Detailed results of HR41_XRay_em_bcr3 by PSVS
Output from PDBStat
Output from PROCHECK
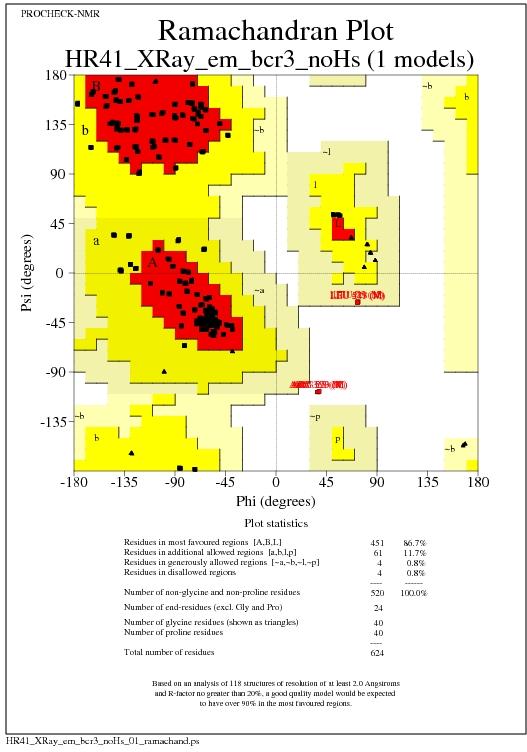
Ramachandran Plot for all models
Text summary of Ramachandran Plot
+----------<<< P R O C H E C K S U M M A R Y >>>----------+
| |
| HR41_XRay_em_bcr3_noHs_000.rin 0.0 624 residues |
| |
*| Ramachandran plot: 86.7% core 11.7% allow 0.8% gener 0.8% disall |
| |
*| All Ramachandrans: 20 labelled residues (out of 600) |
+| Chi1-chi2 plots: 12 labelled residues (out of 416) |
JPEG image for all model Ramachandran Plot
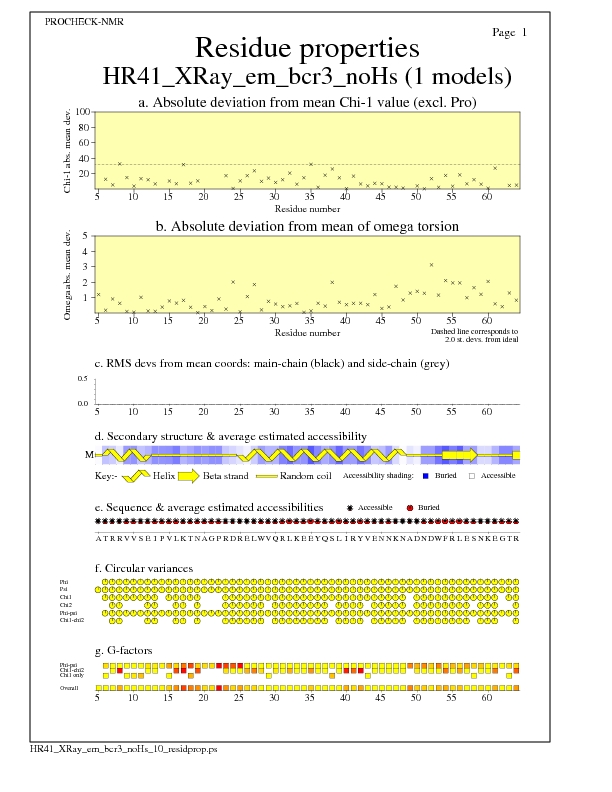
Residue Properties for all models
JPEG for all model Residue Properties - page $num_n
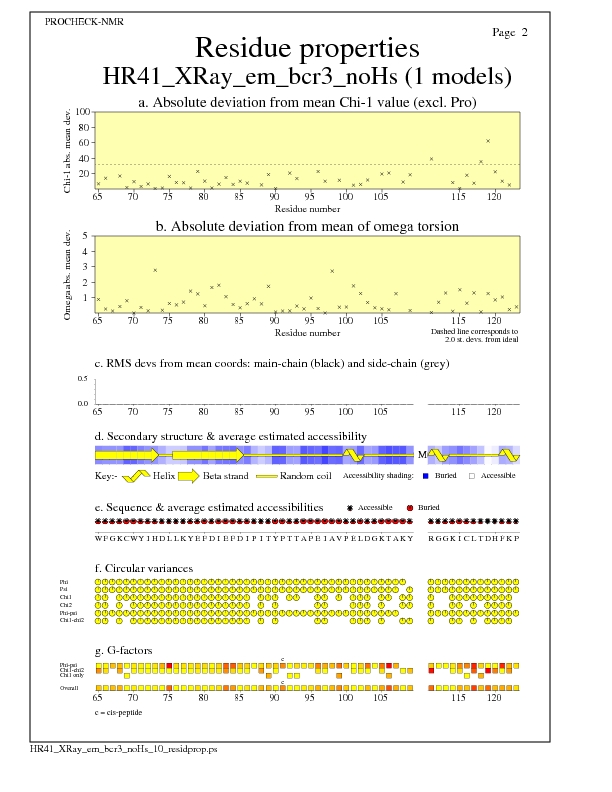
JPEG for all model Residue Properties - page $num_n
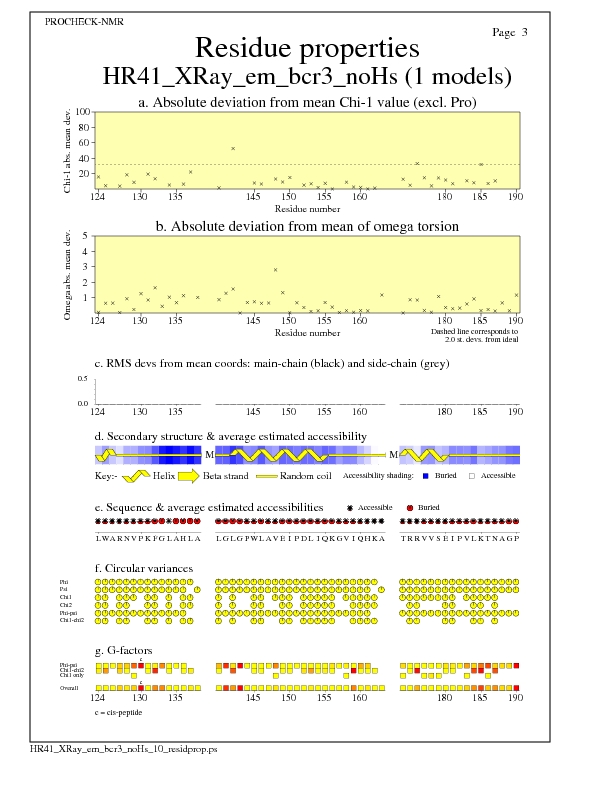
JPEG for all model Residue Properties - page $num_n
JPEG for all model Residue Properties - page $num_n
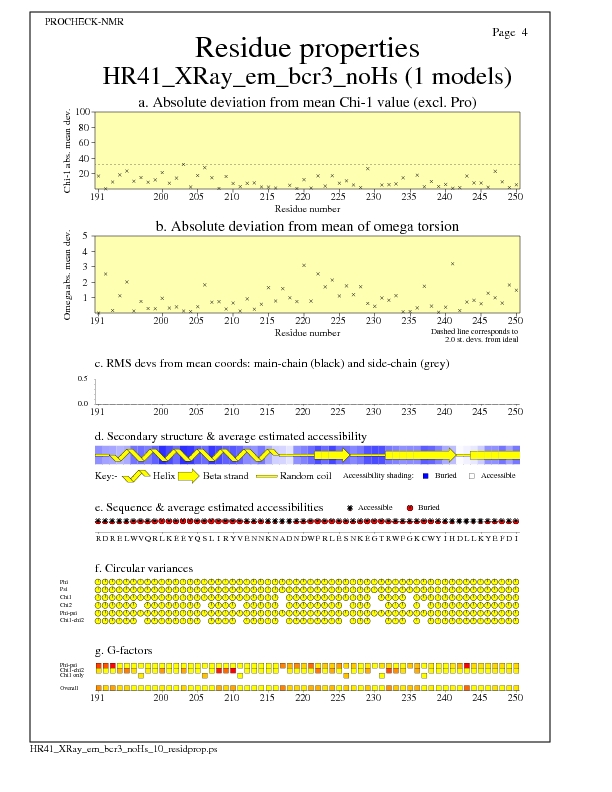
JPEG for all model Residue Properties - page $num_n
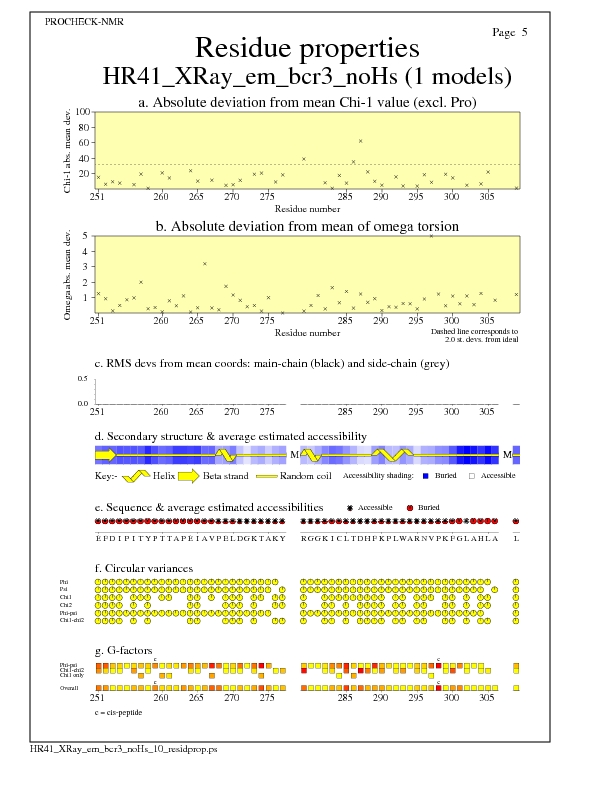
JPEG for all model Residue Properties - page $num_n
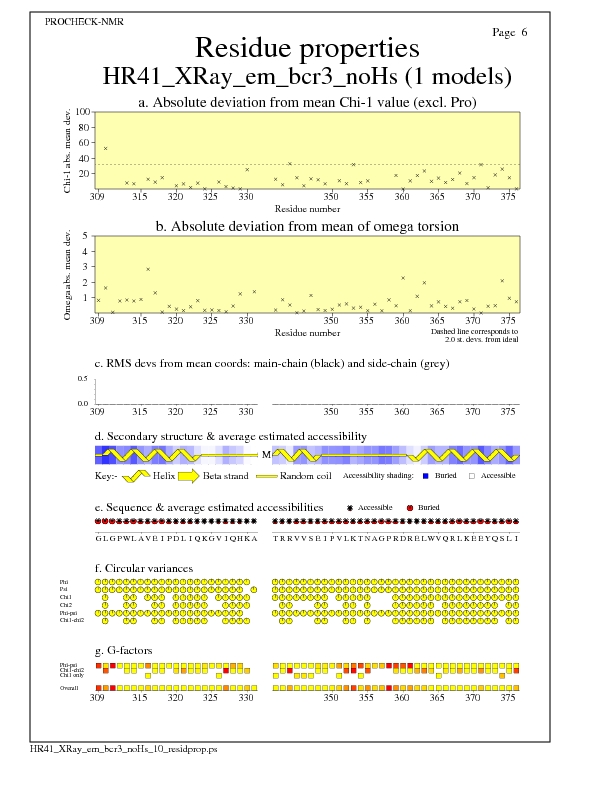
JPEG for all model Residue Properties - page $num_n
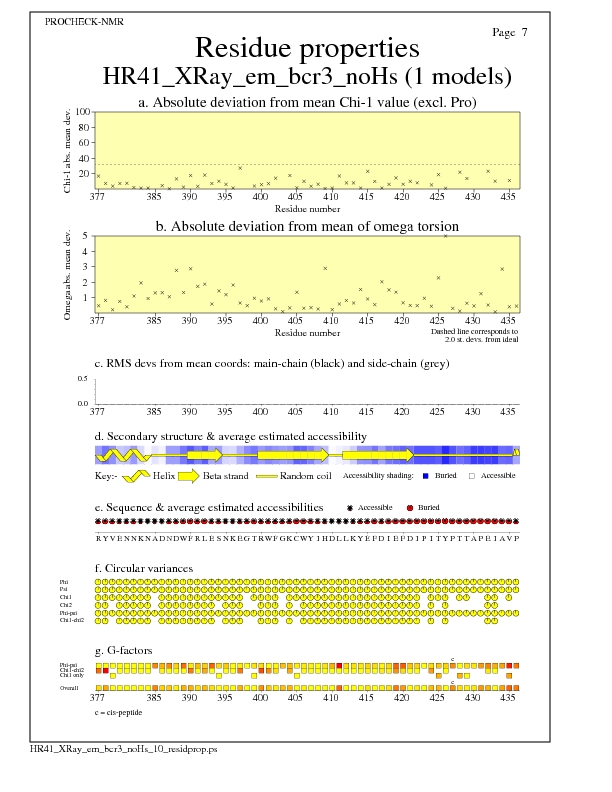
JPEG for all model Residue Properties - page $num_n
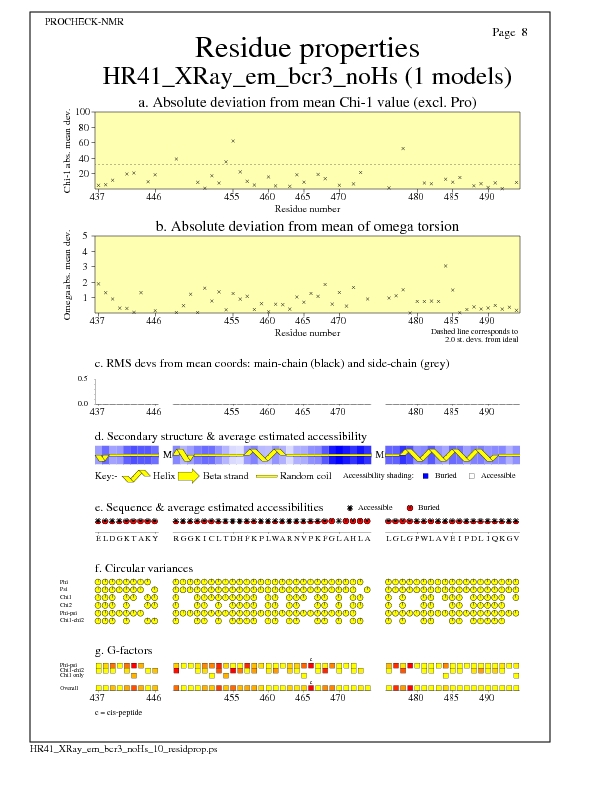
JPEG for all model Residue Properties - page $num_n
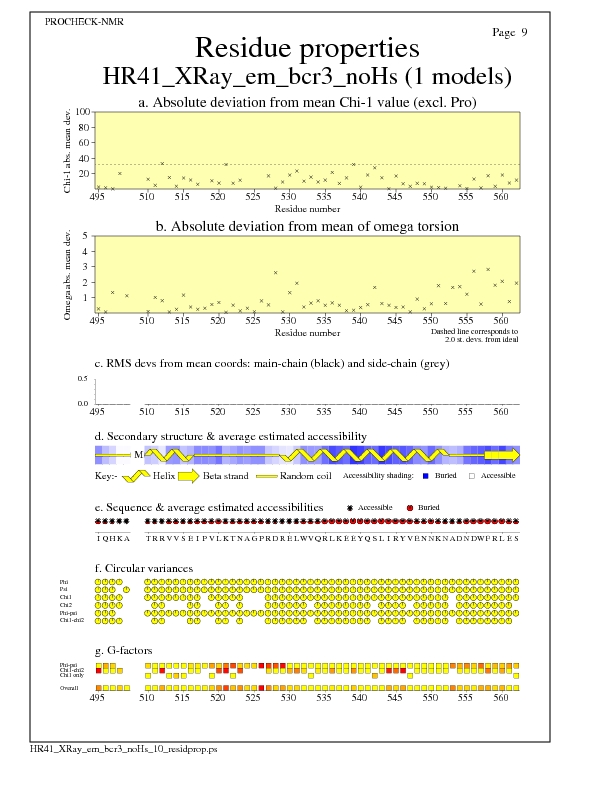
JPEG for all model Residue Properties - page $num_n
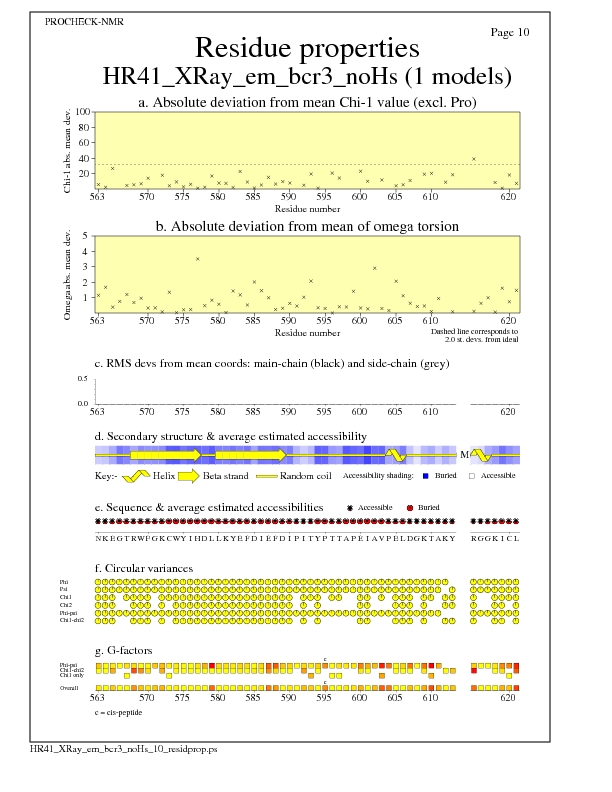
JPEG for all model Residue Properties - page $num_n
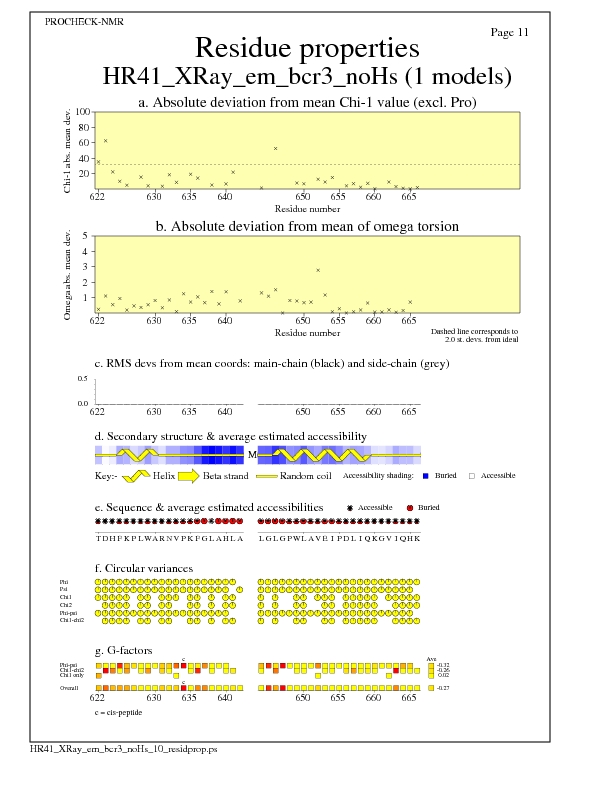
Model Secondary Structures from Procheck
JPEG for Model Secondary Structures - page $num_n

