M 97 VAL N : -0.546: 0
: 2637:M 5 PHE O :M 6 SER CB : -0.542: 0
: 2637:M 1 MET N :M 1 MET SD : -0.523: 0
: 2637:M 6 SER O :M 9 TYR N : -0.485: 0
: 2637:M 6 SER O :M 7 VAL C : -0.480: 0
: 2637:M 8 PRO 1HD :M 7 VAL N : -0.479: 0
: 2637:M 5 PHE C :M 4 LEU O : -0.447: 0
: 2637:M 6 SER OG :M 6 SER O : -0.425: 0
: 2637:M 9 TYR OH :M 1 MET N : -0.420: 0
: 2637:M 8 PRO O :M 12 GLU CG : -0.409: 0
: 2637:M 96 SER CB :M 95 PHE O : -0.517: 0
: 2637:M 44 ASP N :M 40 THR O : -0.512: 0
: 2637:M 148 HIS CG :M 144 LYS O : -0.508: 0
: 2637:M 150 ASP OD1 :M 151 GLU N : -0.469: 0
: 2637:M 144 LYS O :M 148 HIS CD2 : -0.421: 0
: 2637:M 148 HIS C :M 150 ASP N : -0.419: 0
: 2637:M 152 ALA N :M 151 GLU CG : -0.406: 0
: 2637:M 165 HIS CE1 :M 167 HIS CE1 : -0.500: 0
: 2637:M 165 HIS O :M 166 HIS CG : -0.476: 0
: 2637:M 167 HIS ND1 :M 167 HIS N : -0.472: 0
: 2637:M 166 HIS O :M 167 HIS O : -0.443: 0
: 2637:M 166 HIS CG :M 165 HIS C : -0.412: 0
: 2637:M 80 HIS ND1 :M 80 HIS N : -0.495: 0
: 2637:M 131 LEU CB :M 127 VAL O : -0.491: 0
: 2637:M 127 VAL O :M 131 LEU N : -0.411: 0
: 2637:M 48 LYS 1HG :M 50 ALA H : -0.490: 0
: 2637:M 110 MET CB :M 106 GLN O : -0.479: 0
: 2637:M 17 HIS NE2 :M 13 ASN OD1 : -0.476: 0
: 2637:M 45 GLN 1HE2 :M 55 ARG CZ : -0.476: 0
: 2637:M 54 LYS 1HG :M 51 VAL O : -0.471: 0
: 2637:M 54 LYS CG :M 55 ARG N : -0.466: 0
: 2637:M 58 HIS CG :M 54 LYS O : -0.411: 0
: 2637:M 17 HIS CE1 :M 13 ASN OD1 : -0.402: 0
: 2637:M 60 ASP OD1 :M 61 GLU N : -0.441: 0
: 2637:M 62 ALA N :M 61 GLU CG : -0.406: 0
: 2637:M 107 HIS CG :M 103 ASN O : -0.427: 0
: 2637:M 103 ASN O :M 107 HIS CD2 : -0.411: 0
: 2637:M 22 GLN O :M 23 SER CB : -0.410: 0
: 2637:M 23 SER O :M 23 SER OG : -0.410: 0
: 2637:M 105 LYS O :M 109 GLU CB : -0.404: 0
: 2637:M 109 GLU N :M 108 ILE CG1 : -0.404: 0
: 2637:M 145 ARG N :M 145 ARG CD : -0.401: 0
#sum2 ::25.79 clashscore : 25.79 clashscore B<40
#summary::2637 atoms:2637 atoms B<40:296692 potential dots:18540.0 A^2:68 bumps:68 bumps B<40:598.2 score
List of bad contacts calculated by MAGE for model $num_n
/farm/software/bin/probe
: 2637:M 104 LEU N :M 104 LEU 2HD1 : -0.667: 0
: 2637:M 104 LEU 1HD1 :M 123 TYR CE1 : -0.614: 0
: 2637:M 104 LEU N :M 104 LEU CD1 : -0.538: 0
: 2637:M 100 PHE CE2 :M 123 TYR OH : -0.455: 0
: 2637:M 127 VAL 1HG2 :M 123 TYR CE1 : -0.406: 0
: 2637:M 57 GLN 2HE2 :M 53 LEU 2HD1 : -0.662: 0
: 2637:M 57 GLN 2HE2 :M 53 LEU CD1 : -0.572: 0
: 2637:M 53 LEU 3HD1 :M 53 LEU O : -0.474: 0
: 2637:M 52 VAL O :M 56 ILE 1HG1 : -0.460: 0
: 2637:M 49 ASN O :M 53 LEU 1HB : -0.445: 0
: 2637:M 53 LEU N :M 52 VAL 3HG2 : -0.440: 0
: 2637:M 53 LEU C :M 53 LEU 3HD1 : -0.437: 0
: 2637:M 53 LEU CD1 :M 53 LEU C : -0.430: 0
: 2637:M 27 ILE C :M 29 ALA H : -0.600: 0
: 2637:M 27 ILE 3HG2 :M 28 HIS N : -0.506: 0
: 2637:M 29 ALA 2HB :M 26 LYS O : -0.478: 0
: 2637:M 28 HIS N :M 27 ILE CG2 : -0.471: 0
: 2637:M 27 ILE C :M 29 ALA N : -0.425: 0
: 2637:M 98 PRO CD :M 97 VAL N : -0.599: 0
: 2637:M 66 VAL 1HG2 :M 30 MET SD : -0.584: 0
: 2637:M 30 MET O :M 34 TYR N : -0.527: 0
: 2637:M 34 TYR CB :M 30 MET O : -0.451: 0
: 2637:M 120 MET SD :M 121 ASN N : -0.572: 0
: 2637:M 129 SER N :M 128 VAL 3HG2 : -0.527: 0
: 2637:M 124 TYR CB :M 120 MET O : -0.498: 0
: 2637:M 128 VAL 1HG1 :M 124 TYR CE1 : -0.447: 0
: 2637:M 120 MET O :M 124 TYR N : -0.446: 0
: 2637:M 129 SER N :M 128 VAL CG2 : -0.427: 0
: 2637:M 120 MET C :M 120 MET SD : -0.415: 0
: 2637:M 167 HIS N :M 166 HIS CG : -0.550: 0
: 2637:M 167 HIS O :M 166 HIS CD2 : -0.421: 0
: 2637:M 8 PRO CD :M 7 VAL N : -0.546: 0
: 2637:M 8 PRO 1HD :M 7 VAL N : -0.467: 0
: 2637:M 144 LYS CG :M 145 ARG N : -0.530: 0
: 2637:M 131 LEU O :M 134 ASP O : -0.504: 0
: 2637:M 134 ASP O :M 135 GLN C : -0.434: 0
: 2637:M 131 LEU 2HD2 :M 145 ARG NH1 : -0.433: 0
: 2637:M 135 GLN 1HE2 :M 145 ARG 2HH1 : -0.431: 0
: 2637:M 134 ASP O :M 135 GLN CG : -0.429: 0
: 2637:M 135 GLN C :M 137 THR H : -0.409: 0
: 2637:M 137 THR 3HG2 :M 138 LYS N : -0.407: 0
: 2637:M 72 LYS C :M 71 SER O : -0.518: 0
: 2637:M 37 VAL O :M 41 LEU CB : -0.506: 0
: 2637:M 37 VAL O :M 41 LEU N : -0.458: 0
: 2637:M 110 MET O :M 111 ASN CB : -0.494: 0
: 2637:M 111 ASN CA :M 107 HIS O : -0.471: 0
: 2637:M 20 MET O :M 21 ASN CB : -0.493: 0
: 2637:M 21 ASN CA :M 17 HIS O : -0.472: 0
: 2637:M 21 ASN OD1 :M 25 ASP N : -0.430: 0
: 2637:M 25 ASP H :M 24 GLU CG : -0.428: 0
: 2637:M 25 ASP CA :M 21 ASN OD1 : -0.408: 0
: 2637:M 25 ASP CG :M 24 GLU O : -0.400: 0
: 2637:M 62 ALA 3HB :M 59 LEU O : -0.489: 0
: 2637:M 54 LYS CG :M 55 ARG N : -0.489: 0
: 2637:M 68 ARG CG :M 64 ASN O : -0.485: 0
: 2637:M 63 TYR CE2 :M 64 ASN OD1 : -0.430: 0
: 2637:M 64 ASN CG :M 63 TYR CE2 : -0.425: 0
: 2637:M 117 ILE C :M 119 ALA H : -0.484: 0
: 2637:M 117 ILE C :M 119 ALA N : -0.405: 0
: 2637:M 160 GLU O :M 162 LYS N : -0.482: 0
: 2637:M 44 ASP O :M 45 GLN 2HB : -0.474: 0
: 2637:M 61 GLU OE1 :M 58 HIS CD2 : -0.465: 0
: 2637:M 58 HIS ND1 :M 58 HIS C : -0.419: 0
: 2637:M 73 LEU C :M 75 HIS H : -0.460: 0
: 2637:M 75 HIS N :M 73 LEU C : -0.460: 0
: 2637:M 163 LEU 3HD1 :M 163 LEU O : -0.459: 0
: 2637:M 163 LEU C :M 163 LEU CD1 : -0.424: 0
: 2637:M 163 LEU 3HD1 :M 163 LEU C : -0.420: 0
: 2637:M 33 TYR C :M 33 TYR CD1 : -0.434: 0
: 2637:M 10 PHE CE2 :M 33 TYR OH : -0.427: 0
: 2637:M 108 ILE CG1 :M 109 GLU N : -0.429: 0
: 2637:M 19 GLU CB :M 15 LYS O : -0.418: 0
: 2637:M 39 SER O :M 43 GLN CB : -0.412: 0
: 2637:M 79 HIS O :M 79 HIS CG : -0.410: 0
#sum2 ::28.06 clashscore : 28.06 clashscore B<40
#summary::2637 atoms:2637 atoms B<40:296240 potential dots:18520.0 A^2:74 bumps:74 bumps B<40:558.3 score
List of bad contacts calculated by MAGE for model $num_n
/farm/software/bin/probe
: 2637:M 141 VAL 2HG2 :M 138 LYS O : -0.736: 0
: 2637:M 140 ALA H :M 138 LYS 2HG : -0.461: 0
: 2637:M 138 LYS 2HB :M 141 VAL 3HG1 : -0.431: 0
: 2637:M 138 LYS C :M 140 ALA N : -0.429: 0
: 2637:M 138 LYS C :M 140 ALA H : -0.409: 0
: 2637:M 42 VAL O :M 42 VAL 2HG1 : -0.654: 0
: 2637:M 42 VAL O :M 42 VAL CG1 : -0.455: 0
: 2637:M 70 GLU C :M 69 GLY O : -0.651: 0
: 2637:M 71 SER H :M 70 GLU CG : -0.450: 0
: 2637:M 157 LYS 1HD :M 117 ILE 1HD1 : -0.615: 0
: 2637:M 120 MET SD :M 121 ASN N : -0.611: 0
: 2637:M 149 LEU 3HD1 :M 149 LEU O : -0.607: 0
: 2637:M 34 TYR O :M 38 VAL 3HG2 : -0.606: 0
: 2637:M 30 MET CG :M 31 ASN N : -0.563: 0
: 2637:M 124 TYR N :M 120 MET O : -0.558: 0
: 2637:M 124 TYR CD2 :M 149 LEU 2HD1 : -0.509: 0
: 2637:M 120 MET CG :M 121 ASN N : -0.498: 0
: 2637:M 31 ASN H :M 30 MET CG : -0.491: 0
: 2637:M 31 ASN H :M 30 MET 1HG : -0.467: 0
: 2637:M 121 ASN OD1 :M 31 ASN ND2 : -0.463: 0
: 2637:M 120 MET 2HG :M 121 ASN H : -0.437: 0
: 2637:M 34 TYR 1HB :M 30 MET O : -0.429: 0
: 2637:M 124 TYR 1HB :M 120 MET O : -0.418: 0
: 2637:M 121 ASN H :M 120 MET CG : -0.416: 0
: 2637:M 98 PRO CD :M 97 VAL N : -0.582: 0
: 2637:M 145 ARG CZ :M 131 LEU CD1 : -0.564: 0
: 2637:M 131 LEU 2HD1 :M 145 ARG NE : -0.426: 0
: 2637:M 131 LEU CD1 :M 145 ARG NE : -0.416: 0
: 2637:M 72 LYS C :M 74 GLU H : -0.563: 0
: 2637:M 8 PRO CD :M 7 VAL N : -0.535: 0