JPEG for Model Secondary Structures - page $num_n

JPEG for Model Secondary Structures - page $num_n

JPEG for Model Secondary Structures - page $num_n
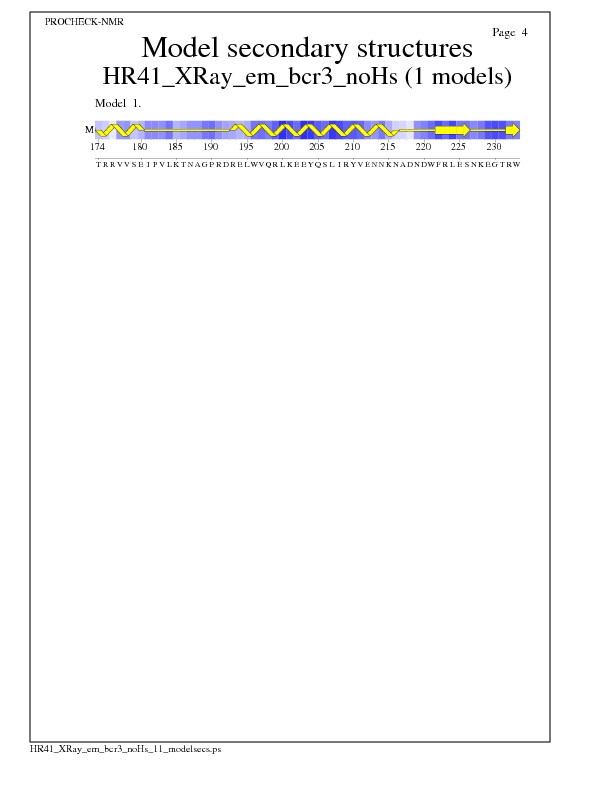

JPEG for Model Secondary Structures - page $num_n
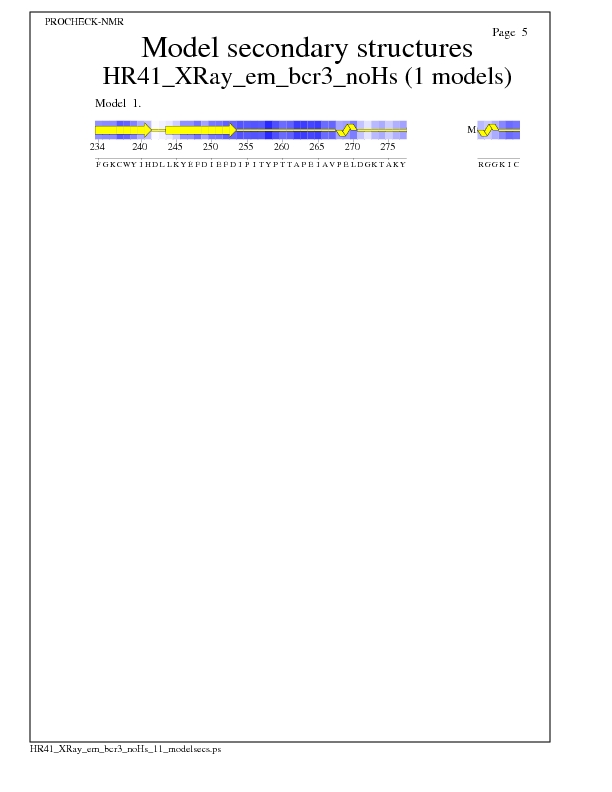

JPEG for Model Secondary Structures - page $num_n

JPEG for Model Secondary Structures - page $num_n
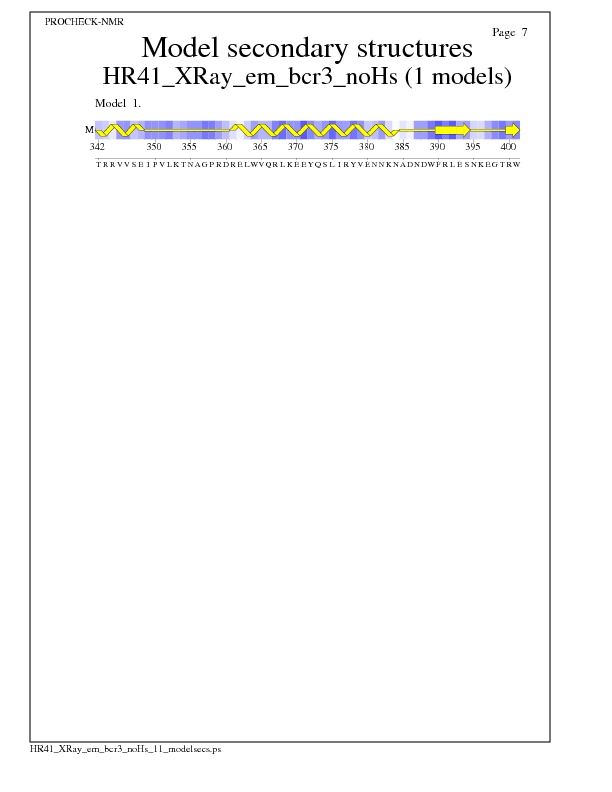

JPEG for Model Secondary Structures - page $num_n
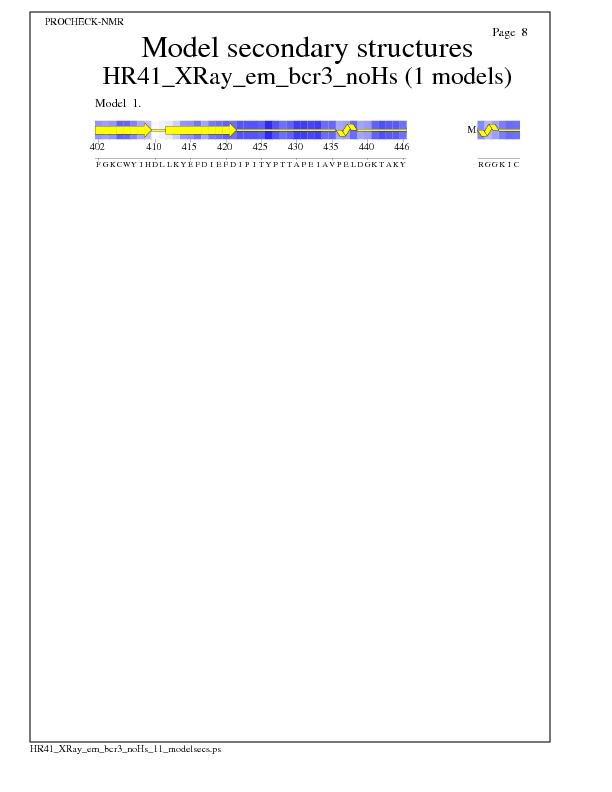

JPEG for Model Secondary Structures - page $num_n

JPEG for Model Secondary Structures - page $num_n
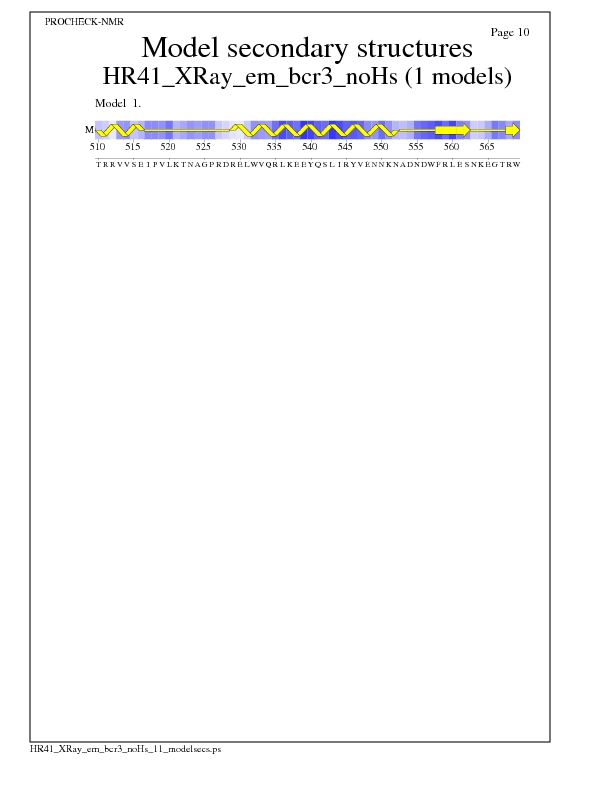

JPEG for Model Secondary Structures - page $num_n
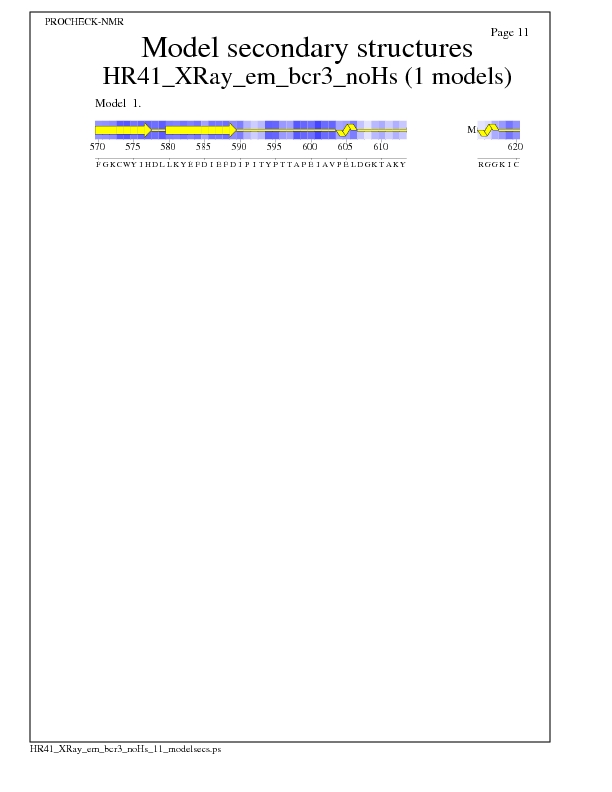

JPEG for Model Secondary Structures - page $num_n
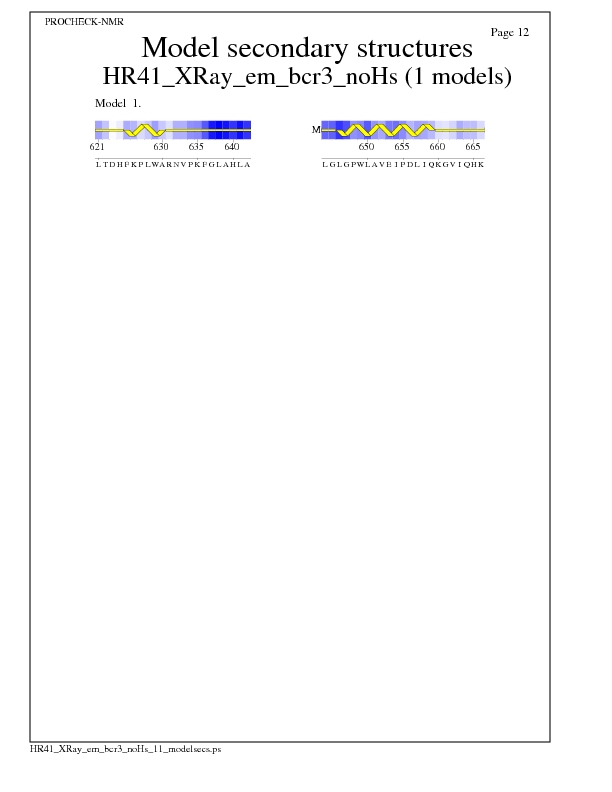

Ramachandran Plots for each residue
JPEG for residue Ramachandran Plots - page $num_n
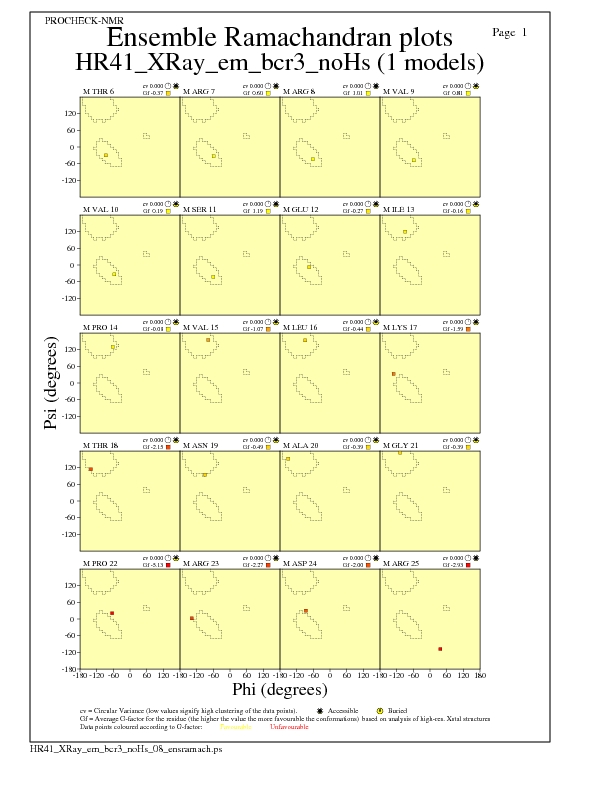
JPEG for residue Ramachandran Plots - page $num_n
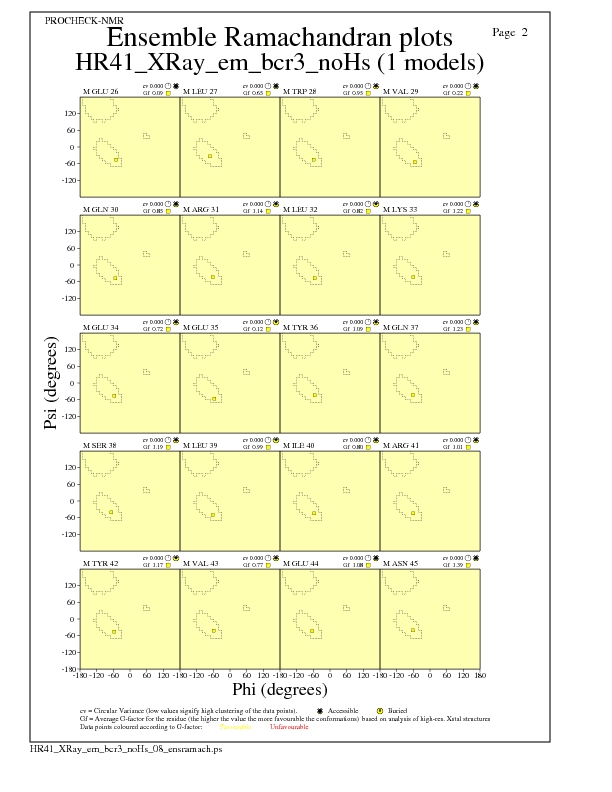
JPEG for residue Ramachandran Plots - page $num_n
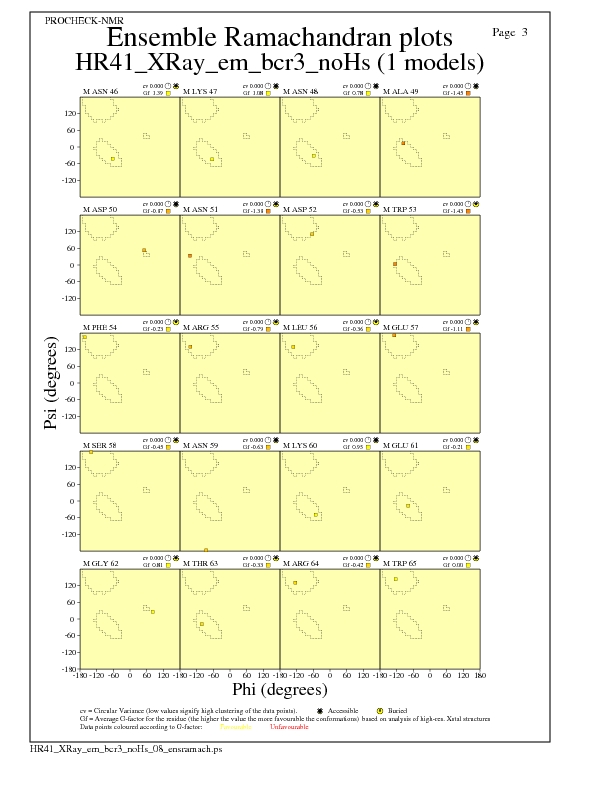
JPEG for residue Ramachandran Plots - page $num_n
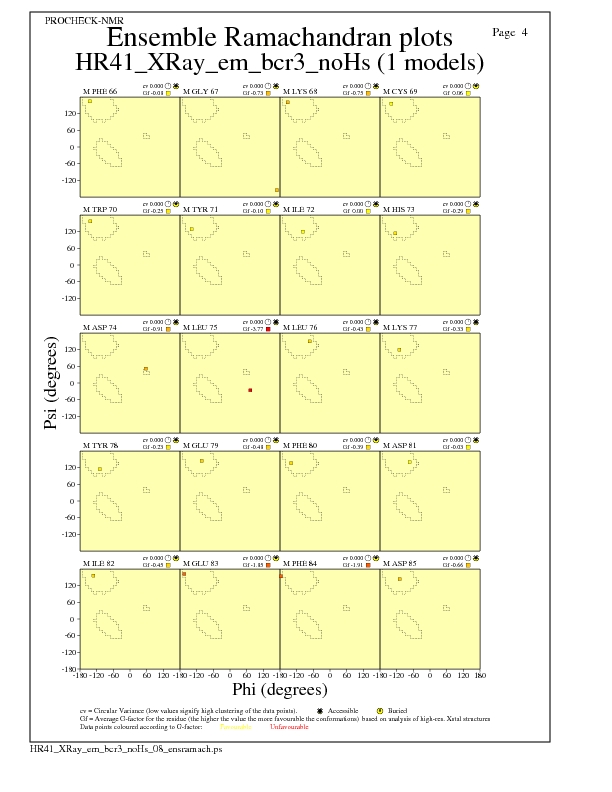
JPEG for residue Ramachandran Plots - page $num_n
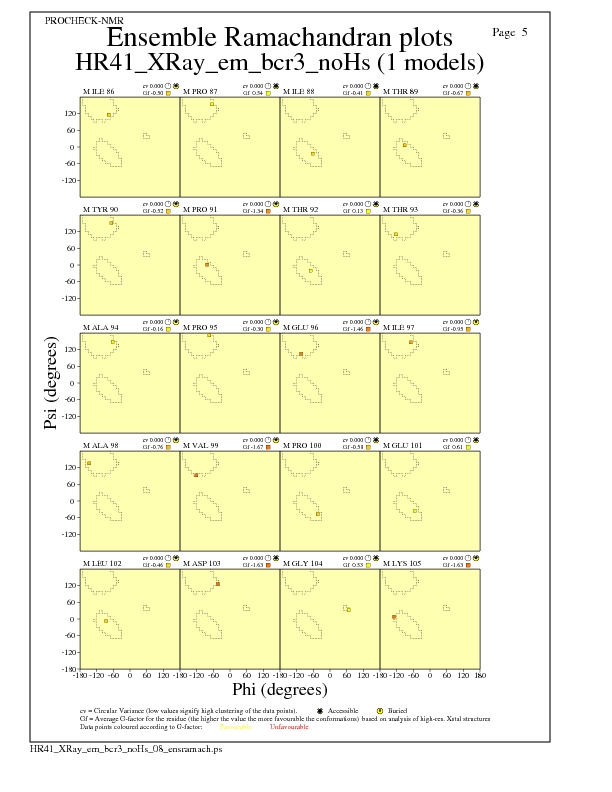
JPEG for residue Ramachandran Plots - page $num_n
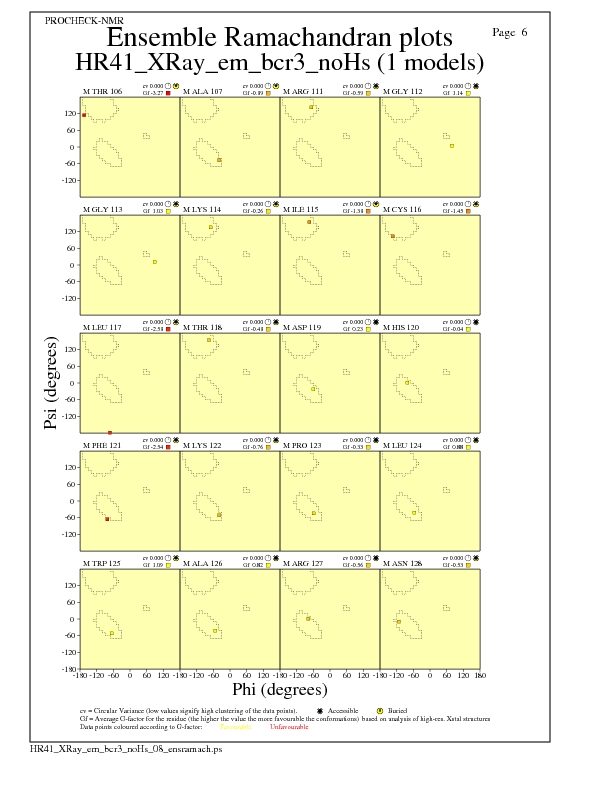
JPEG for residue Ramachandran Plots - page $num_n
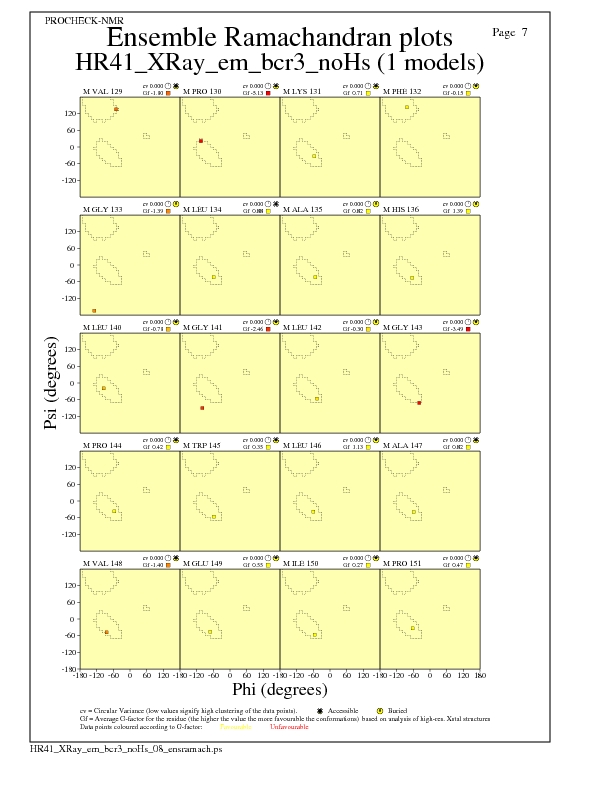
JPEG for residue Ramachandran Plots - page $num_n
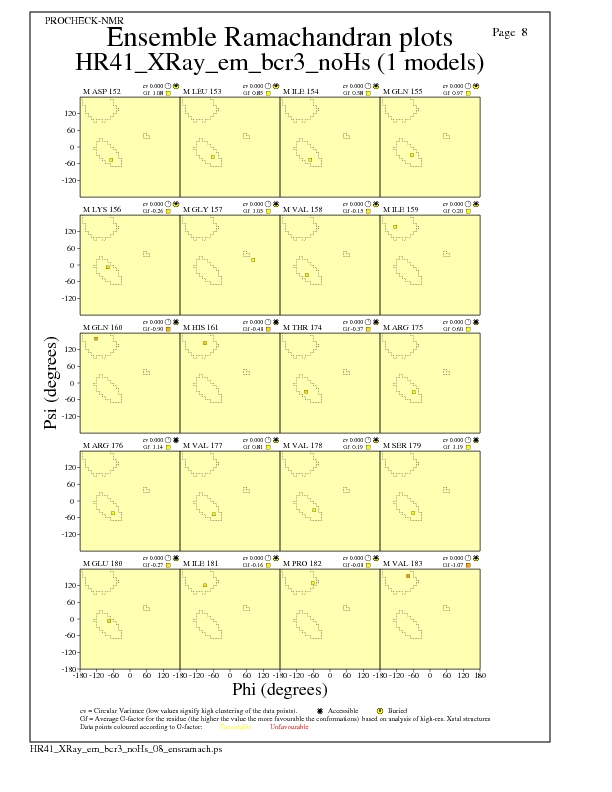
JPEG for residue Ramachandran Plots - page $num_n
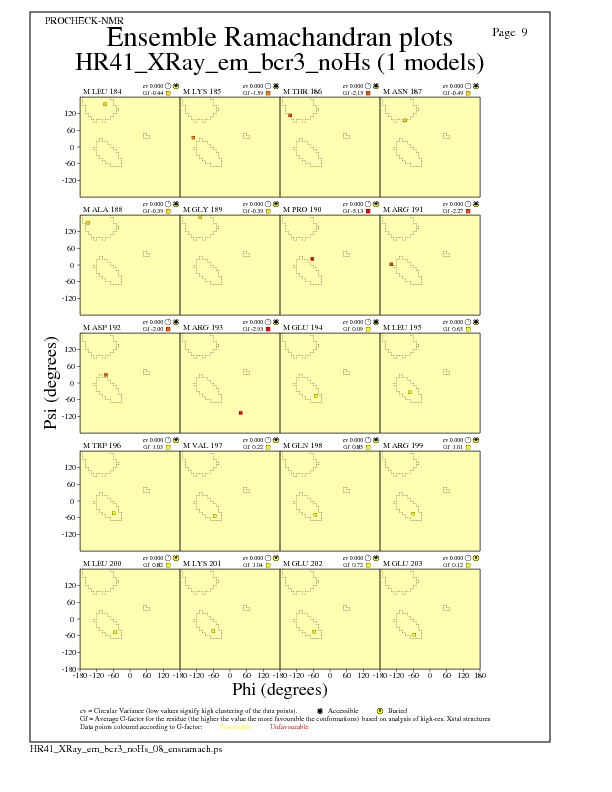
JPEG for residue Ramachandran Plots - page $num_n
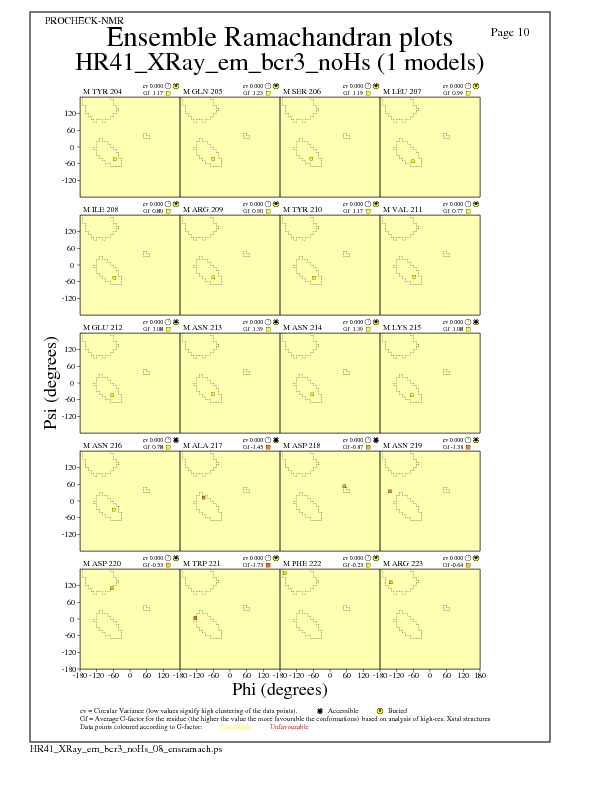
JPEG for residue Ramachandran Plots - page $num_n
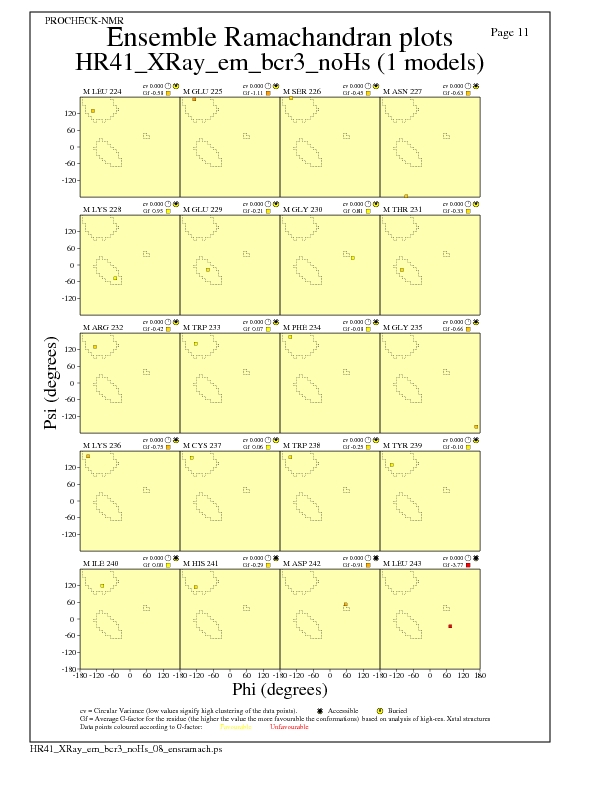

JPEG for residue Ramachandran Plots - page $num_n
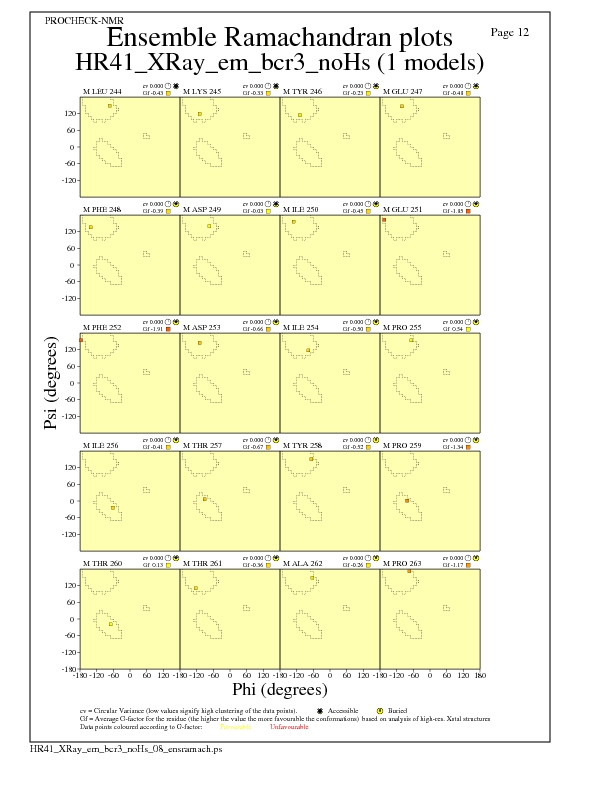
JPEG for residue Ramachandran Plots - page $num_n
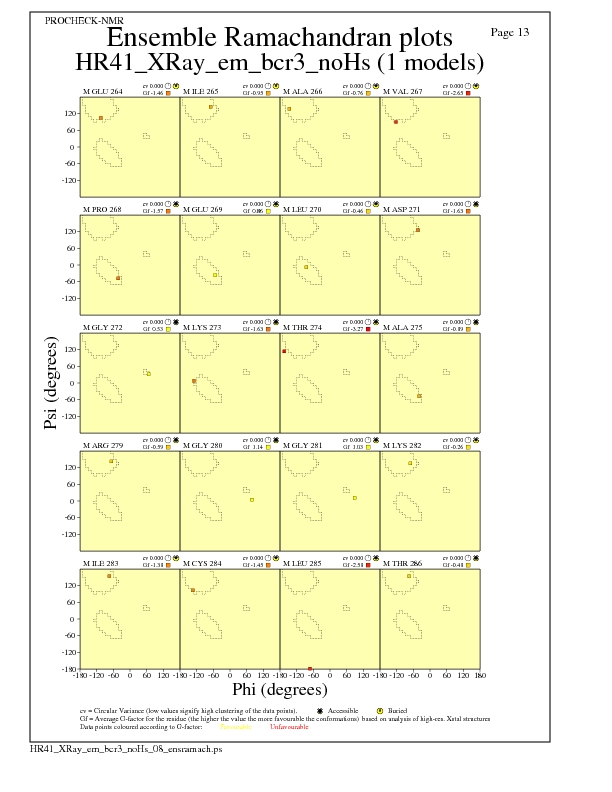
JPEG for residue Ramachandran Plots - page $num_n
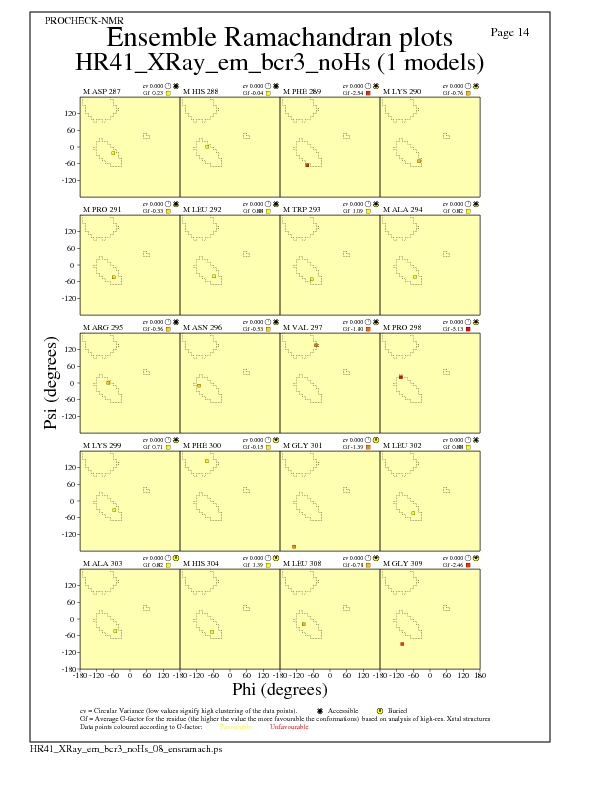
JPEG for residue Ramachandran Plots - page $num_n
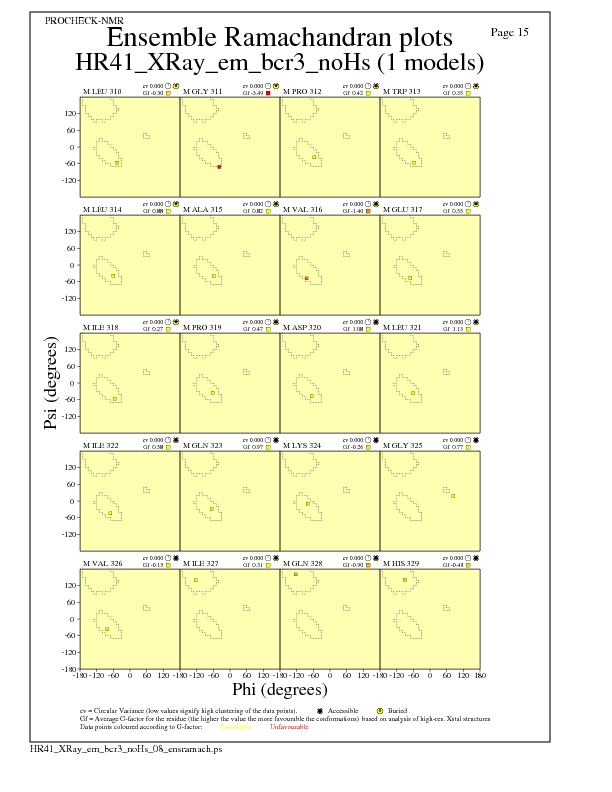
JPEG for residue Ramachandran Plots - page $num_n
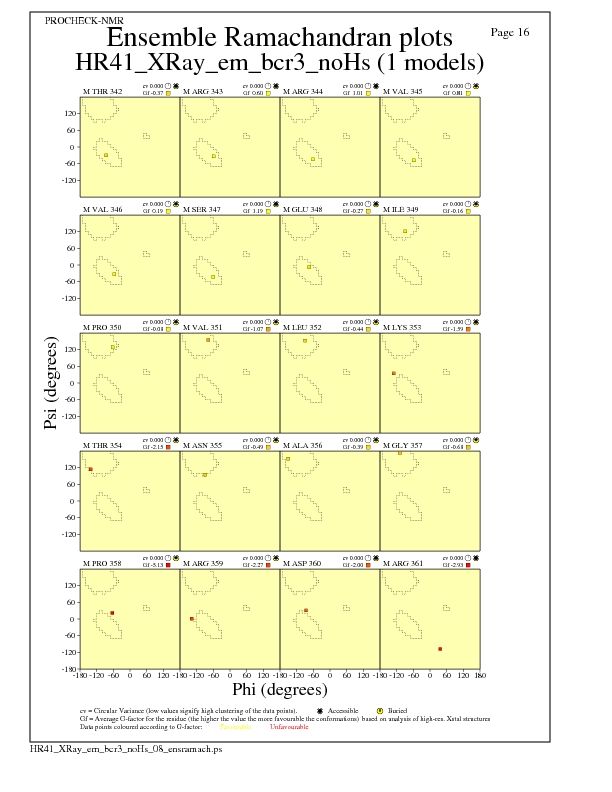
JPEG for residue Ramachandran Plots - page $num_n
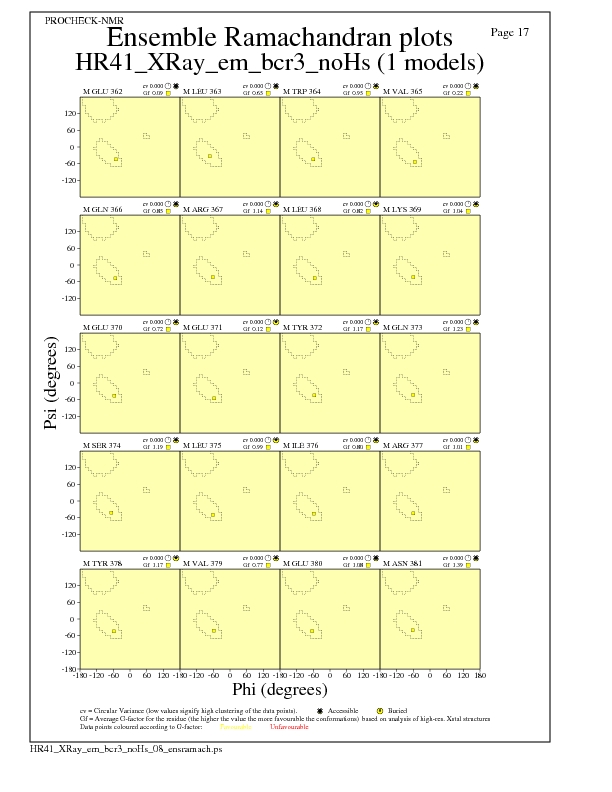
JPEG for residue Ramachandran Plots - page $num_n
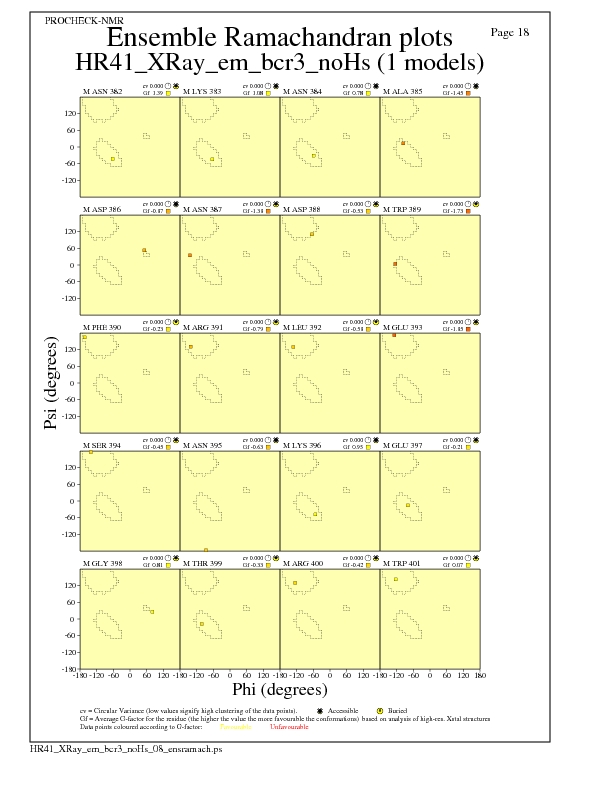
JPEG for residue Ramachandran Plots - page $num_n
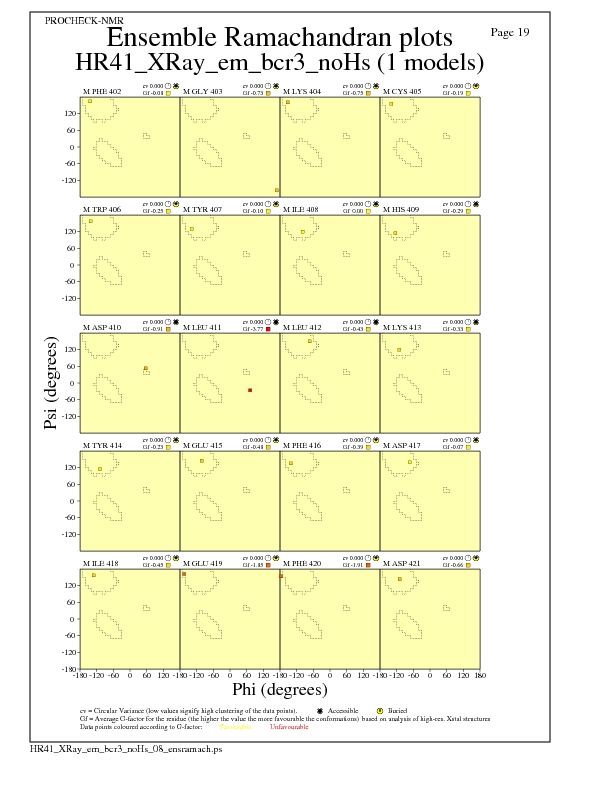
JPEG for residue Ramachandran Plots - page $num_n
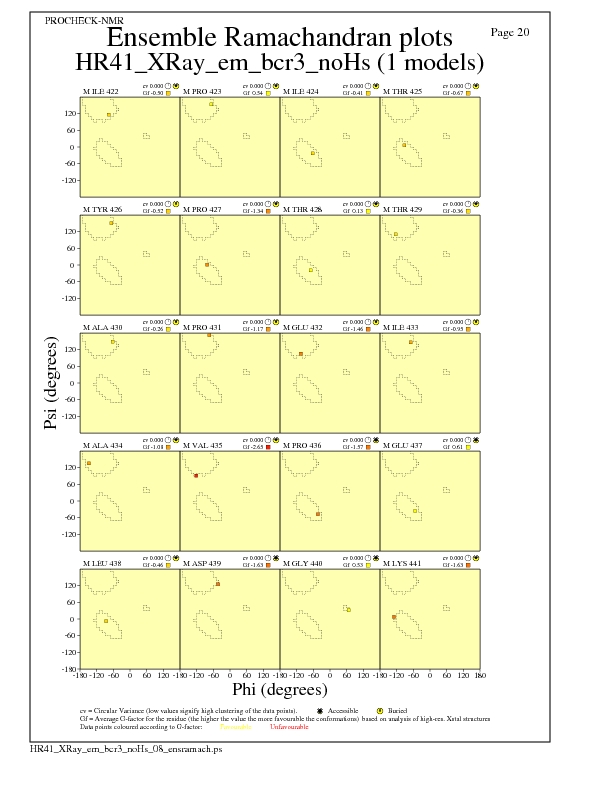
JPEG for residue Ramachandran Plots - page $num_n
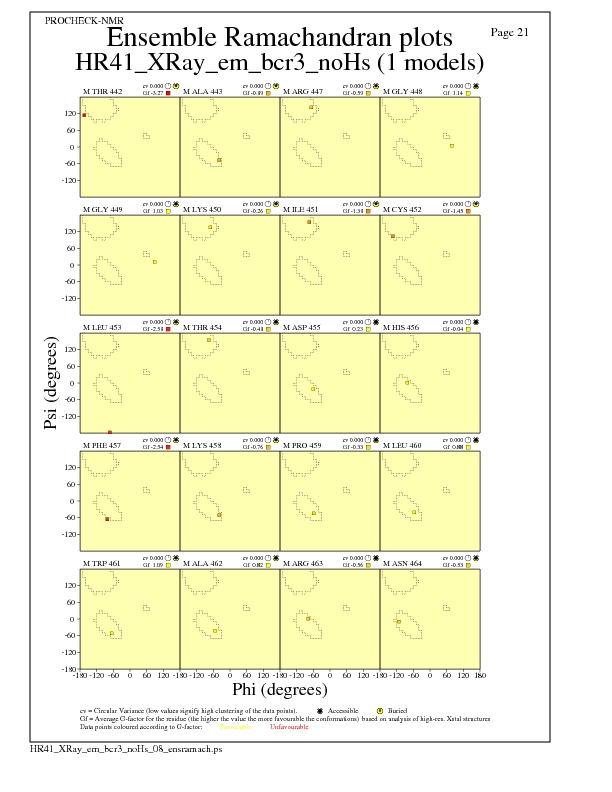
JPEG for residue Ramachandran Plots - page $num_n
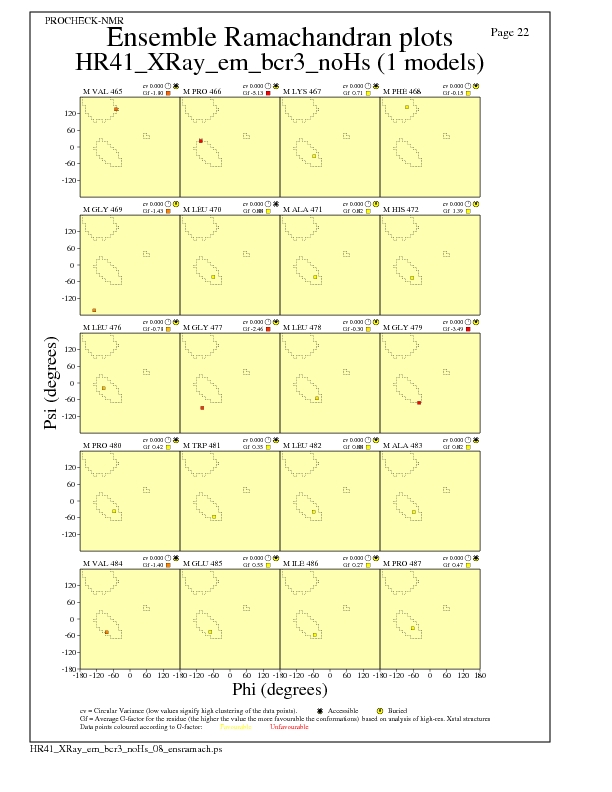
JPEG for residue Ramachandran Plots - page $num_n
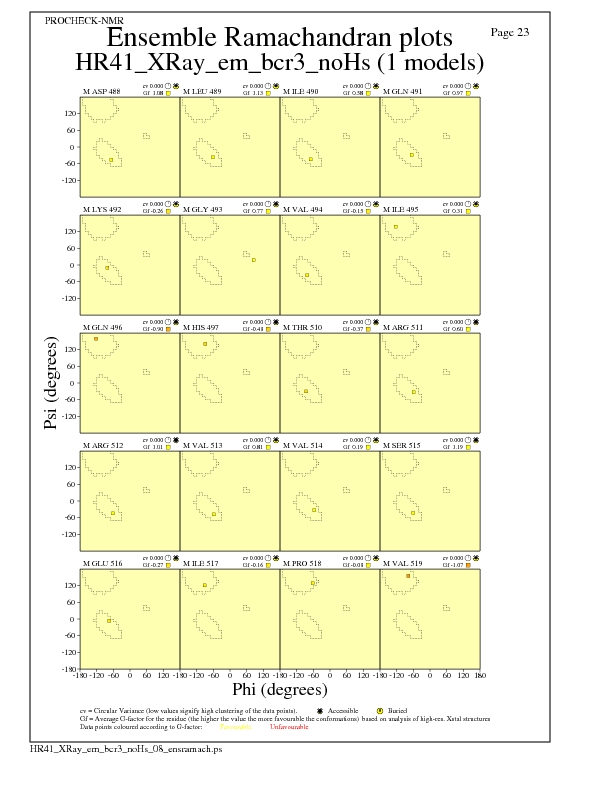
JPEG for residue Ramachandran Plots - page $num_n
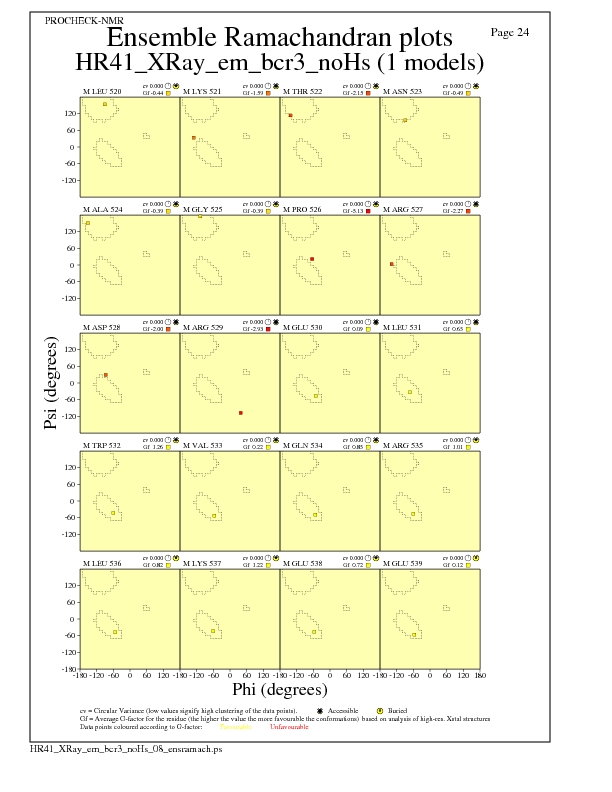
JPEG for residue Ramachandran Plots - page $num_n
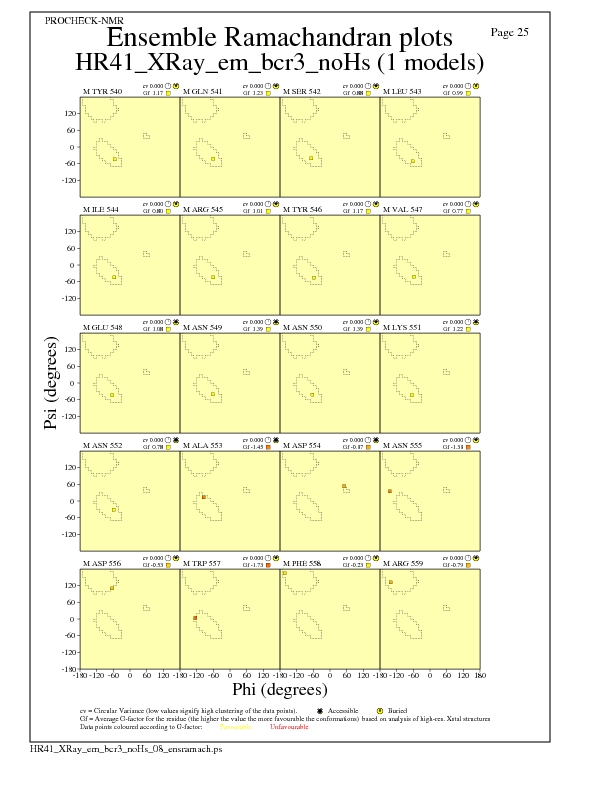
JPEG for residue Ramachandran Plots - page $num_n
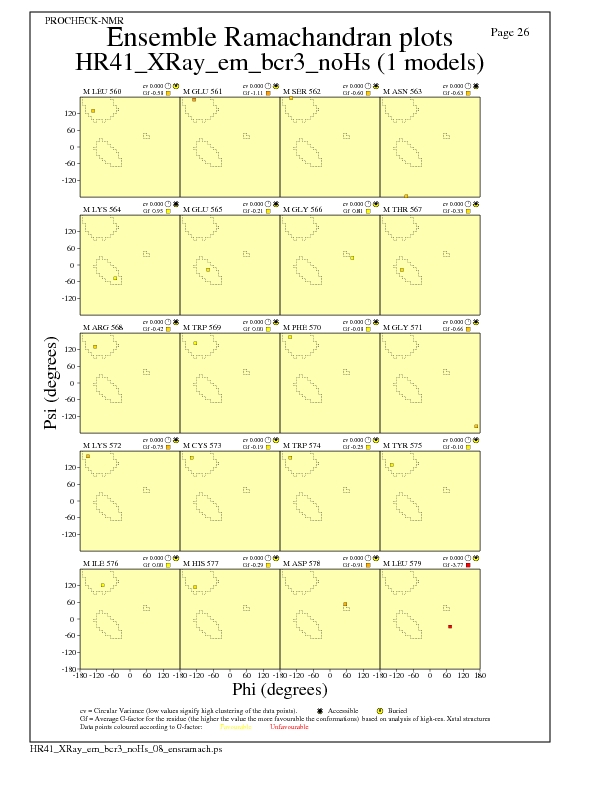

JPEG for residue Ramachandran Plots - page $num_n
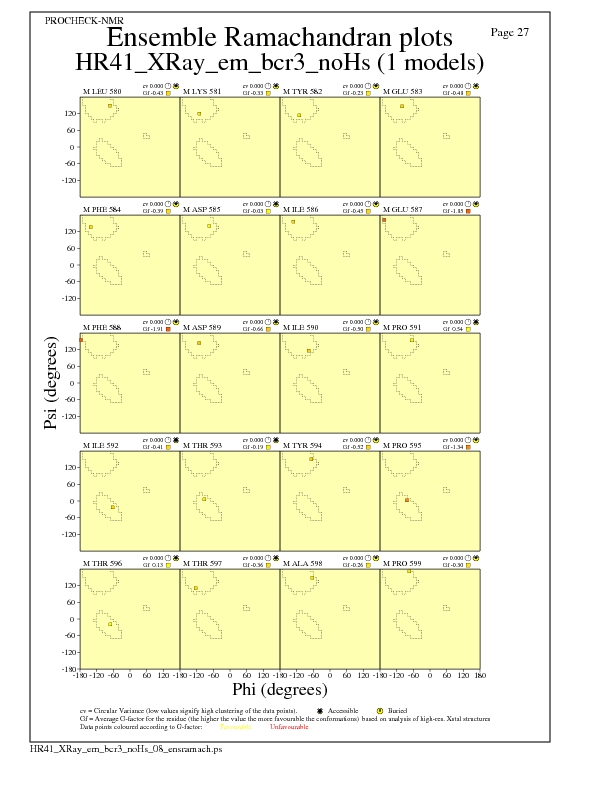
JPEG for residue Ramachandran Plots - page $num_n
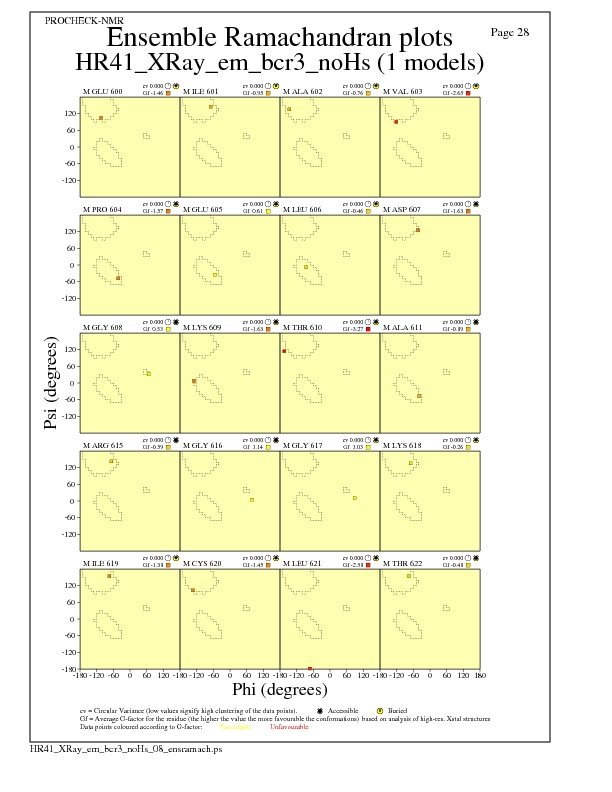