: 2637:M 61 GLU OE2 :M 7 VAL CG1 : -0.478: 0
: 2637:M 60 ASP OD1 :M 61 GLU N : -0.451: 0
: 2637:M 61 GLU CG :M 62 ALA N : -0.439: 0
: 2637:M 94 LEU N :M 94 LEU 2HD1 : -0.524: 0
: 2637:M 94 LEU CD1 :M 94 LEU N : -0.467: 0
: 2637:M 17 HIS CE1 :M 24 GLU C : -0.519: 0
: 2637:M 17 HIS CD2 :M 25 ASP O : -0.424: 0
: 2637:M 46 LEU N :M 44 ASP O : -0.518: 0
: 2637:M 59 LEU O :M 59 LEU 3HD2 : -0.509: 0
: 2637:M 78 HIS CG :M 79 HIS N : -0.507: 0
: 2637:M 78 HIS ND1 :M 79 HIS N : -0.416: 0
: 2637:M 10 PHE CD2 :M 33 TYR OH : -0.459: 0
: 2637:M 29 ALA N :M 27 ILE C : -0.456: 0
: 2637:M 27 ILE C :M 29 ALA H : -0.448: 0
: 2637:M 33 TYR CB :M 29 ALA O : -0.420: 0
: 2637:M 101 ILE O :M 104 LEU N : -0.450: 0
: 2637:M 143 LEU 3HD1 :M 143 LEU O : -0.444: 0
: 2637:M 63 TYR CD1 :M 63 TYR C : -0.440: 0
: 2637:M 128 VAL O :M 132 VAL 2HG1 : -0.433: 0
: 2637:M 22 GLN O :M 23 SER OG : -0.432: 0
: 2637:M 22 GLN O :M 23 SER CB : -0.401: 0
: 2637:M 19 GLU N :M 18 ILE CG1 : -0.430: 0
: 2637:M 18 ILE H :M 18 ILE 3HG2 : -0.406: 0
: 2637:M 167 HIS N :M 166 HIS CG : -0.423: 0
: 2637:M 53 LEU N :M 52 VAL 3HG2 : -0.421: 0
: 2637:M 58 HIS CG :M 54 LYS O : -0.413: 0
: 2637:M 100 PHE CD2 :M 123 TYR OH : -0.412: 0
: 2637:M 6 SER H :M 4 LEU C : -0.410: 0
: 2637:M 11 ILE O :M 14 LEU N : -0.401: 0
#sum2 ::22.37 clashscore : 22.37 clashscore B<40
#summary::2637 atoms:2637 atoms B<40:296667 potential dots:18540.0 A^2:59 bumps:59 bumps B<40:530.8 score
List of bad contacts calculated by MAGE for model $num_n
/farm/software/bin/probe
: 2637:M 44 ASP O :M 45 GLN O : -0.938: 0
: 2637:M 44 ASP O :M 45 GLN C : -0.546: 0
: 2637:M 51 VAL 2HG2 :M 48 LYS O : -0.723: 0
: 2637:M 47 THR HB :M 51 VAL 1HG2 : -0.503: 0
: 2637:M 48 LYS C :M 50 ALA N : -0.477: 0
: 2637:M 50 ALA H :M 48 LYS 2HG : -0.454: 0
: 2637:M 48 LYS C :M 50 ALA H : -0.408: 0
: 2637:M 14 LEU N :M 14 LEU 2HD1 : -0.690: 0
: 2637:M 14 LEU N :M 14 LEU CD1 : -0.540: 0
: 2637:M 33 TYR CE2 :M 14 LEU 1HD1 : -0.531: 0
: 2637:M 14 LEU CD1 :M 10 PHE O : -0.462: 0
: 2637:M 14 LEU H :M 14 LEU CD1 : -0.440: 0
: 2637:M 66 VAL 1HG2 :M 14 LEU 3HD2 : -0.411: 0
: 2637:M 33 TYR CD2 :M 14 LEU 1HD1 : -0.407: 0
: 2637:M 10 PHE CD2 :M 33 TYR OH : -0.401: 0
: 2637:M 159 GLY H :M 157 LYS C : -0.642: 0
: 2637:M 35 ARG NH2 :M 153 TYR CD2 : -0.615: 0
: 2637:M 117 ILE C :M 119 ALA H : -0.574: 0
: 2637:M 117 ILE O :M 120 MET SD : -0.561: 0
: 2637:M 117 ILE CG2 :M 157 LYS NZ : -0.556: 0
: 2637:M 159 GLY N :M 157 LYS C : -0.542: 0
: 2637:M 120 MET SD :M 121 ASN N : -0.540: 0
: 2637:M 163 LEU O :M 165 HIS N : -0.522: 0
: 2637:M 149 LEU O :M 149 LEU 3HD2 : -0.519: 0
: 2637:M 120 MET O :M 124 TYR N : -0.516: 0
: 2637:M 117 ILE CG2 :M 157 LYS 1HZ : -0.510: 0
: 2637:M 157 LYS O :M 159 GLY N : -0.485: 0
: 2637:M 120 MET SD :M 120 MET N : -0.472: 0
: 2637:M 124 TYR CB :M 120 MET O : -0.469: 0
: 2637:M 163 LEU N :M 159 GLY O : -0.463: 0
: 2637:M 154 ASN N :M 153 TYR CG : -0.424: 0
: 2637:M 153 TYR CD1 :M 153 TYR C : -0.418: 0
: 2637:M 153 TYR O :M 157 LYS 1HG : -0.417: 0
: 2637:M 124 TYR CD2 :M 149 LEU 2HD2 : -0.407: 0
: 2637:M 108 ILE 1HD1 :M 160 GLU OE2 : -0.607: 0
: 2637:M 108 ILE 2HD1 :M 161 SER HA : -0.547: 0
: 2637:M 108 ILE 2HG2 :M 116 LYS O : -0.404: 0
: 2637:M 30 MET SD :M 31 ASN N : -0.583: 0
: 2637:M 34 TYR CB :M 30 MET O : -0.488: 0
: 2637:M 30 MET CG :M 31 ASN N : -0.469: 0
: 2637:M 30 MET O :M 34 TYR N : -0.441: 0
: 2637:M 30 MET C :M 30 MET SD : -0.418: 0
: 2637:M 8 PRO CD :M 7 VAL N : -0.565: 0
: 2637:M 7 VAL 3HG2 :M 7 VAL H : -0.404: 0
: 2637:M 98 PRO CD :M 97 VAL N : -0.564: 0
: 2637:M 97 VAL 3HG2 :M 97 VAL H : -0.438: 0
: 2637:M 127 VAL O :M 131 LEU N : -0.550: 0
: 2637:M 131 LEU CB :M 127 VAL O : -0.469: 0
: 2637:M 135 GLN H :M 134 ASP CG : -0.542: 0
: 2637:M 135 GLN C :M 134 ASP O : -0.506: 0
: 2637:M 135 GLN N :M 134 ASP OD1 : -0.472: 0
: 2637:M 136 LEU N :M 134 ASP O : -0.463: 0
: 2637:M 74 GLU N :M 73 LEU 2HD1 : -0.534: 0
: 2637:M 74 GLU C :M 76 HIS N : -0.444: 0
: 2637:M 74 GLU C :M 76 HIS H : -0.409: 0
: 2637:M 1 MET N :M 1 MET SD : -0.529: 0
: 2637:M 166 HIS N :M 164 GLU O : -0.527: 0
: 2637:M 128 VAL 3HG1 :M 129 SER N : -0.524: 0
: 2637:M 129 SER N :M 128 VAL CG1 : -0.487: 0
: 2637:M 39 SER N :M 38 VAL 3HG2 : -0.513: 0
: 2637:M 39 SER N :M 38 VAL CG2 : -0.410: 0
: 2637:M 148 HIS ND1 :M 144 LYS O : -0.476: 0
: 2637:M 70 GLU OE1 :M 26 LYS NZ : -0.473: 0
: 2637:M 70 GLU CD :M 71 SER N : -0.449: 0
: 2637:M 67 LYS 3HZ :M 67 LYS 2HB : -0.469: 0
: 2637:M 67 LYS NZ :M 67 LYS CB : -0.431: 0
: 2637:M 49 ASN 2HD2 :M 140 ALA CB : -0.467: 0
: 2637:M 20 MET O :M 21 ASN CB : -0.462: 0
: 2637:M 21 ASN CA :M 17 HIS O : -0.439: 0
: 2637:M 3 GLU O :M 2 SER O : -0.458: 0
: 2637:M 3 GLU CD :M 3 GLU N : -0.426: 0
: 2637:M 77 HIS O :M 78 HIS O : -0.455: 0
: 2637:M 60 ASP OD1 :M 61 GLU N : -0.452: 0
: 2637:M 54 LYS O :M 58 HIS CD2 : -0.452: 0
: 2637:M 57 GLN O :M 60 ASP CG : -0.423: 0
: 2637:M 100 PHE CD2 :M 123 TYR OH : -0.451: 0
: 2637:M 12 GLU C :M 12 GLU OE1 : -0.439: 0
: 2637:M 106 GLN O :M 110 MET N : -0.431: 0
: 2637:M 110 MET CB :M 106 GLN O : -0.425: 0
: 2637:M 114 GLU OE2 :M 122 SER CB : -0.428: 0
: 2637:M 114 GLU CG :M 115 ASP H : -0.407: 0
: 2637:M 111 ASN CA :M 107 HIS O : -0.422: 0
: 2637:M 93 GLU O :M 92 SER O : -0.420: 0
: 2637:M 170 HIS ND1 :M 170 HIS C : -0.419: 0
: 2637:M 41 LEU 2HD2 :M 55 ARG 1HD : -0.418: 0
: 2637:M 109 GLU CB :M 105 LYS O : -0.412: 0
: 2637:M 19 GLU N :M 18 ILE CG1 : -0.401: 0
#sum2 ::32.99 clashscore : 32.99 clashscore B<40
#summary::2637 atoms:2637 atoms B<40:296476 potential dots:18530.0 A^2:87 bumps:87 bumps B<40:548.6 score
List of bad contacts calculated by MAGE for model $num_n
/farm/software/bin/probe
: 2637:M 51 VAL 2HG2 :M 48 LYS O : -0.826: 0
: 2637:M 51 VAL 1HG2 :M 47 THR 2HG2 : -0.531: 0
: 2637:M 52 VAL 3HG1 :M 51 VAL 3HG2 : -0.485: 0
: 2637:M 51 VAL 1HG2 :M 47 THR CG2 : -0.459: 0
: 2637:M 48 LYS 2HG :M 50 ALA H : -0.407: 0
: 2637:M 141 VAL 2HG2 :M 138 LYS O : -0.816: 0
: 2637:M 141 VAL 1HG2 :M 137 THR 2HG2 : -0.506: 0
: 2637:M 138 LYS N :M 137 THR 3HG2 : -0.480: 0
: 2637:M 141 VAL 1HG2 :M 137 THR CG2 : -0.446: 0
: 2637:M 138 LYS C :M 140 ALA N : -0.433: 0
: 2637:M 138 LYS N :M 137 THR CG2 : -0.404: 0
: 2637:M 55 ARG 1HH1 :M 41 LEU 1HD2 : -0.772: 0
: 2637:M 41 LEU 1HD2 :M 55 ARG NH1 : -0.628: 0
: 2637:M 54 LYS CG :M 55 ARG N : -0.494: 0
: 2637:M 41 LEU 3HD1 :M 41 LEU O : -0.488: 0
: 2637:M 11 ILE 1HG2 :M 65 LYS 2HD : -0.653: 0
: 2637:M 8 PRO CD :M 7 VAL N : -0.628: 0
: 2637:M 7 VAL 3HG2 :M 7 VAL H : -0.414: 0
: 2637:M 45 GLN H :M 44 ASP CG : -0.604: 0
: 2637:M 45 GLN C :M 44 ASP O : -0.504: 0
: 2637:M 46 LEU N :M 44 ASP O : -0.484: 0
: 2637:M 40 THR O :M 44 ASP N : -0.417: 0
: 2637:M 27 ILE C :M 29 ALA H : -0.589: 0
: 2637:M 27 ILE C :M 29 ALA N : -0.428: 0
: 2637:M 33 TYR CE1 :M 59 LEU 1HD1 : -0.586: 0
: 2637:M 30 MET CG :M 31 ASN N : -0.516: 0
: 2637:M 30 MET O :M 34 TYR N : -0.493: 0
: 2637:M 33 TYR CD1 :M 33 TYR C : -0.488: 0
: 2637:M 59 LEU CD1 :M 33 TYR OH : -0.450: 0
: 2637:M 14 LEU 1HD2 :M 33 TYR CG : -0.434: 0