JPEG for residue Ramachandran Plots - page $num_n
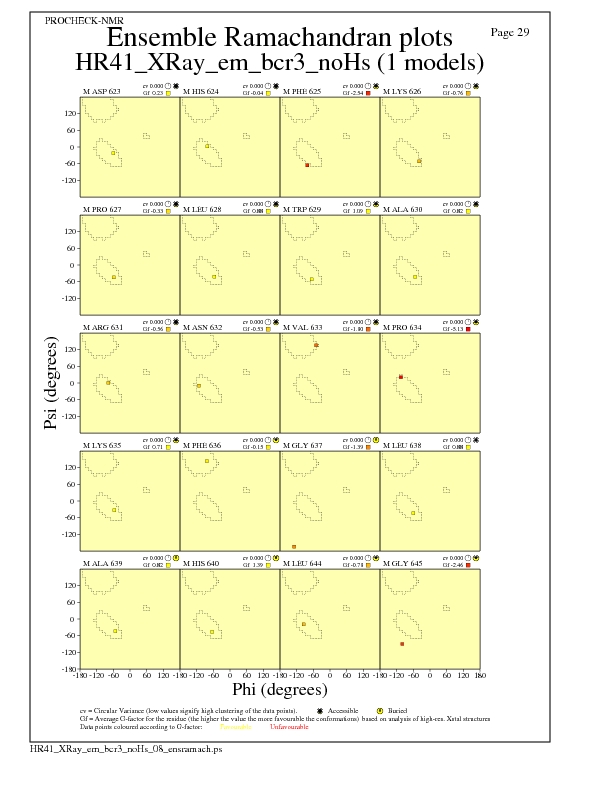
JPEG for residue Ramachandran Plots - page $num_n
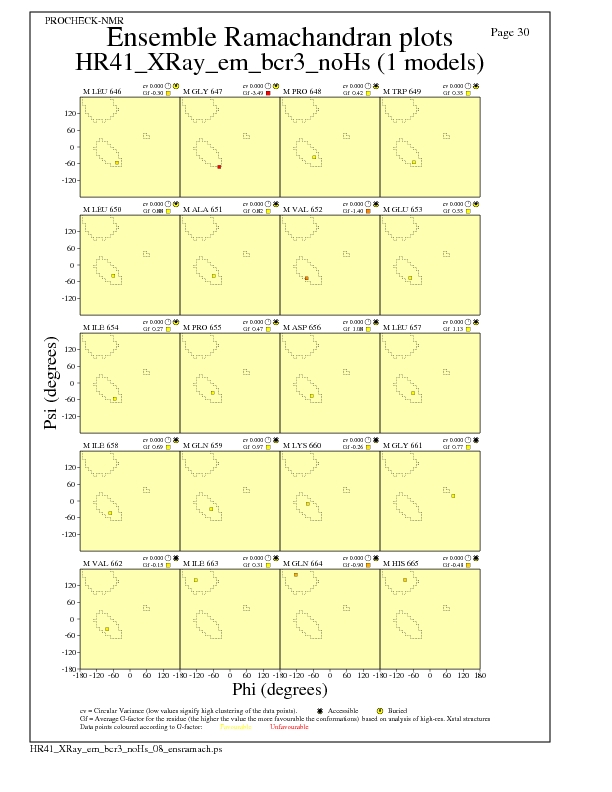
Ramachandran analysis for each residue from Molprobity
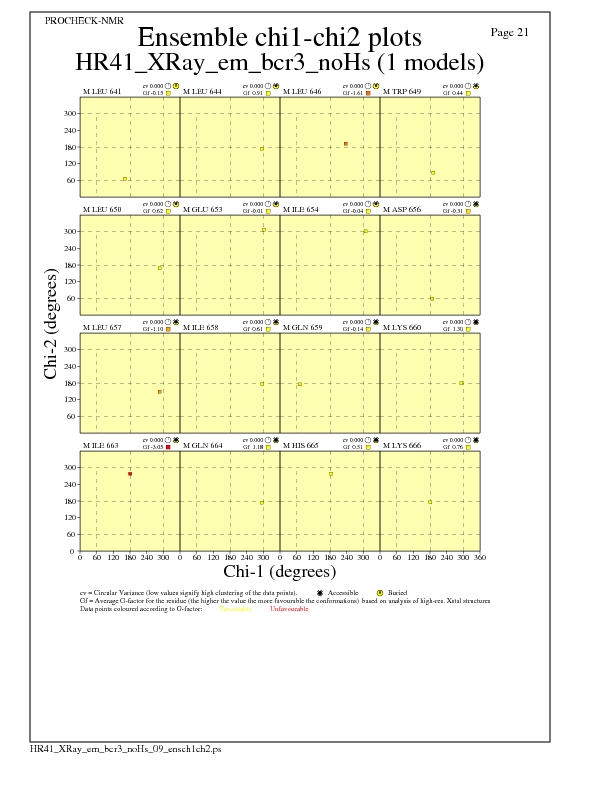
Chi1-Chi2 Plots for each residue
JPEG for residue Chi1-Chi2 Plots - page $num_n
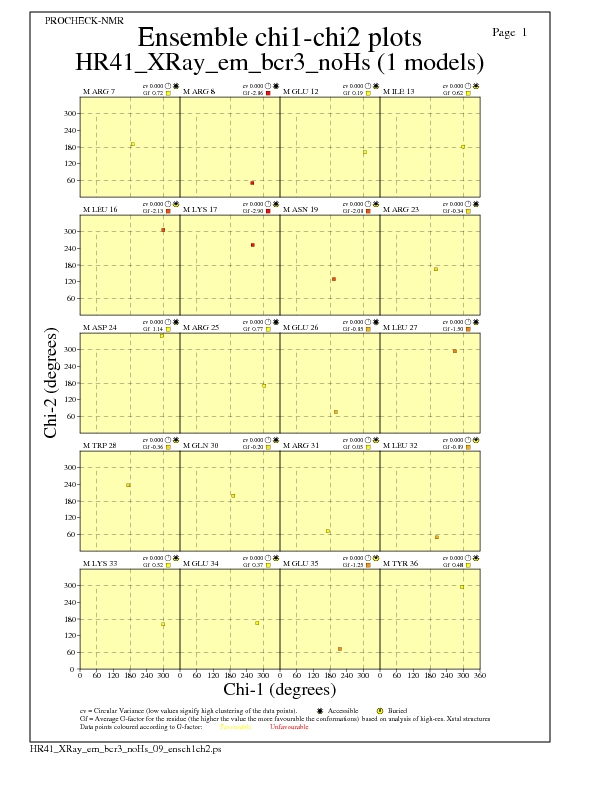
JPEG for residue Chi1-Chi2 Plots - page $num_n
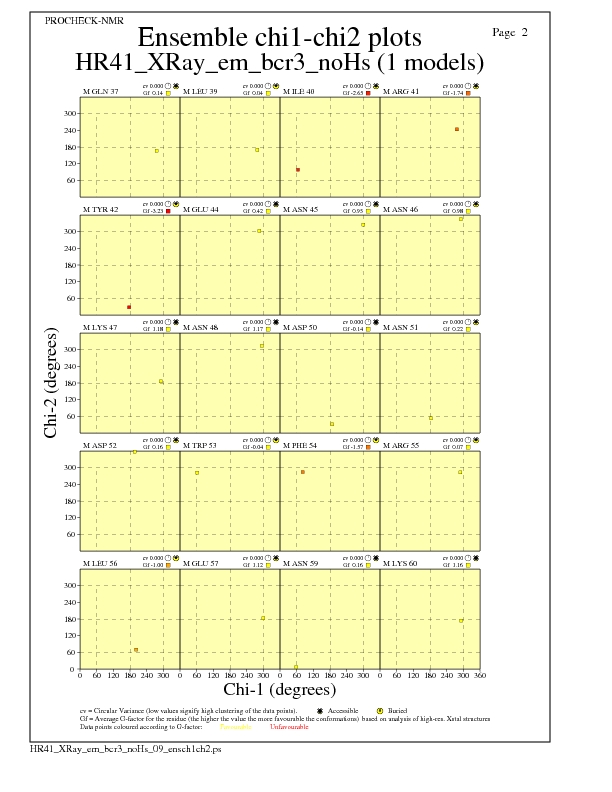
JPEG for residue Chi1-Chi2 Plots - page $num_n
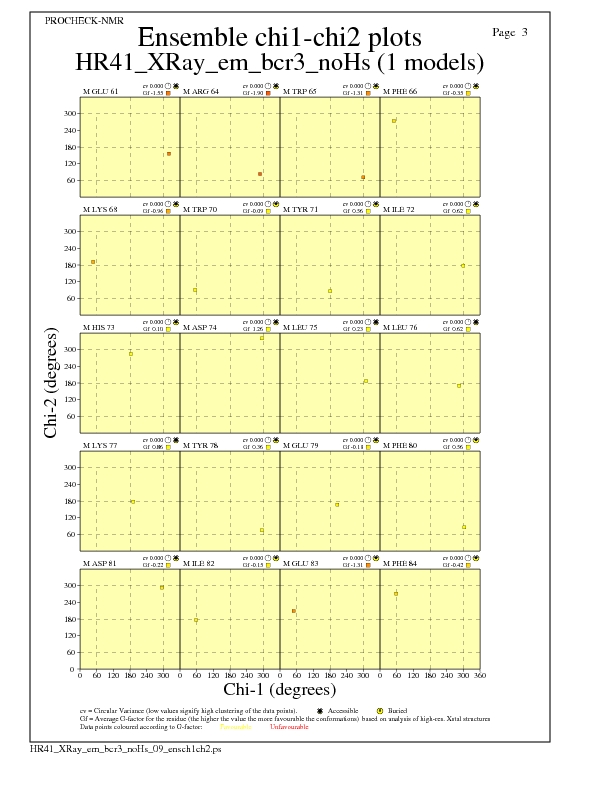
JPEG for residue Chi1-Chi2 Plots - page $num_n
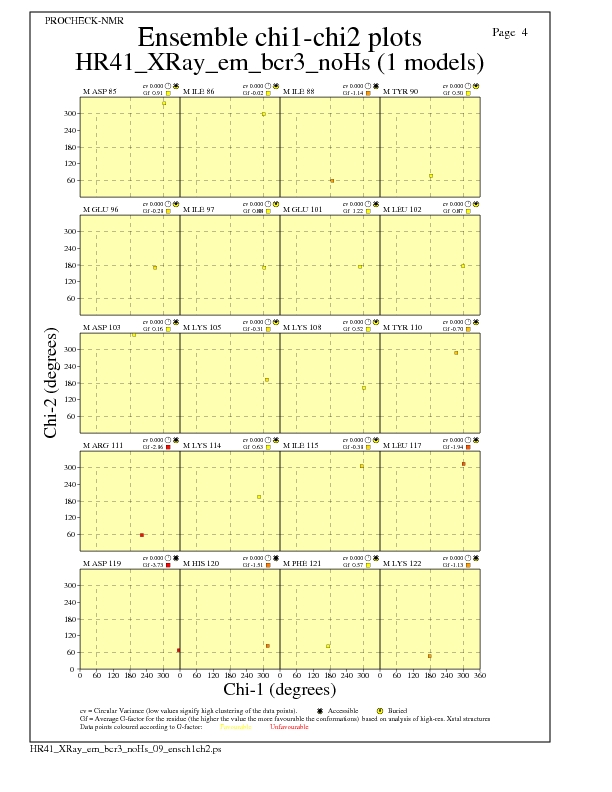
JPEG for residue Chi1-Chi2 Plots - page $num_n
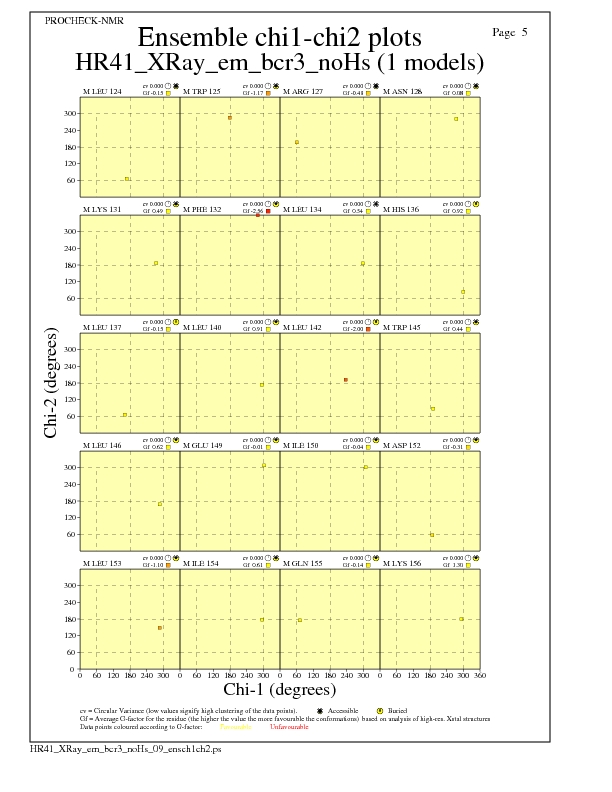
JPEG for residue Chi1-Chi2 Plots - page $num_n
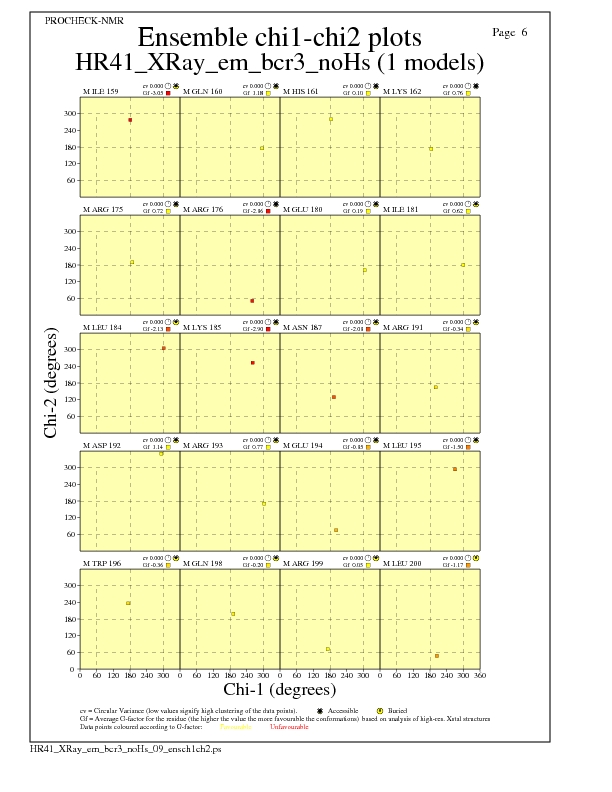
JPEG for residue Chi1-Chi2 Plots - page $num_n
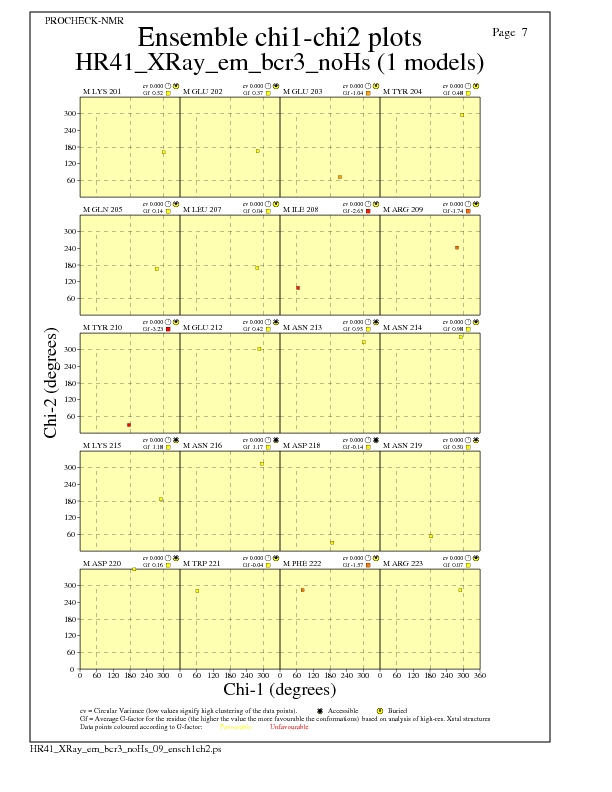
JPEG for residue Chi1-Chi2 Plots - page $num_n
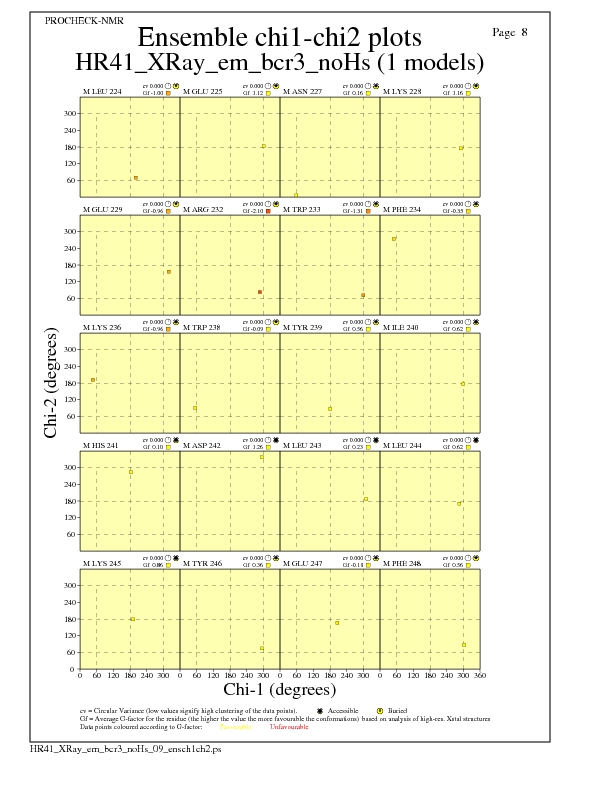
JPEG for residue Chi1-Chi2 Plots - page $num_n
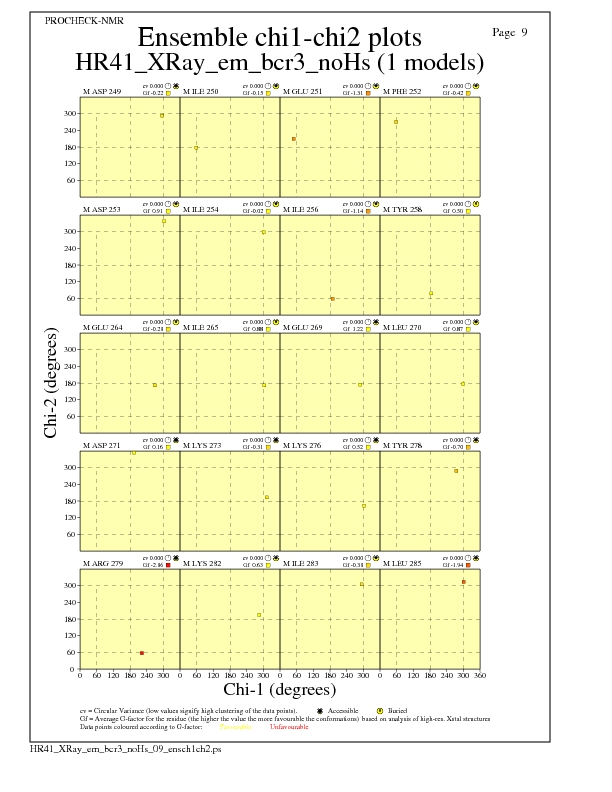
JPEG for residue Chi1-Chi2 Plots - page $num_n
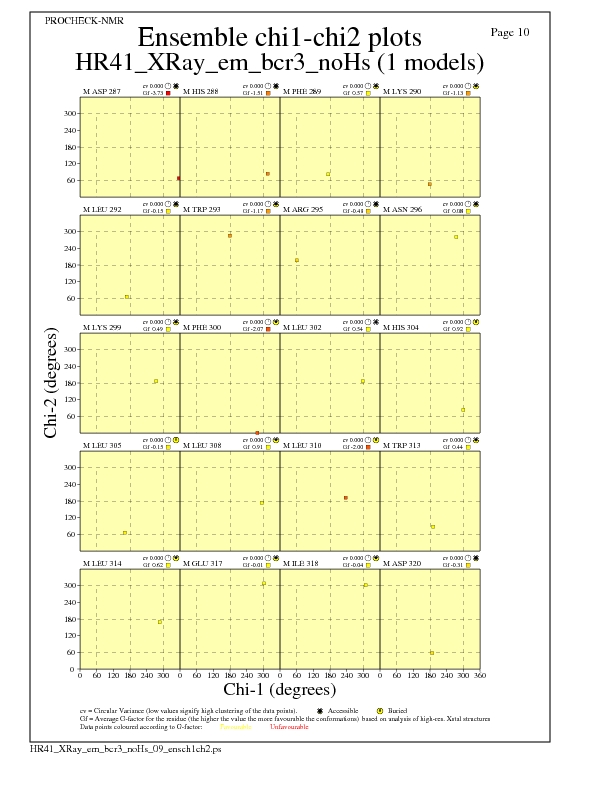
JPEG for residue Chi1-Chi2 Plots - page $num_n
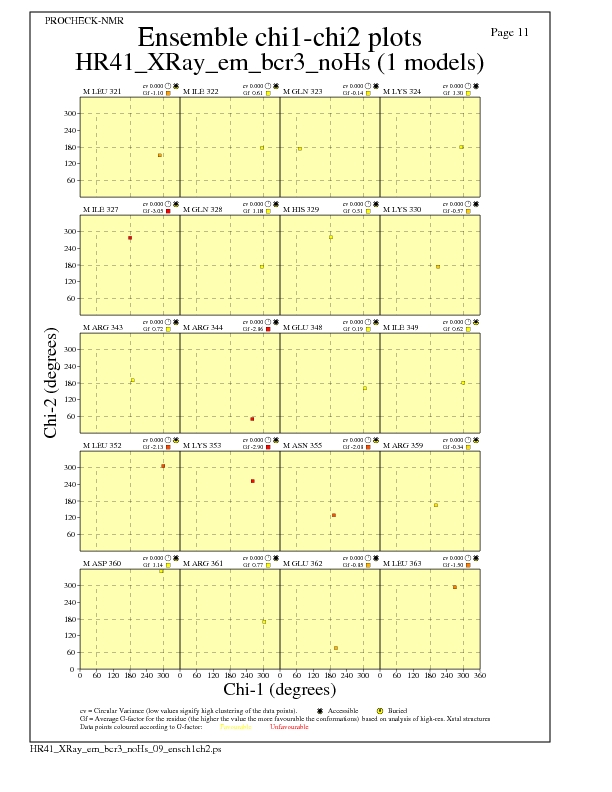
JPEG for residue Chi1-Chi2 Plots - page $num_n
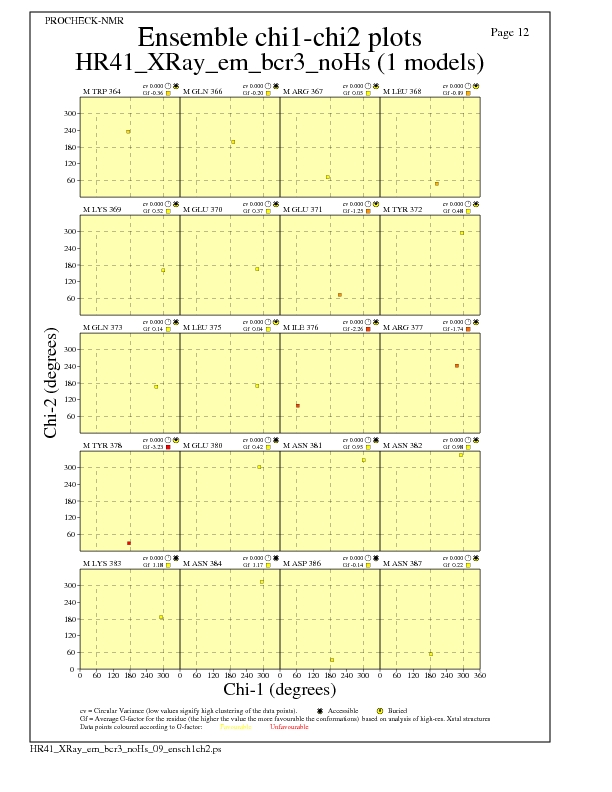
JPEG for residue Chi1-Chi2 Plots - page $num_n
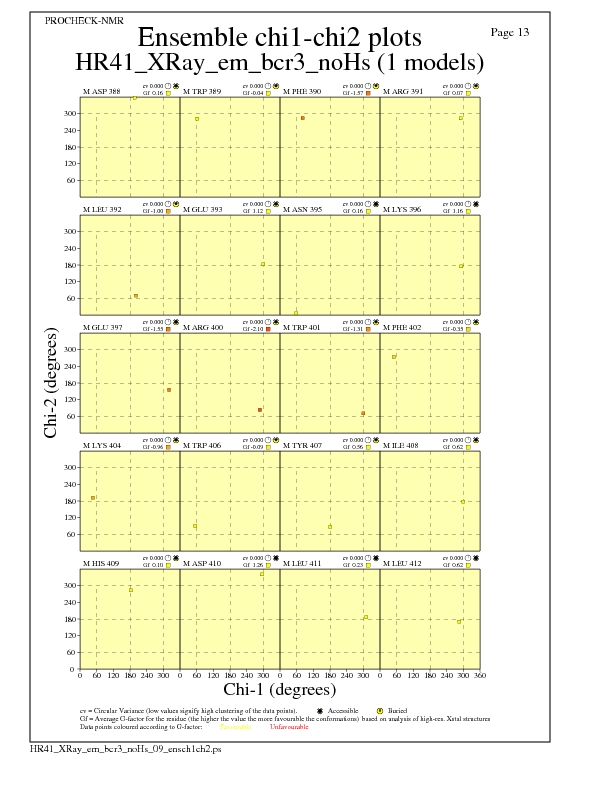
JPEG for residue Chi1-Chi2 Plots - page $num_n
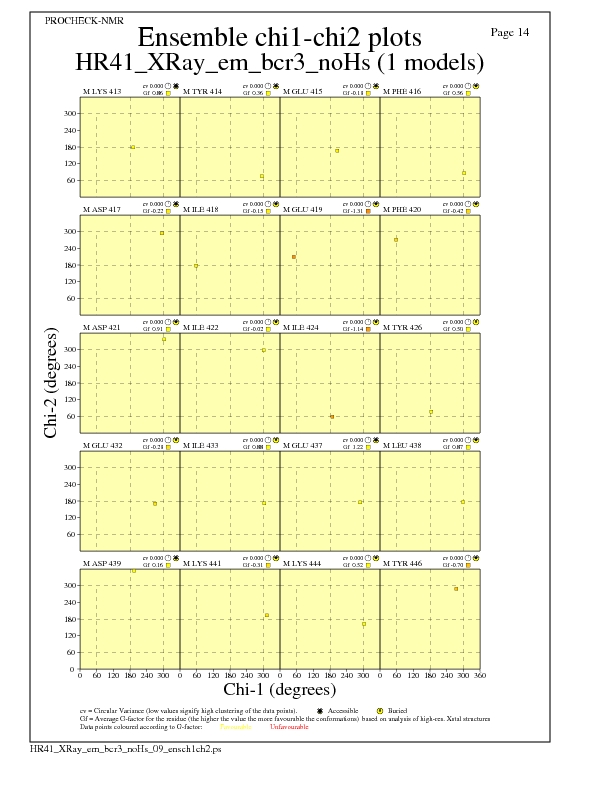
JPEG for residue Chi1-Chi2 Plots - page $num_n
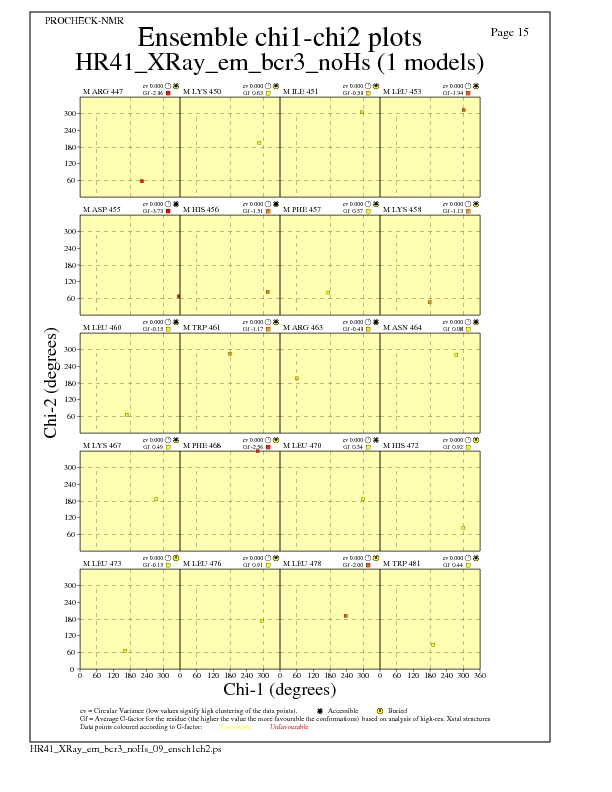
JPEG for residue Chi1-Chi2 Plots - page $num_n
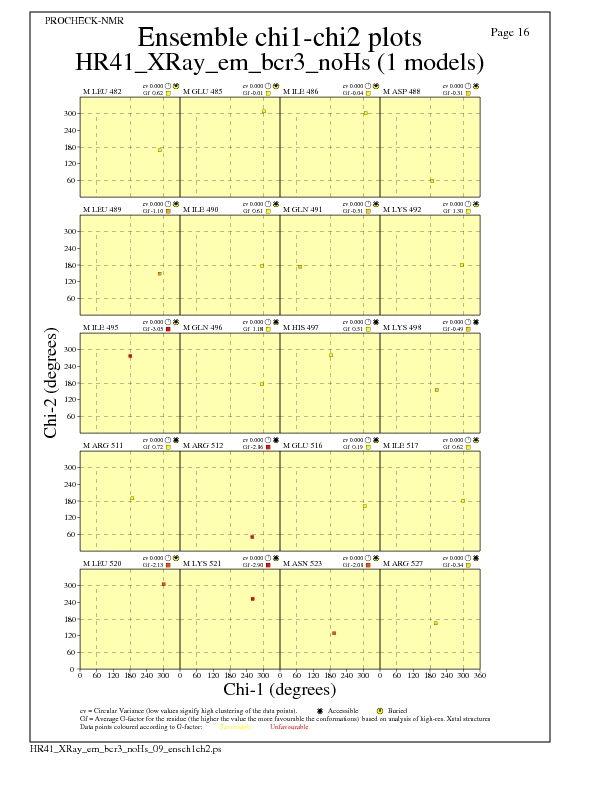
JPEG for residue Chi1-Chi2 Plots - page $num_n
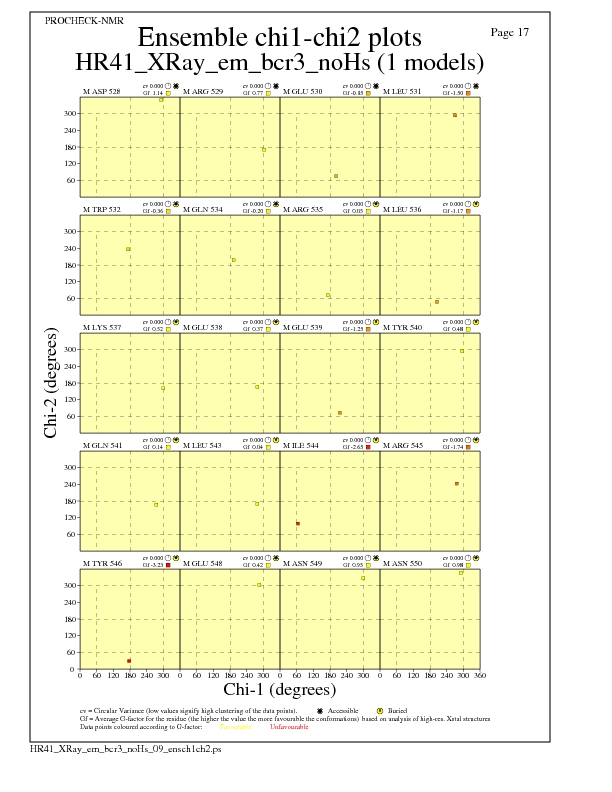
JPEG for residue Chi1-Chi2 Plots - page $num_n
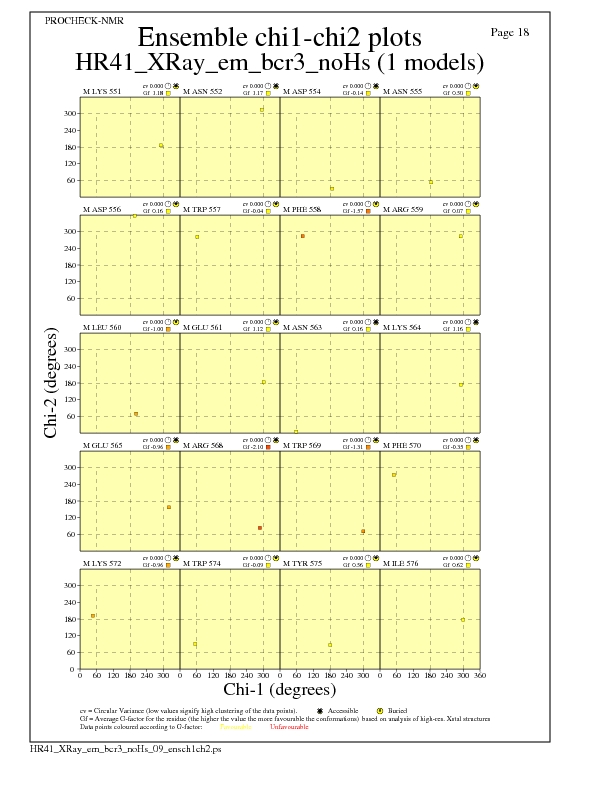
JPEG for residue Chi1-Chi2 Plots - page $num_n
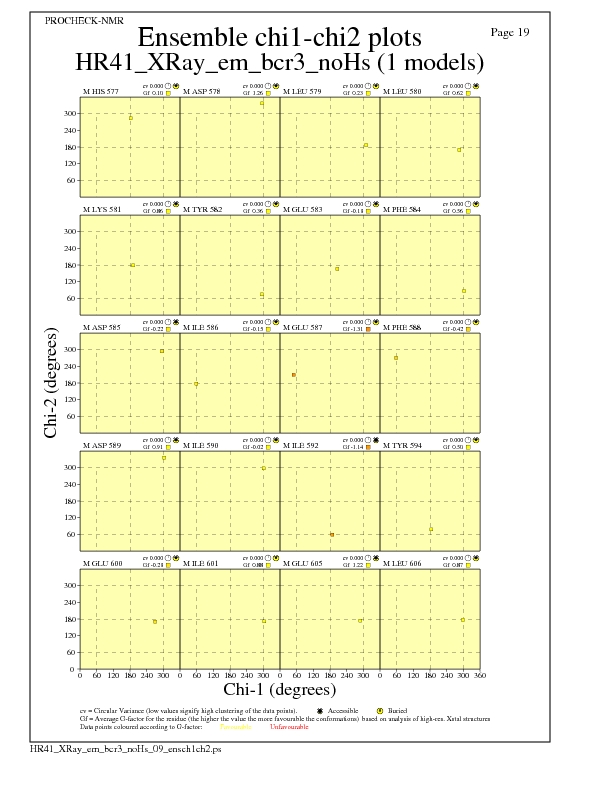
JPEG for residue Chi1-Chi2 Plots - page $num_n
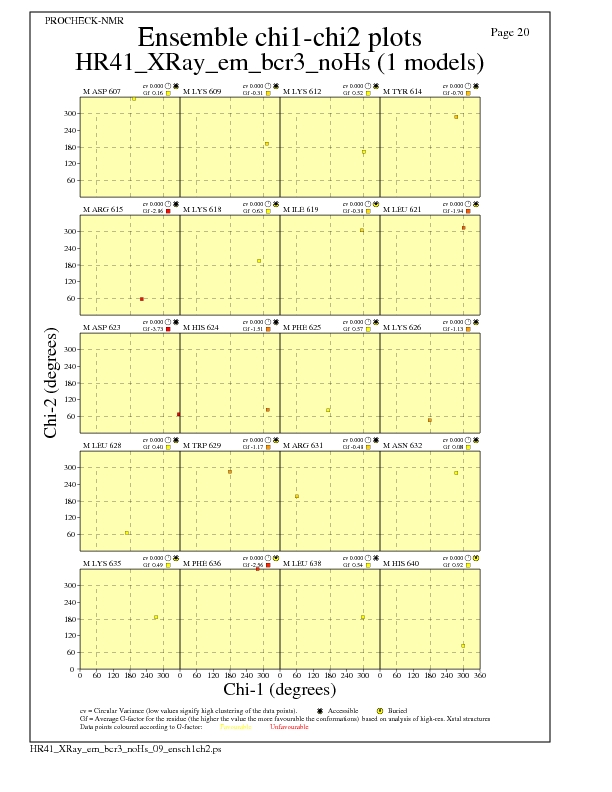
JPEG for residue Chi1-Chi2 Plots - page $num_n
Procheck G-factors for phi-psi for each residue
JPEG image for residue phi-psi G-factors
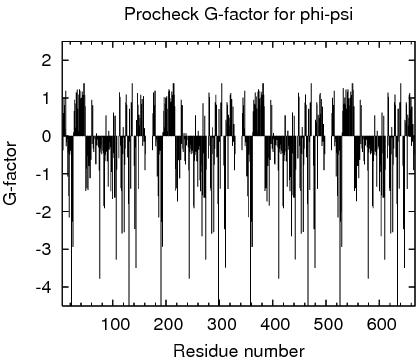
Table of Procheck G-factors for phi-psi for ordered residues
#phipsi_gfactor
#Residue\Model average
6 -0.37
7 0.60
8 1.01
9 0.81
10 0.19
11 1.19
12 -0.27
13 -0.16
14 -0.08
15 -1.07
16 -0.44
17 -1.59
18 -2.15
19 -0.49
20 -0.39
21 -0.39
22 -5.13
23 -2.27
24 -2.00
25 -2.93
26 0.09
27 0.65
28 0.95
29 0.22
30 0.85
31 1.14
32 0.82
33 1.22
34 0.72
35 0.12
36 1.09
37 1.23
38 1.19
39 0.99
40 0.80
41 1.01
42 1.17
43 0.77
44 1.08
45 1.39
46 1.39
47 1.08
48 0.78
49 -1.45
50 -0.87
51 -1.38
52 -0.53
53 -1.43
54 -0.23
55 -0.79
56 -0.36
57 -1.11
58 -0.45
59 -0.63
60 0.95
61 -0.21
62 0.81
63 -0.33
64 -0.42
65 0.00
66 -0.08
67 -0.73
68 -0.75
69 0.06
70 -0.25
71 -0.10
72 0.00
73 -0.29
74 -0.91
75 -3.77
76 -0.43
77 -0.33
78 -0.23
79 -0.48
80 -0.39
81 -0.03
82 -0.45
83 -1.85
84 -1.91
85 -0.66
86 -0.50
87 0.54
88 -0.41
89 -0.67
90 -0.52
91 -1.34
92 0.13
93 -0.36
94 -0.16
95 -0.30
96 -1.46
97 -0.95
98 -0.76
99 -1.67
100 -0.58
101 0.61
102 -0.46
103 -1.63
104 0.53
105 -1.63
106 -3.27
107 -0.89
111 -0.59
112 1.14
113 1.03
114 -0.26
115 -1.38
116 -1.45
117 -2.58
118 -0.48
119 0.23
120 -0.04
121 -2.54
122 -0.76
123 -0.33
124 0.88
125 1.09
126 0.82
127 -0.56
128 -0.53
129 -1.80
130 -5.13