: 2637:M 14 LEU 3HD1 :M 66 VAL CG2 : -0.427: 0
: 2637:M 59 LEU 1HD1 :M 33 TYR CZ : -0.416: 0
: 2637:M 59 LEU 3HD2 :M 34 TYR CE1 : -0.410: 0
: 2637:M 31 ASN H :M 30 MET 1HG : -0.404: 0
: 2637:M 120 MET SD :M 121 ASN N : -0.551: 0
: 2637:M 120 MET O :M 124 TYR N : -0.500: 0
: 2637:M 120 MET CG :M 121 ASN N : -0.498: 0
: 2637:M 121 ASN OD1 :M 125 ARG NH1 : -0.482: 0
: 2637:M 120 MET 2HG :M 121 ASN H : -0.478: 0
: 2637:M 125 ARG CZ :M 121 ASN OD1 : -0.421: 0
: 2637:M 96 SER CB :M 95 PHE O : -0.534: 0
: 2637:M 96 SER O :M 99 TYR N : -0.504: 0
: 2637:M 96 SER O :M 97 VAL C : -0.495: 0
: 2637:M 98 PRO 1HD :M 97 VAL N : -0.483: 0
: 2637:M 99 TYR 2HB :M 96 SER O : -0.470: 0
: 2637:M 108 ILE CG1 :M 109 GLU N : -0.498: 0
: 2637:M 129 SER N :M 128 VAL 3HG2 : -0.497: 0
: 2637:M 129 SER O :M 133 GLN CB : -0.437: 0
: 2637:M 129 SER N :M 128 VAL CG2 : -0.413: 0
: 2637:M 155 LYS O :M 159 GLY N : -0.484: 0
: 2637:M 163 LEU C :M 162 LYS O : -0.478: 0
: 2637:M 28 HIS O :M 28 HIS CG : -0.472: 0
: 2637:M 24 GLU OE2 :M 32 SER CB : -0.470: 0
: 2637:M 100 PHE CD2 :M 123 TYR OH : -0.465: 0
: 2637:M 103 ASN ND2 :M 107 HIS NE2 : -0.465: 0
: 2637:M 103 ASN O :M 107 HIS CD2 : -0.456: 0
: 2637:M 107 HIS CE1 :M 103 ASN OD1 : -0.420: 0
: 2637:M 123 TYR CD2 :M 104 LEU 1HD2 : -0.416: 0
: 2637:M 165 HIS O :M 166 HIS CG : -0.462: 0
: 2637:M 69 GLY N :M 67 LYS C : -0.453: 0
: 2637:M 67 LYS C :M 69 GLY H : -0.410: 0
: 2637:M 63 TYR CD1 :M 64 ASN N : -0.418: 0
: 2637:M 62 ALA N :M 61 GLU CG : -0.417: 0
: 2637:M 131 LEU O :M 134 ASP O : -0.413: 0
: 2637:M 134 ASP O :M 135 GLN C : -0.403: 0
: 2637:M 19 GLU CB :M 15 LYS O : -0.407: 0
: 2637:M 18 ILE 2HG1 :M 19 GLU N : -0.406: 0
: 2637:M 110 MET CB :M 106 GLN O : -0.406: 0
: 2637:M 168 HIS O :M 167 HIS ND1 : -0.405: 0
#sum2 ::26.17 clashscore : 26.17 clashscore B<40
#summary::2637 atoms:2637 atoms B<40:296472 potential dots:18530.0 A^2:69 bumps:69 bumps B<40:528 score
List of bad contacts calculated by MAGE for model $num_n
/farm/software/bin/probe
: 2637:M 34 TYR O :M 38 VAL 3HG2 : -0.746: 0
: 2637:M 30 MET O :M 34 TYR N : -0.552: 0
: 2637:M 38 VAL N :M 37 VAL 3HG2 : -0.529: 0
: 2637:M 38 VAL O :M 42 VAL 2HG1 : -0.506: 0
: 2637:M 27 ILE 3HG2 :M 28 HIS N : -0.505: 0
: 2637:M 34 TYR CB :M 30 MET O : -0.489: 0
: 2637:M 27 ILE O :M 30 MET CG : -0.464: 0
: 2637:M 28 HIS N :M 27 ILE CG2 : -0.441: 0
: 2637:M 31 ASN H :M 30 MET 2HG : -0.440: 0
: 2637:M 38 VAL N :M 37 VAL CG2 : -0.428: 0
: 2637:M 29 ALA N :M 27 ILE C : -0.426: 0
: 2637:M 34 TYR O :M 38 VAL CG2 : -0.421: 0
: 2637:M 34 TYR CE1 :M 59 LEU 3HD2 : -0.405: 0
: 2637:M 29 ALA H :M 27 ILE C : -0.403: 0
: 2637:M 98 PRO 1HD :M 97 VAL N : -0.719: 0
: 2637:M 97 VAL N :M 98 PRO CD : -0.508: 0
: 2637:M 123 TYR CE2 :M 127 VAL 1HG2 : -0.629: 0
: 2637:M 104 LEU 3HD1 :M 156 VAL CG2 : -0.494: 0
: 2637:M 100 PHE CD2 :M 123 TYR OH : -0.434: 0
: 2637:M 123 TYR CD2 :M 104 LEU 1HD2 : -0.427: 0
: 2637:M 117 ILE C :M 119 ALA H : -0.605: 0
: 2637:M 117 ILE O :M 120 MET CG : -0.467: 0
: 2637:M 117 ILE C :M 119 ALA N : -0.445: 0
: 2637:M 124 TYR CB :M 120 MET O : -0.420: 0
: 2637:M 5 PHE H :M 3 GLU C : -0.570: 0
: 2637:M 3 GLU O :M 4 LEU 2HB : -0.425: 0
: 2637:M 3 GLU C :M 5 PHE N : -0.419: 0
: 2637:M 3 GLU O :M 5 PHE N : -0.413: 0
: 2637:M 136 LEU N :M 136 LEU 2HD1 : -0.559: 0
: 2637:M 136 LEU N :M 136 LEU CD1 : -0.438: 0
: 2637:M 149 LEU O :M 149 LEU 3HD2 : -0.555: 0
: 2637:M 60 ASP OD1 :M 61 GLU N : -0.549: 0
: 2637:M 62 ALA O :M 66 VAL 3HG2 : -0.539: 0
: 2637:M 14 LEU 3HD1 :M 66 VAL CG2 : -0.424: 0
: 2637:M 14 LEU 3HD1 :M 66 VAL 1HG2 : -0.421: 0
: 2637:M 62 ALA N :M 61 GLU CG : -0.421: 0
: 2637:M 57 GLN O :M 60 ASP OD1 : -0.420: 0
: 2637:M 57 GLN O :M 60 ASP CG : -0.407: 0
: 2637:M 139 ASN 2HD2 :M 50 ALA CA : -0.534: 0
: 2637:M 138 LYS 1HG :M 140 ALA H : -0.520: 0
: 2637:M 139 ASN OD1 :M 140 ALA N : -0.511: 0
: 2637:M 139 ASN 2HD2 :M 50 ALA N : -0.500: 0
: 2637:M 48 LYS 1HG :M 50 ALA 3HB : -0.482: 0
: 2637:M 8 PRO 1HD :M 7 VAL N : -0.531: 0
: 2637:M 7 VAL N :M 8 PRO CD : -0.499: 0
: 2637:M 163 LEU N :M 163 LEU 2HD2 : -0.530: 0
: 2637:M 163 LEU N :M 163 LEU CD2 : -0.511: 0
: 2637:M 165 HIS CG :M 166 HIS N : -0.517: 0
: 2637:M 129 SER N :M 128 VAL CG1 : -0.514: 0
: 2637:M 128 VAL 3HG1 :M 129 SER N : -0.478: 0
: 2637:M 128 VAL 3HG2 :M 146 ILE CD1 : -0.419: 0
: 2637:M 144 LYS CG :M 145 ARG N : -0.511: 0
: 2637:M 144 LYS 1HG :M 141 VAL O : -0.422: 0
: 2637:M 45 GLN C :M 44 ASP O : -0.499: 0
: 2637:M 44 ASP N :M 40 THR O : -0.493: 0
: 2637:M 44 ASP C :M 45 GLN CG : -0.427: 0
: 2637:M 45 GLN C :M 47 THR H : -0.406: 0
: 2637:M 73 LEU 2HD1 :M 73 LEU N : -0.491: 0
: 2637:M 73 LEU H :M 73 LEU 2HD1 : -0.451: 0
: 2637:M 73 LEU N :M 73 LEU CD1 : -0.420: 0
: 2637:M 41 LEU C :M 43 GLN H : -0.479: 0
: 2637:M 58 HIS ND1 :M 54 LYS O : -0.478: 0
: 2637:M 164 GLU OE2 :M 116 LYS NZ : -0.462: 0
: 2637:M 94 LEU 2HD1 :M 94 LEU C : -0.461: 0
: 2637:M 94 LEU C :M 94 LEU CD1 : -0.435: 0
: 2637:M 95 PHE N :M 94 LEU 2HD1 : -0.419: 0
: 2637:M 18 ILE 2HG1 :M 19 GLU N : -0.456: 0
: 2637:M 153 TYR CD1 :M 154 ASN N : -0.453: 0
: 2637:M 154 ASN O :M 158 ARG CB : -0.444: 0
: 2637:M 153 TYR CD1 :M 153 TYR C : -0.436: 0
: 2637:M 21 ASN CG :M 20 MET O : -0.444: 0
: 2637:M 152 ALA N :M 151 GLU CG : -0.442: 0
: 2637:M 108 ILE 2HG1 :M 109 GLU N : -0.441: 0
: 2637:M 63 TYR CD1 :M 64 ASN N : -0.441: 0
: 2637:M 63 TYR CD1 :M 63 TYR C : -0.439: 0
: 2637:M 109 GLU CB :M 105 LYS O : -0.411: 0
: 2637:M 109 GLU N :M 108 ILE CG1 : -0.408: 0
: 2637:M 110 MET CB :M 106 GLN O : -0.440: 0
: 2637:M 106 GLN O :M 110 MET N : -0.435: 0
: 2637:M 91 MET C :M 91 MET SD : -0.438: 0
: 2637:M 133 GLN O :M 134 ASP C : -0.433: 0
: 2637:M 130 THR O :M 134 ASP N : -0.409: 0
: 2637:M 69 GLY N :M 67 LYS C : -0.424: 0
: 2637:M 69 GLY H :M 67 LYS C : -0.413: 0
: 2637:M 33 TYR CD1 :M 33 TYR C : -0.421: 0
: 2637:M 10 PHE CD2 :M 33 TYR OH : -0.412: 0
: 2637:M 74 GLU O :M 75 HIS O : -0.418: 0
: 2637:M 114 GLU O :M 115 ASP CG : -0.417: 0
#sum2 ::33.37 clashscore : 33.37 clashscore B<40
#summary::2637 atoms:2637 atoms B<40:295793 potential dots:18490.0 A^2:88 bumps:88 bumps B<40:496.3 score
List of bad contacts calculated by MAGE for model $num_n
/farm/software/bin/probe
: 2637:M 157 LYS 1HD :M 117 ILE 1HD1 : -0.658: 0
: 2637:M 157 LYS O :M 160 GLU N : -0.441: 0
: 2637:M 160 GLU O :M 162 LYS N : -0.436: 0
: 2637:M 117 ILE C :M 119 ALA H : -0.415: 0
: 2637:M 117 ILE 3HG2 :M 118 HIS N : -0.401: 0
: 2637:M 98 PRO 1HD :M 97 VAL N : -0.651: 0
: 2637:M 97 VAL N :M 98 PRO CD : -0.455: 0
: 2637:M 14 LEU N :M 14 LEU 2HD1 : -0.620: 0
: 2637:M 14 LEU N :M 14 LEU CD1 : -0.524: 0
: 2637:M 14 LEU 1HD1 :M 33 TYR CD1 : -0.486: 0
: 2637:M 10 PHE CD2 :M 33 TYR OH : -0.433: 0
: 2637:M 14 LEU 1HD1 :M 33 TYR CE1 : -0.427: 0
: 2637:M 27 ILE 1HD1 :M 67 LYS 1HD : -0.619: 0
: 2637:M 27 ILE O :M 30 MET SD : -0.615: 0
: 2637:M 27 ILE C :M 29 ALA H : -0.583: 0
: 2637:M 30 MET SD :M 31 ASN N : -0.560: 0
: 2637:M 30 MET SD :M 30 MET N : -0.527: 0
: 2637:M 30 MET O :M 34 TYR N : -0.495: 0