131 0.71
132 -0.15
133 -1.39
134 0.88
135 0.82
136 1.39
140 -0.78
141 -2.46
142 -0.30
143 -3.49
144 0.42
145 0.35
146 1.13
147 0.82
148 -1.40
149 0.55
150 0.27
151 0.47
152 1.08
153 0.85
154 0.58
155 0.97
156 -0.26
157 1.05
158 -0.15
159 0.20
160 -0.90
161 -0.48
174 -0.37
175 0.60
176 1.14
177 0.81
178 0.19
179 1.19
180 -0.27
181 -0.16
182 -0.08
183 -1.07
184 -0.44
185 -1.59
186 -2.15
187 -0.49
188 -0.39
189 -0.39
190 -5.13
191 -2.27
192 -2.00
193 -2.93
194 0.09
195 0.65
196 1.03
197 0.22
198 0.85
199 1.01
200 0.82
201 1.04
202 0.72
203 0.12
204 1.17
205 1.23
206 1.19
207 0.99
208 0.80
209 0.90
210 1.17
211 0.77
212 1.08
213 1.39
214 1.39
215 1.08
216 0.78
217 -1.45
218 -0.87
219 -1.38
220 -0.53
221 -1.73
222 -0.23
223 -0.64
224 -0.58
225 -1.11
226 -0.45
227 -0.63
228 0.95
229 -0.21
230 0.81
231 -0.33
232 -0.42
233 0.07
234 -0.08
235 -0.66
236 -0.75
237 0.06
238 -0.25
239 -0.10
240 0.00
241 -0.29
242 -0.91
243 -3.77
244 -0.43
245 -0.33
246 -0.23
247 -0.48
248 -0.39
249 -0.03
250 -0.45
251 -1.85
252 -1.91
253 -0.66
254 -0.50
255 0.54
256 -0.41
257 -0.67
258 -0.52
259 -1.34
260 0.13
261 -0.36
262 -0.26
263 -1.17
264 -1.46
265 -0.95
266 -0.76
267 -2.65
268 -1.57
269 0.86
270 -0.46
271 -1.63
272 0.53
273 -1.63
274 -3.27
275 -0.89
279 -0.59
280 1.14
281 1.03
282 -0.26
283 -1.38
284 -1.45
285 -2.58
286 -0.48
287 0.23
288 -0.04
289 -2.54
290 -0.76
291 -0.33
292 0.88
293 1.09
294 0.82
295 -0.56
296 -0.53
297 -1.80
298 -5.13
299 0.71
300 -0.15
301 -1.39
302 0.88
303 0.82
304 1.39
308 -0.78
309 -2.46
310 -0.30
311 -3.49
312 0.42
313 0.35
314 0.88
315 0.82
316 -1.40
317 0.55
318 0.27
319 0.47
320 1.08
321 1.13
322 0.58
323 0.97
324 -0.26
325 0.77
326 -0.15
327 0.31
328 -0.90
329 -0.48
342 -0.37
343 0.60
344 1.01
345 0.81
346 0.19
347 1.19
348 -0.27
349 -0.16
350 -0.08
351 -1.07
352 -0.44
353 -1.59
354 -2.15
355 -0.49
356 -0.39
357 -0.68
358 -5.13
359 -2.27
360 -2.00
361 -2.93
362 0.09
363 0.65
364 0.95
365 0.22
366 0.85
367 1.14
368 0.82
369 1.04
370 0.72
371 0.12
372 1.17
373 1.23
374 1.19
375 0.99
376 0.80
377 1.01
378 1.17
379 0.77
380 1.08
381 1.39
382 1.39
383 1.08
384 0.78
385 -1.45
386 -0.87
387 -1.38
388 -0.53
389 -1.73
390 -0.23
391 -0.79
392 -0.58
393 -1.85
394 -0.45
395 -0.63
396 0.95
397 -0.21
398 0.81
399 -0.33
400 -0.42
401 0.07
402 -0.08
403 -0.73
404 -0.75
405 -0.19
406 -0.25
407 -0.10
408 0.00
409 -0.29
410 -0.91
411 -3.77
412 -0.43
413 -0.33
414 -0.23
415 -0.48
416 -0.39
417 -0.07
418 -0.45
419 -1.85
420 -1.91
421 -0.66
422 -0.50
423 0.54
424 -0.41
425 -0.67
426 -0.52
427 -1.34
428 0.13
429 -0.36
430 -0.26
431 -1.17
432 -1.46
433 -0.95
434 -1.08
435 -2.65
436 -1.57
437 0.61
438 -0.46
439 -1.63
440 0.53
441 -1.63
442 -3.27
443 -0.89
447 -0.59
448 1.14
449 1.03
450 -0.26
451 -1.38
452 -1.45
453 -2.58
454 -0.48
455 0.23
456 -0.04
457 -2.54
458 -0.76
459 -0.33
460 0.88
461 1.09
462 0.82
463 -0.56
464 -0.53
465 -1.80
466 -5.13
467 0.71
468 -0.15
469 -1.43
470 0.88
471 0.82
472 1.39
476 -0.78
477 -2.46
478 -0.30
479 -3.49
480 0.42
481 0.35
482 0.88
483 0.82
484 -1.40
485 0.55
486 0.27
487 0.47
488 1.08
489 1.13
490 0.58
491 0.97
492 -0.26
493 0.77
494 -0.15
495 0.31
496 -0.90
497 -0.48
510 -0.37
511 0.60
512 1.01
513 0.81
514 0.19
515 1.19
516 -0.27
517 -0.16
518 -0.08
519 -1.07
520 -0.44
521 -1.59
522 -2.15
523 -0.49
524 -0.39
525 -0.39
526 -5.13
527 -2.27
528 -2.00
529 -2.93
530 0.09
531 0.65
532 1.26
533 0.22
534 0.85
535 1.01
536 0.82
537 1.22
538 0.72