: 2637:M 34 TYR CB :M 30 MET O : -0.483: 0
: 2637:M 28 HIS N :M 27 ILE 3HG2 : -0.426: 0
: 2637:M 31 ASN N :M 30 MET CG : -0.404: 0
: 2637:M 94 LEU N :M 94 LEU 2HD2 : -0.606: 0
: 2637:M 94 LEU N :M 94 LEU CD2 : -0.551: 0
: 2637:M 94 LEU O :M 96 SER N : -0.536: 0
: 2637:M 94 LEU H :M 94 LEU 2HD2 : -0.482: 0
: 2637:M 43 GLN O :M 44 ASP C : -0.595: 0
: 2637:M 120 MET SD :M 121 ASN N : -0.574: 0
: 2637:M 129 SER N :M 128 VAL 3HG2 : -0.561: 0
: 2637:M 124 TYR CB :M 120 MET O : -0.485: 0
: 2637:M 120 MET O :M 124 TYR N : -0.459: 0
: 2637:M 128 VAL CG2 :M 129 SER N : -0.459: 0
: 2637:M 128 VAL 1HG1 :M 124 TYR CE1 : -0.446: 0
: 2637:M 120 MET CG :M 121 ASN N : -0.437: 0
: 2637:M 42 VAL 3HG2 :M 38 VAL O : -0.567: 0
: 2637:M 39 SER N :M 38 VAL 3HG2 : -0.550: 0
: 2637:M 42 VAL O :M 42 VAL 2HG1 : -0.535: 0
: 2637:M 42 VAL O :M 42 VAL CG1 : -0.467: 0
: 2637:M 39 SER N :M 38 VAL CG2 : -0.452: 0
: 2637:M 38 VAL O :M 42 VAL CG2 : -0.441: 0
: 2637:M 8 PRO 1HD :M 7 VAL N : -0.543: 0
: 2637:M 9 TYR N :M 7 VAL O : -0.518: 0
: 2637:M 6 SER OG :M 7 VAL N : -0.495: 0
: 2637:M 9 TYR N :M 7 VAL C : -0.435: 0
: 2637:M 91 MET SD :M 91 MET N : -0.536: 0
: 2637:M 48 LYS 1HG :M 50 ALA H : -0.524: 0
: 2637:M 37 VAL O :M 41 LEU N : -0.518: 0
: 2637:M 37 VAL O :M 41 LEU CB : -0.435: 0
: 2637:M 165 HIS CG :M 166 HIS N : -0.517: 0
: 2637:M 73 LEU C :M 75 HIS H : -0.495: 0
: 2637:M 75 HIS ND1 :M 76 HIS N : -0.436: 0
: 2637:M 73 LEU C :M 75 HIS N : -0.412: 0
: 2637:M 24 GLU OE2 :M 32 SER CB : -0.494: 0
: 2637:M 18 ILE CG1 :M 19 GLU N : -0.489: 0
: 2637:M 26 LYS O :M 18 ILE 2HG2 : -0.442: 0
: 2637:M 148 HIS CG :M 144 LYS O : -0.473: 0
: 2637:M 144 LYS NZ :M 144 LYS CB : -0.414: 0
: 2637:M 45 GLN O :M 46 LEU C : -0.469: 0
: 2637:M 134 ASP O :M 135 GLN CB : -0.468: 0
: 2637:M 134 ASP O :M 135 GLN 2HB : -0.461: 0
: 2637:M 134 ASP O :M 131 LEU O : -0.402: 0
: 2637:M 135 GLN O :M 136 LEU C : -0.401: 0
: 2637:M 110 MET O :M 111 ASN CB : -0.467: 0
: 2637:M 111 ASN 1HD2 :M 115 ASP HA : -0.427: 0
: 2637:M 111 ASN CA :M 107 HIS O : -0.424: 0
: 2637:M 143 LEU N :M 142 VAL 3HG2 : -0.466: 0
: 2637:M 61 GLU OE1 :M 58 HIS CD2 : -0.464: 0
: 2637:M 100 PHE CD2 :M 123 TYR OH : -0.456: 0
: 2637:M 123 TYR CD1 :M 123 TYR C : -0.414: 0
: 2637:M 164 GLU C :M 163 LEU O : -0.452: 0
: 2637:M 163 LEU 3HD1 :M 163 LEU HA : -0.432: 0
: 2637:M 139 ASN ND2 :M 53 LEU CD1 : -0.449: 0
: 2637:M 169 HIS CG :M 168 HIS O : -0.443: 0
: 2637:M 169 HIS CG :M 168 HIS C : -0.432: 0
: 2637:M 138 LYS 1HG :M 140 ALA H : -0.417: 0
#sum2 ::28.06 clashscore : 28.06 clashscore B<40
#summary::2637 atoms:2637 atoms B<40:296061 potential dots:18500.0 A^2:74 bumps:74 bumps B<40:552.2 score
List of bad contacts calculated by MAGE for model $num_n
/farm/software/bin/probe
: 2637:M 42 VAL O :M 42 VAL 2HG1 : -0.730: 0
: 2637:M 147 GLN 1HE2 :M 42 VAL CG1 : -0.671: 0
: 2637:M 42 VAL O :M 42 VAL CG1 : -0.527: 0
: 2637:M 124 TYR O :M 128 VAL 3HG2 : -0.682: 0
: 2637:M 27 ILE O :M 30 MET SD : -0.599: 0
: 2637:M 30 MET SD :M 30 MET N : -0.594: 0
: 2637:M 38 VAL 1HG2 :M 34 TYR CE1 : -0.586: 0
: 2637:M 120 MET SD :M 121 ASN N : -0.562: 0
: 2637:M 124 TYR N :M 120 MET O : -0.562: 0
: 2637:M 128 VAL N :M 127 VAL 3HG2 : -0.532: 0
: 2637:M 30 MET SD :M 31 ASN N : -0.517: 0
: 2637:M 30 MET O :M 34 TYR N : -0.498: 0
: 2637:M 125 ARG NH1 :M 34 TYR CZ : -0.494: 0
: 2637:M 128 VAL N :M 127 VAL CG2 : -0.492: 0
: 2637:M 121 ASN ND2 :M 31 ASN OD1 : -0.478: 0
: 2637:M 128 VAL CG2 :M 124 TYR O : -0.474: 0
: 2637:M 34 TYR OH :M 125 ARG NH1 : -0.462: 0
: 2637:M 34 TYR CB :M 30 MET O : -0.450: 0
: 2637:M 121 ASN 1HD2 :M 31 ASN CG : -0.450: 0
: 2637:M 120 MET CG :M 121 ASN N : -0.436: 0
: 2637:M 124 TYR 1HB :M 120 MET O : -0.434: 0
: 2637:M 27 ILE 3HG2 :M 28 HIS N : -0.405: 0
: 2637:M 31 ASN CG :M 121 ASN ND2 : -0.400: 0
: 2637:M 53 LEU C :M 53 LEU 3HD2 : -0.653: 0
: 2637:M 53 LEU N :M 52 VAL 3HG2 : -0.580: 0
: 2637:M 52 VAL O :M 56 ILE 1HG1 : -0.539: 0
: 2637:M 49 ASN O :M 53 LEU CB : -0.491: 0
: 2637:M 53 LEU C :M 53 LEU CD2 : -0.467: 0
: 2637:M 53 LEU N :M 52 VAL CG2 : -0.459: 0
: 2637:M 52 VAL 2HG2 :M 48 LYS O : -0.407: 0
: 2637:M 98 PRO CD :M 97 VAL N : -0.639: 0
: 2637:M 97 VAL N :M 98 PRO 1HD : -0.436: 0
: 2637:M 145 ARG CZ :M 131 LEU 2HD2 : -0.624: 0
: 2637:M 165 HIS NE2 :M 166 HIS NE2 : -0.575: 0
: 2637:M 165 HIS NE2 :M 166 HIS CE1 : -0.550: 0
: 2637:M 165 HIS NE2 :M 166 HIS CD2 : -0.532: 0
: 2637:M 166 HIS CG :M 165 HIS CD2 : -0.522: 0
: 2637:M 166 HIS CD2 :M 165 HIS CD2 : -0.472: 0
: 2637:M 165 HIS CG :M 166 HIS N : -0.467: 0
: 2637:M 9 TYR N :M 7 VAL O : -0.569: 0
: 2637:M 8 PRO 1HD :M 7 VAL N : -0.516: 0
: 2637:M 9 TYR N :M 7 VAL C : -0.447: 0
: 2637:M 7 VAL O :M 8 PRO C : -0.427: 0
: 2637:M 134 ASP N :M 130 THR O : -0.546: 0
: 2637:M 63 TYR CD1 :M 64 ASN N : -0.517: 0
: 2637:M 64 ASN ND2 :M 61 GLU O : -0.509: 0
: 2637:M 67 LYS N :M 66 VAL CG1 : -0.479: 0
: 2637:M 66 VAL 2HG1 :M 63 TYR O : -0.466: 0
: 2637:M 160 GLU CD :M 161 SER N : -0.516: 0
: 2637:M 161 SER N :M 160 GLU CG : -0.501: 0
: 2637:M 161 SER H :M 160 GLU CG : -0.500: 0
: 2637:M 161 SER H :M 160 GLU 2HG : -0.465: 0
: 2637:M 156 VAL O :M 160 GLU CG : -0.450: 0
: 2637:M 160 GLU OE1 :M 117 ILE 1HD1 : -0.410: 0
: 2637:M 117 ILE C :M 119 ALA H : -0.405: 0
: 2637:M 95 PHE H :M 93 GLU C : -0.489: 0
: 2637:M 95 PHE N :M 93 GLU C : -0.450: 0
: 2637:M 93 GLU O :M 92 SER O : -0.437: 0
: 2637:M 138 LYS 1HG :M 140 ALA H : -0.480: 0
: 2637:M 143 LEU N :M 142 VAL 3HG2 : -0.427: 0
: 2637:M 142 VAL 2HG2 :M 138 LYS O : -0.412: 0
: 2637:M 110 MET C :M 111 ASN ND2 : -0.477: 0
: 2637:M 107 HIS O :M 111 ASN OD1 : -0.439: 0
: 2637:M 111 ASN O :M 112 GLN C : -0.434: 0
: 2637:M 110 MET C :M 111 ASN 2HD2 : -0.422: 0
: 2637:M 107 HIS O :M 111 ASN HA : -0.416: 0
: 2637:M 111 ASN CA :M 107 HIS O : -0.412: 0
: 2637:M 155 LYS O :M 159 GLY N : -0.467: 0
: 2637:M 123 TYR O :M 123 TYR CD1 : -0.465: 0
: 2637:M 104 LEU 1HD2 :M 123 TYR CD1 : -0.439: 0
: 2637:M 123 TYR CD1 :M 123 TYR C : -0.411: 0
: 2637:M 4 LEU C :M 4 LEU 3HD2 : -0.440: 0
: 2637:M 4 LEU C :M 4 LEU CD2 : -0.417: 0
: 2637:M 109 GLU CB :M 105 LYS O : -0.420: 0
: 2637:M 148 HIS CG :M 144 LYS O : -0.417: 0
: 2637:M 55 ARG N :M 54 LYS CG : -0.411: 0
: 2637:M 18 ILE 2HG1 :M 19 GLU N : -0.405: 0
: 2637:M 19 GLU N :M 18 ILE CG1 : -0.400: 0
: 2637:M 26 LYS O :M 29 ALA 2HB : -0.400: 0
: 2637:M 70 GLU OE2 :M 26 LYS NZ : -0.400: 0
: 2637:M 58 HIS ND1 :M 59 LEU N : -0.400: 0
: 2637:M 36 SER O :M 40 THR OG1 : -0.400: 0
#sum2 ::31.10 clashscore : 31.10 clashscore B<40
#summary::2637 atoms:2637 atoms B<40:296246 potential dots:18520.0 A^2:82 bumps:82 bumps B<40:471.2 score
Output from PDB validation software
Summary from PDB validation
May. 10, 18:57:34 2013
[ Text modified to reflect that this was run under PSVS - Aneerban Bhattacharya: Dec 2005 ]
The following checks were made on :
-----------------------------------------
CLOSE CONTACTS
==> Distances smaller than 2.2 Angstroms are considered as close contacts
for heavy atoms, 1.6 Angstroms for hydrogens.
Chain Atom Res Seq Chain Atom Res Seq Mol_ID Distance
-------------------------------------------------------------------------
A HD1 HIS 78 - A H HIS 80 15 Dist = 1.26
B 1HD2 ASN 13 - B HE2 HIS 17 5 Dist = 1.37
B O SER 71 - B H LEU 73 8 Dist = 1.38
B 1HD2 ASN 13 - B HE2 HIS 17 17 Dist = 1.38
B 1HE2 GLN 45 - B HE ARG 55 11 Dist = 1.39
B O MET 30 - B H TYR 34 2 Dist = 1.40
B O VAL 37 - B H LEU 41 16 Dist = 1.41
A O VAL 37 - A H LEU 41 19 Dist = 1.42
A O MET 30 - A H TYR 34 20 Dist = 1.43
A O SER 6 - A H TYR 9 13 Dist = 1.44
B O MET 30 - B H TYR 34 6 Dist = 1.44
A O MET 30 - A H TYR 34 18 Dist = 1.44
A O MET 30 - A H TYR 34 12 Dist = 1.44
B O MET 30 - B H TYR 34 1 Dist = 1.45
A O MET 30 - A H TYR 34 7 Dist = 1.45
A O VAL 37 - A H LEU 41 14 Dist = 1.46
B O VAL 37 - B H LEU 41 8 Dist = 1.46
A O MET 30 - A H TYR 34 3 Dist = 1.46
A O MET 30 - A H TYR 34 8 Dist = 1.47
B 2HZ LYS 65 - B 1HH2 ARG 68 6 Dist = 1.47
A O MET 30 - A H TYR 34 9 Dist = 1.47
A HE2 HIS 28 - A 2HD2 ASN 31 8 Dist = 1.47
B O MET 30 - B H TYR 34 9 Dist = 1.47
B O MET 30 - B H TYR 34 19 Dist = 1.47
A O MET 30 - A H TYR 34 13 Dist = 1.47
B O VAL 37 - B H LEU 41 11 Dist = 1.48
A O MET 30 - A H TYR 34 19 Dist = 1.48
B O MET 30 - B H TYR 34 16 Dist = 1.48
B O LEU 73 - B H HIS 75 16 Dist = 1.49
B O VAL 37 - B H LEU 41 20 Dist = 1.49
A O VAL 37 - A H LEU 41 11 Dist = 1.49
A HE2 HIS 28 - A 2HD2 ASN 31 7 Dist = 1.49
A O THR 40 - A H ASP 44 1 Dist = 1.49
A O MET 30 - A H TYR 34 6 Dist = 1.49
A 1H MET 1 - A HH TYR 9 13 Dist = 1.49
B O MET 30 - B H TYR 34 3 Dist = 1.49
A O MET 30 - A H TYR 34 16 Dist = 1.49
A 2HE2 GLN 16 - A 2HE2 GLN 22 9 Dist = 1.50
B O MET 30 - B H TYR 34 12 Dist = 1.50
B O MET 30 - B H TYR 34 10 Dist = 1.50
B HD1 HIS 75 - B H HIS 77 5 Dist = 1.50
B O VAL 37 - B H LEU 41 13 Dist = 1.50
A O THR 40 - A H ASP 44 8 Dist = 1.50
B O MET 30 - B H TYR 34 13 Dist = 1.50
B O SER 6 - B H TYR 9 5 Dist = 1.51
A O MET 30 - A H TYR 34 5 Dist = 1.51
B O MET 30 - B H TYR 34 18 Dist = 1.51
B O MET 30 - B H TYR 34 14 Dist = 1.51
B O MET 30 - B H TYR 34 7 Dist = 1.51
B O SER 6 - B H TYR 9 17 Dist = 1.51
B O ALA 50 - B H LYS 54 1 Dist = 1.52
A O MET 30 - A H TYR 34 15 Dist = 1.52
A O ASN 49 - A H LEU 53 14 Dist = 1.52
A 2HD2 ASN 31 - B HE ARG 35 12 Dist = 1.52
A O MET 30 - A H TYR 34 10 Dist = 1.53
A O MET 30 - A H TYR 34 2 Dist = 1.53
A O VAL 37 - A H LEU 41 8 Dist = 1.53
A O VAL 37 - A H LEU 41 3 Dist = 1.54
B O THR 40 - B H ASP 44 18 Dist = 1.54
A O VAL 37 - A H LEU 41 18 Dist = 1.54
A O LEU 4 - A H SER 6 15 Dist = 1.55
A O ALA 50 - A H LYS 54 5 Dist = 1.55
A 2HZ LYS 65 - A 1HH2 ARG 68 4 Dist = 1.55
B O LEU 59 - B H TYR 63 11 Dist = 1.55
A O ASN 49 - A H LEU 53 20 Dist = 1.56
B HD1 HIS 28 - B 2HD2 ASN 31 9 Dist = 1.56
B O MET 30 - B H TYR 34 4 Dist = 1.56
A O MET 30 - A H TYR 34 4 Dist = 1.56
A O ALA 50 - A H LYS 54 10 Dist = 1.56
B O LEU 4 - B H SER 6 1 Dist = 1.57
B 1HE2 GLN 45 - B 2HH1 ARG 55 14 Dist = 1.57
B 3HZ LYS 72 - B HE2 HIS 78 14 Dist = 1.58
A O GLU 3 - A H PHE 5 18 Dist = 1.58
A O VAL 37 - A H LEU 41 17 Dist = 1.58
B 1HE2 GLN 45 - B 2HH1 ARG 55 10 Dist = 1.58
B O HIS 28 - B H SER 32 13 Dist = 1.58
A O VAL 66 - A H GLU 70 16 Dist = 1.58
B O HIS 75 - B H HIS 77 11 Dist = 1.59
B O MET 30 - B H TYR 34 11 Dist = 1.59
B O MET 30 - B H TYR 34 8 Dist = 1.59
B O ASN 49 - B H LEU 53 19 Dist = 1.59
B 1HH2 ARG 55 - B HD1 HIS 58 4 Dist = 1.59
A 3HZ LYS 65 - A 1HH2 ARG 68 7 Dist = 1.60
A O SER 71 - A H LEU 73 8 Dist = 1.60
A O MET 1 - A H GLU 3 10 Dist = 1.60
A O ASN 64 - A H ARG 68 8 Dist = 1.60
DISTANCES AND ANGLES
We have checked your intra and intermolecular distances and angles with the
procedures currently in place at PDB:
==> Bond and angle checks are performed by first computing the average rms
error for all bonds and angles relative to standard values for nucleotide
units [L. Clowney et al., Geometric Parameters in Nucleic Acids: Nitrogenous
Bases, J.Am.Chem.Soc. 1996, 118, 509-518; A. Gelbin et al., Geometric
Parameters in Nucleic Acids: Sugar and Phosphate Constituents, J.Am.Chem.Soc.
1996, 118, 519-529] and amino acid units [R.A. Engh and R. Huber, Accurate
Bond and Angle Parameters for X-ray protein structure refinement, Acta
Crystallogr. 1991, A47, 392-400]. Any bond or angle which deviates from the
dictionary values by more than six times this computed rms error is
identified as an outlier.
*** Covalent Bond Lengths:
The RMS deviation for covalent bonds relative to the standard
dictionary is 0.013 Angstroms
All covalent bonds lie within a 6.0*RMSD range about the
standard dictionary values.
*** Covalent Angle Values:
The RMS deviation for covalent angles relative to the standard
dictionary is 1.5 degrees.
All covalent bond angles lie within a 6.0*RMSD range about the
standard dictionary values.
TORSION ANGLES
The torsion angle distributions have been checked. The postscript file of the
conformation rings showing the torsion angle distributions will be sent in a
separate E-mail message.
CHIRALITY
The chirality has been checked. O1P, O2P, and hydrogen atoms which do not
follow the convention defined in the IUBMB (Liebecq, C. Compendium of
Biochemical Nomenclature and Related Documents, 2nd ed.; Portland Press:
London and Chapel Hill, 1992) and IUPAC nomenclature (J.L. Markley, A. Bax,
Y. Arata, C.W. Hilbers, R. Kaptein, B.D. Sykes, P.E. Wright and K. W�thrich,
Recommendations for the Presentation of NMR Structures of Proteins and
Nucleic Acids, Pure & Appl. Chem., Vol. 70, pp. 117-142, 1998) have been
standardized. Any other stereochemical violations are listed below.
E/Z NOMENCLATURE
E/Z nomenclature of hydrogens and/or nitrogens on Arg, Asn or Gln residues needs
to be corrected to conform with the standard for E/Z orientation presented in
[J.L. Markley, et al., Recommendations for the Presentation of NMR Structures of
Proteins and Nucleic Acids, Pure & Appl. Chem., 1998, 70, 117-142].
Model Chain Residue Residue Atom Name Original
Name Number Atom Name
----- ----- ------- ------- -------- ---------
1 A ASN 13 1HD2
1 A ASN 13 2HD2
1 A GLN 16 1HE2
1 A GLN 16 2HE2
1 A ASN 21 1HD2
1 A ASN 21 2HD2
1 A GLN 22 1HE2
1 A GLN 22 2HE2
1 A ASN 31 1HD2
1 A ASN 31 2HD2
1 A GLN 43 1HE2
1 A GLN 43 2HE2
1 A GLN 45 1HE2
1 A GLN 45 2HE2
1 A ASN 49 1HD2
1 A ASN 49 2HD2
1 A GLN 57 1HE2
1 A GLN 57 2HE2
1 A ASN 64 1HD2
1 A ASN 64 2HD2
1 B ASN 13 1HD2
1 B ASN 13 2HD2
1 B GLN 16 1HE2
1 B GLN 16 2HE2
1 B ASN 21 1HD2
1 B ASN 21 2HD2
1 B GLN 22 1HE2
1 B GLN 22 2HE2
1 B ASN 31 1HD2
1 B ASN 31 2HD2
1 B GLN 43 1HE2
1 B GLN 43 2HE2
1 B GLN 45 1HE2
1 B GLN 45 2HE2
1 B ASN 49 1HD2
1 B ASN 49 2HD2
1 B GLN 57 1HE2
1 B GLN 57 2HE2
1 B ASN 64 1HD2
1 B ASN 64 2HD2
2 A ASN 13 1HD2
2 A ASN 13 2HD2
2 A GLN 16 1HE2
2 A GLN 16 2HE2
2 A ASN 21 1HD2
2 A ASN 21 2HD2
2 A GLN 22 1HE2
2 A GLN 22 2HE2
2 A ASN 31 1HD2
2 A ASN 31 2HD2
2 A GLN 43 1HE2
2 A GLN 43 2HE2
2 A GLN 45 1HE2
2 A GLN 45 2HE2
2 A ASN 49 1HD2
2 A ASN 49 2HD2
2 A GLN 57 1HE2
2 A GLN 57 2HE2
2 A ASN 64 1HD2
2 A ASN 64 2HD2
2 B ASN 13 1HD2
2 B ASN 13 2HD2
2 B GLN 16 1HE2
2 B GLN 16 2HE2
2 B ASN 21 1HD2
2 B ASN 21 2HD2
2 B GLN 22 1HE2
2 B GLN 22 2HE2
2 B ASN 31 1HD2
2 B ASN 31 2HD2
2 B GLN 43 1HE2
2 B GLN 43 2HE2
2 B GLN 45 1HE2
2 B GLN 45 2HE2
2 B ASN 49 1HD2
2 B ASN 49 2HD2
2 B GLN 57 1HE2
2 B GLN 57 2HE2
2 B ASN 64 1HD2
2 B ASN 64 2HD2
3 A ASN 13 1HD2
3 A ASN 13 2HD2
3 A GLN 16 1HE2
3 A GLN 16 2HE2
3 A ASN 21 1HD2
3 A ASN 21 2HD2
3 A GLN 22 1HE2
3 A GLN 22 2HE2
3 A ASN 31 1HD2
3 A ASN 31 2HD2
3 A GLN 43 1HE2
3 A GLN 43 2HE2
3 A GLN 45 1HE2
3 A GLN 45 2HE2
3 A ASN 49 1HD2
3 A ASN 49 2HD2
3 A GLN 57 1HE2
3 A GLN 57 2HE2
3 A ASN 64 1HD2
3 A ASN 64 2HD2
3 B ASN 13 1HD2
3 B ASN 13 2HD2
3 B GLN 16 1HE2
3 B GLN 16 2HE2
3 B ASN 21 1HD2
3 B ASN 21 2HD2
3 B GLN 22 1HE2
3 B GLN 22 2HE2
3 B ASN 31 1HD2
3 B ASN 31 2HD2
3 B GLN 43 1HE2
3 B GLN 43 2HE2
3 B GLN 45 1HE2
3 B GLN 45 2HE2
3 B ASN 49 1HD2
3 B ASN 49 2HD2
3 B GLN 57 1HE2
3 B GLN 57 2HE2
3 B ASN 64 1HD2
3 B ASN 64 2HD2
4 A ASN 13 1HD2
4 A ASN 13 2HD2
4 A GLN 16 1HE2
4 A GLN 16 2HE2
4 A ASN 21 1HD2
4 A ASN 21 2HD2
4 A GLN 22 1HE2
4 A GLN 22 2HE2
4 A ASN 31 1HD2
4 A ASN 31 2HD2
4 A GLN 43 1HE2
4 A GLN 43 2HE2
4 A GLN 45 1HE2
4 A GLN 45 2HE2
4 A ASN 49 1HD2
4 A ASN 49 2HD2
4 A GLN 57 1HE2
4 A GLN 57 2HE2
4 A ASN 64 1HD2
4 A ASN 64 2HD2
4 B ASN 13 1HD2
4 B ASN 13 2HD2
4 B GLN 16 1HE2
4 B GLN 16 2HE2
4 B ASN 21 1HD2
4 B ASN 21 2HD2
4 B GLN 22 1HE2
4 B GLN 22 2HE2
4 B ASN 31 1HD2
4 B ASN 31 2HD2
4 B GLN 43 1HE2
4 B GLN 43 2HE2
4 B GLN 45 1HE2
4 B GLN 45 2HE2
4 B ASN 49 1HD2
4 B ASN 49 2HD2
4 B GLN 57 1HE2
4 B GLN 57 2HE2
4 B ASN 64 1HD2
4 B ASN 64 2HD2
5 A ASN 13 1HD2
5 A ASN 13 2HD2
5 A GLN 16 1HE2
5 A GLN 16 2HE2
5 A ASN 21 1HD2
5 A ASN 21 2HD2
5 A GLN 22 1HE2
5 A GLN 22 2HE2
5 A ASN 31 1HD2
5 A ASN 31 2HD2
5 A GLN 43 1HE2
5 A GLN 43 2HE2
5 A GLN 45 1HE2
5 A GLN 45 2HE2
5 A ASN 49 1HD2
5 A ASN 49 2HD2
5 A GLN 57 1HE2
5 A GLN 57 2HE2
5 A ASN 64 1HD2
5 A ASN 64 2HD2
5 B ASN 13 1HD2
5 B ASN 13 2HD2
5 B GLN 16 1HE2
5 B GLN 16 2HE2
5 B ASN 21 1HD2
5 B ASN 21 2HD2
5 B GLN 22 1HE2
5 B GLN 22 2HE2
5 B ASN 31 1HD2
5 B ASN 31 2HD2
5 B GLN 43 1HE2
5 B GLN 43 2HE2
5 B GLN 45 1HE2
5 B GLN 45 2HE2
5 B ASN 49 1HD2
5 B ASN 49 2HD2
5 B GLN 57 1HE2
5 B GLN 57 2HE2
5 B ASN 64 1HD2
5 B ASN 64 2HD2
6 A ASN 13 1HD2
6 A ASN 13 2HD2
6 A GLN 16 1HE2
6 A GLN 16 2HE2
6 A ASN 21 1HD2
6 A ASN 21 2HD2
6 A GLN 22 1HE2
6 A GLN 22 2HE2
6 A ASN 31 1HD2
6 A ASN 31 2HD2
6 A GLN 43 1HE2
6 A GLN 43 2HE2
6 A GLN 45 1HE2
6 A GLN 45 2HE2
6 A ASN 49 1HD2
6 A ASN 49 2HD2
6 A GLN 57 1HE2
6 A GLN 57 2HE2
6 A ASN 64 1HD2
6 A ASN 64 2HD2
6 B ASN 13 1HD2
6 B ASN 13 2HD2
6 B GLN 16 1HE2
6 B GLN 16 2HE2
6 B ASN 21 1HD2
6 B ASN 21 2HD2
6 B GLN 22 1HE2
6 B GLN 22 2HE2
6 B ASN 31 1HD2
6 B ASN 31 2HD2
6 B GLN 43 1HE2
6 B GLN 43 2HE2
6 B GLN 45 1HE2
6 B GLN 45 2HE2
6 B ASN 49 1HD2
6 B ASN 49 2HD2
6 B GLN 57 1HE2
6 B GLN 57 2HE2
6 B ASN 64 1HD2
6 B ASN 64 2HD2
7 A ASN 13 1HD2
7 A ASN 13 2HD2
7 A GLN 16 1HE2
7 A GLN 16 2HE2
7 A ASN 21 1HD2
7 A ASN 21 2HD2
7 A GLN 22 1HE2
7 A GLN 22 2HE2
7 A ASN 31 1HD2
7 A ASN 31 2HD2
7 A GLN 43 1HE2
7 A GLN 43 2HE2
7 A GLN 45 1HE2
7 A GLN 45 2HE2
7 A ASN 49 1HD2
7 A ASN 49 2HD2
7 A GLN 57 1HE2
7 A GLN 57 2HE2
7 A ASN 64 1HD2
7 A ASN 64 2HD2
7 B ASN 13 1HD2
7 B ASN 13 2HD2
7 B GLN 16 1HE2
7 B GLN 16 2HE2
7 B ASN 21 1HD2
7 B ASN 21 2HD2
7 B GLN 22 1HE2
7 B GLN 22 2HE2
7 B ASN 31 1HD2
7 B ASN 31 2HD2
7 B GLN 43 1HE2
7 B GLN 43 2HE2
7 B GLN 45 1HE2
7 B GLN 45 2HE2
7 B ASN 49 1HD2
7 B ASN 49 2HD2
7 B GLN 57 1HE2
7 B GLN 57 2HE2
7 B ASN 64 1HD2
7 B ASN 64 2HD2
8 A ASN 13 1HD2
8 A ASN 13 2HD2
8 A GLN 16 1HE2
8 A GLN 16 2HE2
8 A ASN 21 1HD2
8 A ASN 21 2HD2
8 A GLN 22 1HE2
8 A GLN 22 2HE2
8 A ASN 31 1HD2
8 A ASN 31 2HD2
8 A GLN 43 1HE2
8 A GLN 43 2HE2
8 A GLN 45 1HE2
8 A GLN 45 2HE2
8 A ASN 49 1HD2
8 A ASN 49 2HD2
8 A GLN 57 1HE2
8 A GLN 57 2HE2
8 A ASN 64 1HD2
8 A ASN 64 2HD2
8 B ASN 13 1HD2
8 B ASN 13 2HD2
8 B GLN 16 1HE2
8 B GLN 16 2HE2
8 B ASN 21 1HD2
8 B ASN 21 2HD2
8 B GLN 22 1HE2
8 B GLN 22 2HE2
8 B ASN 31 1HD2
8 B ASN 31 2HD2
8 B GLN 43 1HE2
8 B GLN 43 2HE2
8 B GLN 45 1HE2
8 B GLN 45 2HE2
8 B ASN 49 1HD2
8 B ASN 49 2HD2
8 B GLN 57 1HE2
8 B GLN 57 2HE2
8 B ASN 64 1HD2
8 B ASN 64 2HD2
9 A ASN 13 1HD2
9 A ASN 13 2HD2
9 A GLN 16 1HE2
9 A GLN 16 2HE2
9 A ASN 21 1HD2
9 A ASN 21 2HD2
9 A GLN 22 1HE2
9 A GLN 22 2HE2
9 A ASN 31 1HD2
9 A ASN 31 2HD2
9 A GLN 43 1HE2
9 A GLN 43 2HE2
9 A GLN 45 1HE2
9 A GLN 45 2HE2
9 A ASN 49 1HD2
9 A ASN 49 2HD2
9 A GLN 57 1HE2
9 A GLN 57 2HE2
9 A ASN 64 1HD2
9 A ASN 64 2HD2
9 B ASN 13 1HD2
9 B ASN 13 2HD2
9 B GLN 16 1HE2
9 B GLN 16 2HE2
9 B ASN 21 1HD2
9 B ASN 21 2HD2
9 B GLN 22 1HE2
9 B GLN 22 2HE2
9 B ASN 31 1HD2
9 B ASN 31 2HD2
9 B GLN 43 1HE2
9 B GLN 43 2HE2
9 B GLN 45 1HE2
9 B GLN 45 2HE2
9 B ASN 49 1HD2
9 B ASN 49 2HD2
9 B GLN 57 1HE2
9 B GLN 57 2HE2
9 B ASN 64 1HD2
9 B ASN 64 2HD2
10 A ASN 13 1HD2
10 A ASN 13 2HD2
10 A GLN 16 1HE2
10 A GLN 16 2HE2
10 A ASN 21 1HD2
10 A ASN 21 2HD2
10 A GLN 22 1HE2
10 A GLN 22 2HE2
10 A ASN 31 1HD2
10 A ASN 31 2HD2
10 A GLN 43 1HE2
10 A GLN 43 2HE2
10 A GLN 45 1HE2
10 A GLN 45 2HE2
10 A ASN 49 1HD2
10 A ASN 49 2HD2
10 A GLN 57 1HE2
10 A GLN 57 2HE2
10 A ASN 64 1HD2
10 A ASN 64 2HD2
10 B ASN 13 1HD2
10 B ASN 13 2HD2
10 B GLN 16 1HE2
10 B GLN 16 2HE2
10 B ASN 21 1HD2
10 B ASN 21 2HD2
10 B GLN 22 1HE2
10 B GLN 22 2HE2
10 B ASN 31 1HD2
10 B ASN 31 2HD2
10 B GLN 43 1HE2
10 B GLN 43 2HE2
10 B GLN 45 1HE2
10 B GLN 45 2HE2
10 B ASN 49 1HD2
10 B ASN 49 2HD2
10 B GLN 57 1HE2
10 B GLN 57 2HE2
10 B ASN 64 1HD2
10 B ASN 64 2HD2
11 A ASN 13 1HD2
11 A ASN 13 2HD2
11 A GLN 16 1HE2
11 A GLN 16 2HE2
11 A ASN 21 1HD2
11 A ASN 21 2HD2
11 A GLN 22 1HE2
11 A GLN 22 2HE2
11 A ASN 31 1HD2
11 A ASN 31 2HD2
11 A GLN 43 1HE2
11 A GLN 43 2HE2
11 A GLN 45 1HE2
11 A GLN 45 2HE2
11 A ASN 49 1HD2
11 A ASN 49 2HD2
11 A GLN 57 1HE2
11 A GLN 57 2HE2
11 A ASN 64 1HD2
11 A ASN 64 2HD2
11 B ASN 13 1HD2
11 B ASN 13 2HD2
11 B GLN 16 1HE2
11 B GLN 16 2HE2
11 B ASN 21 1HD2
11 B ASN 21 2HD2
11 B GLN 22 1HE2
11 B GLN 22 2HE2
11 B ASN 31 1HD2
11 B ASN 31 2HD2
11 B GLN 43 1HE2
11 B GLN 43 2HE2
11 B GLN 45 1HE2
11 B GLN 45 2HE2
11 B ASN 49 1HD2
11 B ASN 49 2HD2
11 B GLN 57 1HE2
11 B GLN 57 2HE2
11 B ASN 64 1HD2
11 B ASN 64 2HD2
12 A ASN 13 1HD2
12 A ASN 13 2HD2
12 A GLN 16 1HE2
12 A GLN 16 2HE2
12 A ASN 21 1HD2
12 A ASN 21 2HD2
12 A GLN 22 1HE2
12 A GLN 22 2HE2
12 A ASN 31 1HD2
12 A ASN 31 2HD2
12 A GLN 43 1HE2
12 A GLN 43 2HE2
12 A GLN 45 1HE2
12 A GLN 45 2HE2
12 A ASN 49 1HD2
12 A ASN 49 2HD2
12 A GLN 57 1HE2
12 A GLN 57 2HE2
12 A ASN 64 1HD2
12 A ASN 64 2HD2
12 B ASN 13 1HD2
12 B ASN 13 2HD2
12 B GLN 16 1HE2
12 B GLN 16 2HE2
12 B ASN 21 1HD2
12 B ASN 21 2HD2
12 B GLN 22 1HE2
12 B GLN 22 2HE2
12 B ASN 31 1HD2
12 B ASN 31 2HD2
12 B GLN 43 1HE2
12 B GLN 43 2HE2
12 B GLN 45 1HE2
12 B GLN 45 2HE2
12 B ASN 49 1HD2
12 B ASN 49 2HD2
12 B GLN 57 1HE2
12 B GLN 57 2HE2
12 B ASN 64 1HD2
12 B ASN 64 2HD2
13 A ASN 13 1HD2
13 A ASN 13 2HD2
13 A GLN 16 1HE2
13 A GLN 16 2HE2
13 A ASN 21 1HD2
13 A ASN 21 2HD2
13 A GLN 22 1HE2
13 A GLN 22 2HE2
13 A ASN 31 1HD2
13 A ASN 31 2HD2
13 A GLN 43 1HE2
13 A GLN 43 2HE2
13 A GLN 45 1HE2
13 A GLN 45 2HE2
13 A ASN 49 1HD2
13 A ASN 49 2HD2
13 A GLN 57 1HE2
13 A GLN 57 2HE2
13 A ASN 64 1HD2
13 A ASN 64 2HD2
13 B ASN 13 1HD2
13 B ASN 13 2HD2
13 B GLN 16 1HE2
13 B GLN 16 2HE2
13 B ASN 21 1HD2
13 B ASN 21 2HD2
13 B GLN 22 1HE2
13 B GLN 22 2HE2
13 B ASN 31 1HD2
13 B ASN 31 2HD2
13 B GLN 43 1HE2
13 B GLN 43 2HE2
13 B GLN 45 1HE2
13 B GLN 45 2HE2
13 B ASN 49 1HD2
13 B ASN 49 2HD2
13 B GLN 57 1HE2
13 B GLN 57 2HE2
13 B ASN 64 1HD2
13 B ASN 64 2HD2
14 A ASN 13 1HD2
14 A ASN 13 2HD2
14 A GLN 16 1HE2
14 A GLN 16 2HE2
14 A ASN 21 1HD2
14 A ASN 21 2HD2
14 A GLN 22 1HE2
14 A GLN 22 2HE2
14 A ASN 31 1HD2
14 A ASN 31 2HD2
14 A GLN 43 1HE2
14 A GLN 43 2HE2
14 A GLN 45 1HE2
14 A GLN 45 2HE2
14 A ASN 49 1HD2
14 A ASN 49 2HD2
14 A GLN 57 1HE2
14 A GLN 57 2HE2
14 A ASN 64 1HD2
14 A ASN 64 2HD2
14 B ASN 13 1HD2
14 B ASN 13 2HD2
14 B GLN 16 1HE2
14 B GLN 16 2HE2
14 B ASN 21 1HD2
14 B ASN 21 2HD2
14 B GLN 22 1HE2
14 B GLN 22 2HE2
14 B ASN 31 1HD2
14 B ASN 31 2HD2
14 B GLN 43 1HE2
14 B GLN 43 2HE2
14 B GLN 45 1HE2
14 B GLN 45 2HE2
14 B ASN 49 1HD2
14 B ASN 49 2HD2
14 B GLN 57 1HE2
14 B GLN 57 2HE2
14 B ASN 64 1HD2
14 B ASN 64 2HD2
15 A ASN 13 1HD2
15 A ASN 13 2HD2
15 A GLN 16 1HE2
15 A GLN 16 2HE2
15 A ASN 21 1HD2
15 A ASN 21 2HD2
15 A GLN 22 1HE2
15 A GLN 22 2HE2
15 A ASN 31 1HD2
15 A ASN 31 2HD2
15 A GLN 43 1HE2
15 A GLN 43 2HE2
15 A GLN 45 1HE2
15 A GLN 45 2HE2
15 A ASN 49 1HD2
15 A ASN 49 2HD2
15 A GLN 57 1HE2
15 A GLN 57 2HE2
15 A ASN 64 1HD2
15 A ASN 64 2HD2
15 B ASN 13 1HD2
15 B ASN 13 2HD2
15 B GLN 16 1HE2
15 B GLN 16 2HE2
15 B ASN 21 1HD2
15 B ASN 21 2HD2
15 B GLN 22 1HE2
15 B GLN 22 2HE2
15 B ASN 31 1HD2
15 B ASN 31 2HD2
15 B GLN 43 1HE2
15 B GLN 43 2HE2
15 B GLN 45 1HE2
15 B GLN 45 2HE2
15 B ASN 49 1HD2
15 B ASN 49 2HD2
15 B GLN 57 1HE2
15 B GLN 57 2HE2
15 B ASN 64 1HD2
15 B ASN 64 2HD2
16 A ASN 13 1HD2
16 A ASN 13 2HD2
16 A GLN 16 1HE2
16 A GLN 16 2HE2
16 A ASN 21 1HD2
16 A ASN 21 2HD2
16 A GLN 22 1HE2
16 A GLN 22 2HE2
16 A ASN 31 1HD2
16 A ASN 31 2HD2
16 A GLN 43 1HE2
16 A GLN 43 2HE2
16 A GLN 45 1HE2
16 A GLN 45 2HE2
16 A ASN 49 1HD2
16 A ASN 49 2HD2
16 A GLN 57 1HE2
16 A GLN 57 2HE2
16 A ASN 64 1HD2
16 A ASN 64 2HD2
16 B ASN 13 1HD2
16 B ASN 13 2HD2
16 B GLN 16 1HE2
16 B GLN 16 2HE2
16 B ASN 21 1HD2
16 B ASN 21 2HD2
16 B GLN 22 1HE2
16 B GLN 22 2HE2
16 B ASN 31 1HD2
16 B ASN 31 2HD2
16 B GLN 43 1HE2
16 B GLN 43 2HE2
16 B GLN 45 1HE2
16 B GLN 45 2HE2
16 B ASN 49 1HD2
16 B ASN 49 2HD2
16 B GLN 57 1HE2
16 B GLN 57 2HE2
16 B ASN 64 1HD2
16 B ASN 64 2HD2
17 A ASN 13 1HD2
17 A ASN 13 2HD2
17 A GLN 16 1HE2
17 A GLN 16 2HE2
17 A ASN 21 1HD2
17 A ASN 21 2HD2
17 A GLN 22 1HE2
17 A GLN 22 2HE2
17 A ASN 31 1HD2
17 A ASN 31 2HD2
17 A GLN 43 1HE2
17 A GLN 43 2HE2
17 A GLN 45 1HE2
17 A GLN 45 2HE2
17 A ASN 49 1HD2
17 A ASN 49 2HD2
17 A GLN 57 1HE2
17 A GLN 57 2HE2
17 A ASN 64 1HD2
17 A ASN 64 2HD2
17 B ASN 13 1HD2
17 B ASN 13 2HD2
17 B GLN 16 1HE2
17 B GLN 16 2HE2
17 B ASN 21 1HD2
17 B ASN 21 2HD2
17 B GLN 22 1HE2
17 B GLN 22 2HE2
17 B ASN 31 1HD2
17 B ASN 31 2HD2
17 B GLN 43 1HE2
17 B GLN 43 2HE2
17 B GLN 45 1HE2
17 B GLN 45 2HE2
17 B ASN 49 1HD2
17 B ASN 49 2HD2
17 B GLN 57 1HE2
17 B GLN 57 2HE2
17 B ASN 64 1HD2
17 B ASN 64 2HD2
18 A ASN 13 1HD2
18 A ASN 13 2HD2
18 A GLN 16 1HE2
18 A GLN 16 2HE2
18 A ASN 21 1HD2
18 A ASN 21 2HD2
18 A GLN 22 1HE2
18 A GLN 22 2HE2
18 A ASN 31 1HD2
18 A ASN 31 2HD2
18 A GLN 43 1HE2
18 A GLN 43 2HE2
18 A GLN 45 1HE2
18 A GLN 45 2HE2
18 A ASN 49 1HD2
18 A ASN 49 2HD2
18 A GLN 57 1HE2
18 A GLN 57 2HE2
18 A ASN 64 1HD2
18 A ASN 64 2HD2
18 B ASN 13 1HD2
18 B ASN 13 2HD2
18 B GLN 16 1HE2
18 B GLN 16 2HE2
18 B ASN 21 1HD2
18 B ASN 21 2HD2
18 B GLN 22 1HE2
18 B GLN 22 2HE2
18 B ASN 31 1HD2
18 B ASN 31 2HD2
18 B GLN 43 1HE2
18 B GLN 43 2HE2
18 B GLN 45 1HE2
18 B GLN 45 2HE2
18 B ASN 49 1HD2
18 B ASN 49 2HD2
18 B GLN 57 1HE2
18 B GLN 57 2HE2
18 B ASN 64 1HD2
18 B ASN 64 2HD2
19 A ASN 13 1HD2
19 A ASN 13 2HD2
19 A GLN 16 1HE2
19 A GLN 16 2HE2
19 A ASN 21 1HD2
19 A ASN 21 2HD2
19 A GLN 22 1HE2
19 A GLN 22 2HE2
19 A ASN 31 1HD2
19 A ASN 31 2HD2
19 A GLN 43 1HE2
19 A GLN 43 2HE2
19 A GLN 45 1HE2
19 A GLN 45 2HE2
19 A ASN 49 1HD2
19 A ASN 49 2HD2
19 A GLN 57 1HE2
19 A GLN 57 2HE2
19 A ASN 64 1HD2
19 A ASN 64 2HD2
19 B ASN 13 1HD2
19 B ASN 13 2HD2
19 B GLN 16 1HE2
19 B GLN 16 2HE2
19 B ASN 21 1HD2
19 B ASN 21 2HD2
19 B GLN 22 1HE2
19 B GLN 22 2HE2
19 B ASN 31 1HD2
19 B ASN 31 2HD2
19 B GLN 43 1HE2
19 B GLN 43 2HE2
19 B GLN 45 1HE2
19 B GLN 45 2HE2
19 B ASN 49 1HD2
19 B ASN 49 2HD2
19 B GLN 57 1HE2
19 B GLN 57 2HE2
19 B ASN 64 1HD2
19 B ASN 64 2HD2
20 A ASN 13 1HD2
20 A ASN 13 2HD2
20 A GLN 16 1HE2
20 A GLN 16 2HE2
20 A ASN 21 1HD2
20 A ASN 21 2HD2
20 A GLN 22 1HE2
20 A GLN 22 2HE2
20 A ASN 31 1HD2
20 A ASN 31 2HD2
20 A GLN 43 1HE2
20 A GLN 43 2HE2
20 A GLN 45 1HE2
20 A GLN 45 2HE2
20 A ASN 49 1HD2
20 A ASN 49 2HD2
20 A GLN 57 1HE2
20 A GLN 57 2HE2
20 A ASN 64 1HD2
20 A ASN 64 2HD2
20 B ASN 13 1HD2
20 B ASN 13 2HD2
20 B GLN 16 1HE2
20 B GLN 16 2HE2
20 B ASN 21 1HD2
20 B ASN 21 2HD2
20 B GLN 22 1HE2
20 B GLN 22 2HE2
20 B ASN 31 1HD2
20 B ASN 31 2HD2
20 B GLN 43 1HE2
20 B GLN 43 2HE2
20 B GLN 45 1HE2
20 B GLN 45 2HE2
20 B ASN 49 1HD2
20 B ASN 49 2HD2
20 B GLN 57 1HE2
20 B GLN 57 2HE2
20 B ASN 64 1HD2
20 B ASN 64 2HD2
OTHER IMPORTANT ISSUES
==> The following residues have missing atoms:
RES MOD#C SEQ ATOMS
GLU( 1 A 3) HE2
GLU( 1 A 12) HE2
GLU( 1 A 19) HE2
GLU( 1 A 24) HE2
ASP( 1 A 25) HD2
ASP( 1 A 44) HD2
ASP( 1 A 60) HD2
GLU( 1 A 61) HE2
GLU( 1 A 70) HE2
GLU( 1 A 74) HE2
GLU( 1 B 3) HE2
GLU( 1 B 12) HE2
GLU( 1 B 19) HE2
GLU( 1 B 24) HE2
ASP( 1 B 25) HD2
ASP( 1 B 44) HD2
ASP( 1 B 60) HD2
GLU( 1 B 61) HE2
GLU( 1 B 70) HE2
GLU( 1 B 74) HE2
GLU( 2 A 3) HE2
GLU( 2 A 12) HE2
GLU( 2 A 19) HE2
GLU( 2 A 24) HE2
ASP( 2 A 25) HD2
ASP( 2 A 44) HD2
ASP( 2 A 60) HD2
GLU( 2 A 61) HE2
GLU( 2 A 70) HE2
GLU( 2 A 74) HE2
GLU( 2 B 3) HE2
GLU( 2 B 12) HE2
GLU( 2 B 19) HE2
GLU( 2 B 24) HE2
ASP( 2 B 25) HD2
ASP( 2 B 44) HD2
ASP( 2 B 60) HD2
GLU( 2 B 61) HE2
GLU( 2 B 70) HE2
GLU( 2 B 74) HE2
GLU( 3 A 3) HE2
GLU( 3 A 12) HE2
GLU( 3 A 19) HE2
GLU( 3 A 24) HE2
ASP( 3 A 25) HD2
ASP( 3 A 44) HD2
ASP( 3 A 60) HD2
GLU( 3 A 61) HE2
GLU( 3 A 70) HE2
GLU( 3 A 74) HE2
GLU( 3 B 3) HE2
GLU( 3 B 12) HE2
GLU( 3 B 19) HE2
GLU( 3 B 24) HE2
ASP( 3 B 25) HD2
ASP( 3 B 44) HD2
ASP( 3 B 60) HD2
GLU( 3 B 61) HE2
GLU( 3 B 70) HE2
GLU( 3 B 74) HE2
GLU( 4 A 3) HE2
GLU( 4 A 12) HE2
GLU( 4 A 19) HE2
GLU( 4 A 24) HE2
ASP( 4 A 25) HD2
ASP( 4 A 44) HD2
ASP( 4 A 60) HD2
GLU( 4 A 61) HE2
GLU( 4 A 70) HE2
GLU( 4 A 74) HE2
GLU( 4 B 3) HE2
GLU( 4 B 12) HE2
GLU( 4 B 19) HE2
GLU( 4 B 24) HE2
ASP( 4 B 25) HD2
ASP( 4 B 44) HD2
ASP( 4 B 60) HD2
GLU( 4 B 61) HE2
GLU( 4 B 70) HE2
GLU( 4 B 74) HE2
GLU( 5 A 3) HE2
GLU( 5 A 12) HE2
GLU( 5 A 19) HE2
GLU( 5 A 24) HE2
ASP( 5 A 25) HD2
ASP( 5 A 44) HD2
ASP( 5 A 60) HD2
GLU( 5 A 61) HE2
GLU( 5 A 70) HE2
GLU( 5 A 74) HE2
GLU( 5 B 3) HE2
GLU( 5 B 12) HE2
GLU( 5 B 19) HE2
GLU( 5 B 24) HE2
ASP( 5 B 25) HD2
ASP( 5 B 44) HD2
ASP( 5 B 60) HD2
GLU( 5 B 61) HE2
GLU( 5 B 70) HE2
GLU( 5 B 74) HE2
GLU( 6 A 3) HE2
GLU( 6 A 12) HE2
GLU( 6 A 19) HE2
GLU( 6 A 24) HE2
ASP( 6 A 25) HD2
ASP( 6 A 44) HD2
ASP( 6 A 60) HD2
GLU( 6 A 61) HE2
GLU( 6 A 70) HE2
GLU( 6 A 74) HE2
GLU( 6 B 3) HE2
GLU( 6 B 12) HE2
GLU( 6 B 19) HE2
GLU( 6 B 24) HE2
ASP( 6 B 25) HD2
ASP( 6 B 44) HD2
ASP( 6 B 60) HD2
GLU( 6 B 61) HE2
GLU( 6 B 70) HE2
GLU( 6 B 74) HE2
GLU( 7 A 3) HE2
GLU( 7 A 12) HE2
GLU( 7 A 19) HE2
GLU( 7 A 24) HE2
ASP( 7 A 25) HD2
ASP( 7 A 44) HD2
ASP( 7 A 60) HD2
GLU( 7 A 61) HE2
GLU( 7 A 70) HE2
GLU( 7 A 74) HE2
GLU( 7 B 3) HE2
GLU( 7 B 12) HE2
GLU( 7 B 19) HE2
GLU( 7 B 24) HE2
ASP( 7 B 25) HD2
ASP( 7 B 44) HD2
ASP( 7 B 60) HD2
GLU( 7 B 61) HE2
GLU( 7 B 70) HE2
GLU( 7 B 74) HE2
GLU( 8 A 3) HE2
GLU( 8 A 12) HE2
GLU( 8 A 19) HE2
GLU( 8 A 24) HE2
ASP( 8 A 25) HD2
ASP( 8 A 44) HD2
ASP( 8 A 60) HD2
GLU( 8 A 61) HE2
GLU( 8 A 70) HE2
GLU( 8 A 74) HE2
GLU( 8 B 3) HE2
GLU( 8 B 12) HE2
GLU( 8 B 19) HE2
GLU( 8 B 24) HE2
ASP( 8 B 25) HD2
ASP( 8 B 44) HD2
ASP( 8 B 60) HD2
GLU( 8 B 61) HE2
GLU( 8 B 70) HE2
GLU( 8 B 74) HE2
GLU( 9 A 3) HE2
GLU( 9 A 12) HE2
GLU( 9 A 19) HE2
GLU( 9 A 24) HE2
ASP( 9 A 25) HD2
ASP( 9 A 44) HD2
ASP( 9 A 60) HD2
GLU( 9 A 61) HE2
GLU( 9 A 70) HE2
GLU( 9 A 74) HE2
GLU( 9 B 3) HE2
GLU( 9 B 12) HE2
GLU( 9 B 19) HE2
GLU( 9 B 24) HE2
ASP( 9 B 25) HD2
ASP( 9 B 44) HD2
ASP( 9 B 60) HD2
GLU( 9 B 61) HE2
GLU( 9 B 70) HE2
GLU( 9 B 74) HE2
GLU( 10 A 3) HE2
GLU( 10 A 12) HE2
GLU( 10 A 19) HE2
GLU( 10 A 24) HE2
ASP( 10 A 25) HD2
ASP( 10 A 44) HD2
ASP( 10 A 60) HD2
GLU( 10 A 61) HE2
GLU( 10 A 70) HE2
GLU( 10 A 74) HE2
GLU( 10 B 3) HE2
GLU( 10 B 12) HE2
GLU( 10 B 19) HE2
GLU( 10 B 24) HE2
ASP( 10 B 25) HD2
ASP( 10 B 44) HD2
ASP( 10 B 60) HD2
GLU( 10 B 61) HE2
GLU( 10 B 70) HE2
GLU( 10 B 74) HE2
GLU( 11 A 3) HE2
GLU( 11 A 12) HE2
GLU( 11 A 19) HE2
GLU( 11 A 24) HE2
ASP( 11 A 25) HD2
ASP( 11 A 44) HD2
ASP( 11 A 60) HD2
GLU( 11 A 61) HE2
GLU( 11 A 70) HE2
GLU( 11 A 74) HE2
GLU( 11 B 3) HE2
GLU( 11 B 12) HE2
GLU( 11 B 19) HE2
GLU( 11 B 24) HE2
ASP( 11 B 25) HD2
ASP( 11 B 44) HD2
ASP( 11 B 60) HD2
GLU( 11 B 61) HE2
GLU( 11 B 70) HE2
GLU( 11 B 74) HE2
GLU( 12 A 3) HE2
GLU( 12 A 12) HE2
GLU( 12 A 19) HE2
GLU( 12 A 24) HE2
ASP( 12 A 25) HD2
ASP( 12 A 44) HD2
ASP( 12 A 60) HD2
GLU( 12 A 61) HE2
GLU( 12 A 70) HE2
GLU( 12 A 74) HE2
GLU( 12 B 3) HE2
GLU( 12 B 12) HE2
GLU( 12 B 19) HE2
GLU( 12 B 24) HE2
ASP( 12 B 25) HD2
ASP( 12 B 44) HD2
ASP( 12 B 60) HD2
GLU( 12 B 61) HE2
GLU( 12 B 70) HE2
GLU( 12 B 74) HE2
GLU( 13 A 3) HE2
GLU( 13 A 12) HE2
GLU( 13 A 19) HE2
GLU( 13 A 24) HE2
ASP( 13 A 25) HD2
ASP( 13 A 44) HD2
ASP( 13 A 60) HD2
GLU( 13 A 61) HE2
GLU( 13 A 70) HE2
GLU( 13 A 74) HE2
GLU( 13 B 3) HE2
GLU( 13 B 12) HE2
GLU( 13 B 19) HE2
GLU( 13 B 24) HE2
ASP( 13 B 25) HD2
ASP( 13 B 44) HD2
ASP( 13 B 60) HD2
GLU( 13 B 61) HE2
GLU( 13 B 70) HE2
GLU( 13 B 74) HE2
GLU( 14 A 3) HE2
GLU( 14 A 12) HE2
GLU( 14 A 19) HE2
GLU( 14 A 24) HE2
ASP( 14 A 25) HD2
ASP( 14 A 44) HD2
ASP( 14 A 60) HD2
GLU( 14 A 61) HE2
GLU( 14 A 70) HE2
GLU( 14 A 74) HE2
GLU( 14 B 3) HE2
GLU( 14 B 12) HE2
GLU( 14 B 19) HE2
GLU( 14 B 24) HE2
ASP( 14 B 25) HD2
ASP( 14 B 44) HD2
ASP( 14 B 60) HD2
GLU( 14 B 61) HE2
GLU( 14 B 70) HE2
GLU( 14 B 74) HE2
GLU( 15 A 3) HE2
GLU( 15 A 12) HE2
GLU( 15 A 19) HE2
GLU( 15 A 24) HE2
ASP( 15 A 25) HD2
ASP( 15 A 44) HD2
ASP( 15 A 60) HD2
GLU( 15 A 61) HE2
GLU( 15 A 70) HE2
GLU( 15 A 74) HE2
GLU( 15 B 3) HE2
GLU( 15 B 12) HE2
GLU( 15 B 19) HE2
GLU( 15 B 24) HE2
ASP( 15 B 25) HD2
ASP( 15 B 44) HD2
ASP( 15 B 60) HD2
GLU( 15 B 61) HE2
GLU( 15 B 70) HE2
GLU( 15 B 74) HE2
GLU( 16 A 3) HE2
GLU( 16 A 12) HE2
GLU( 16 A 19) HE2
GLU( 16 A 24) HE2
ASP( 16 A 25) HD2
ASP( 16 A 44) HD2
ASP( 16 A 60) HD2
GLU( 16 A 61) HE2
GLU( 16 A 70) HE2
GLU( 16 A 74) HE2
GLU( 16 B 3) HE2
GLU( 16 B 12) HE2
GLU( 16 B 19) HE2
GLU( 16 B 24) HE2
ASP( 16 B 25) HD2
ASP( 16 B 44) HD2
ASP( 16 B 60) HD2
GLU( 16 B 61) HE2
GLU( 16 B 70) HE2
GLU( 16 B 74) HE2
GLU( 17 A 3) HE2
GLU( 17 A 12) HE2
GLU( 17 A 19) HE2
GLU( 17 A 24) HE2
ASP( 17 A 25) HD2
ASP( 17 A 44) HD2
ASP( 17 A 60) HD2
GLU( 17 A 61) HE2
GLU( 17 A 70) HE2
GLU( 17 A 74) HE2
GLU( 17 B 3) HE2
GLU( 17 B 12) HE2
GLU( 17 B 19) HE2
GLU( 17 B 24) HE2
ASP( 17 B 25) HD2
ASP( 17 B 44) HD2
ASP( 17 B 60) HD2
GLU( 17 B 61) HE2
GLU( 17 B 70) HE2
GLU( 17 B 74) HE2
GLU( 18 A 3) HE2
GLU( 18 A 12) HE2
GLU( 18 A 19) HE2
GLU( 18 A 24) HE2
ASP( 18 A 25) HD2
ASP( 18 A 44) HD2
ASP( 18 A 60) HD2
GLU( 18 A 61) HE2
GLU( 18 A 70) HE2
GLU( 18 A 74) HE2
GLU( 18 B 3) HE2
GLU( 18 B 12) HE2
GLU( 18 B 19) HE2
GLU( 18 B 24) HE2
ASP( 18 B 25) HD2
ASP( 18 B 44) HD2
ASP( 18 B 60) HD2
GLU( 18 B 61) HE2
GLU( 18 B 70) HE2
GLU( 18 B 74) HE2
GLU( 19 A 3) HE2
GLU( 19 A 12) HE2
GLU( 19 A 19) HE2
GLU( 19 A 24) HE2
ASP( 19 A 25) HD2
ASP( 19 A 44) HD2
ASP( 19 A 60) HD2
GLU( 19 A 61) HE2
GLU( 19 A 70) HE2
GLU( 19 A 74) HE2
GLU( 19 B 3) HE2
GLU( 19 B 12) HE2
GLU( 19 B 19) HE2
GLU( 19 B 24) HE2
ASP( 19 B 25) HD2
ASP( 19 B 44) HD2
ASP( 19 B 60) HD2
GLU( 19 B 61) HE2
GLU( 19 B 70) HE2
GLU( 19 B 74) HE2
GLU( 20 A 3) HE2
GLU( 20 A 12) HE2
GLU( 20 A 19) HE2
GLU( 20 A 24) HE2
ASP( 20 A 25) HD2
ASP( 20 A 44) HD2
ASP( 20 A 60) HD2
GLU( 20 A 61) HE2
GLU( 20 A 70) HE2
GLU( 20 A 74) HE2
GLU( 20 B 3) HE2
GLU( 20 B 12) HE2
GLU( 20 B 19) HE2
GLU( 20 B 24) HE2
ASP( 20 B 25) HD2
ASP( 20 B 44) HD2
ASP( 20 B 60) HD2
GLU( 20 B 61) HE2
GLU( 20 B 70) HE2
GLU( 20 B 74) HE2





