539 0.12
540 1.17
541 1.23
542 0.88
543 0.99
544 0.80
545 1.01
546 1.17
547 0.77
548 1.08
549 1.39
550 1.39
551 1.22
552 0.78
553 -1.45
554 -0.87
555 -1.38
556 -0.53
557 -1.73
558 -0.23
559 -0.79
560 -0.58
561 -1.11
562 -0.60
563 -0.63
564 0.95
565 -0.21
566 0.81
567 -0.33
568 -0.42
569 0.00
570 -0.08
571 -0.66
572 -0.75
573 -0.19
574 -0.25
575 -0.10
576 0.00
577 -0.29
578 -0.91
579 -3.77
580 -0.43
581 -0.33
582 -0.23
583 -0.48
584 -0.39
585 -0.03
586 -0.45
587 -1.85
588 -1.91
589 -0.66
590 -0.50
591 0.54
592 -0.41
593 -0.19
594 -0.52
595 -1.34
596 0.13
597 -0.36
598 -0.26
599 -0.30
600 -1.46
601 -0.95
602 -0.76
603 -2.65
604 -1.57
605 0.61
606 -0.46
607 -1.63
608 0.53
609 -1.63
610 -3.27
611 -0.89
615 -0.59
616 1.14
617 1.03
618 -0.26
619 -1.38
620 -1.45
621 -2.58
622 -0.48
623 0.23
624 -0.04
625 -2.54
626 -0.76
627 -0.33
628 0.88
629 1.09
630 0.82
631 -0.56
632 -0.53
633 -1.80
634 -5.13
635 0.71
636 -0.15
637 -1.39
638 0.88
639 0.82
640 1.39
644 -0.78
645 -2.46
646 -0.30
647 -3.49
648 0.42
649 0.35
650 0.88
651 0.82
652 -1.40
653 0.55
654 0.27
655 0.47
656 1.08
657 1.13
658 0.69
659 0.97
660 -0.26
661 0.77
662 -0.15
663 0.31
664 -0.90
665 -0.48
#Reported_Model_Average -0.321
#Overall_Average_Reported -0.321
Procheck G-factors for all dihedral angles for each residue
JPEG image for residue all dihedral G-factors
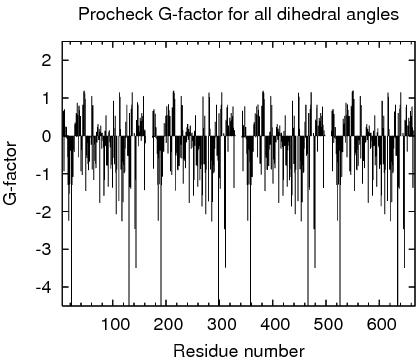
Table of Procheck G-factors for all dihedrals for ordered residues
#alldih_gfactor
#Residue\Model average
5 0.00
6 0.13
7 0.66
8 -0.93
9 0.70
10 -0.03
11 0.24
12 -0.04
13 0.23
14 -0.08
15 -0.55
16 -1.29
17 -2.24
18 -1.52
19 -1.28
20 -0.39
21 -0.39
22 -5.13
23 -1.30
24 -0.43
25 -1.08
26 -0.38
27 -0.42
28 0.30
29 0.36
30 0.32
31 0.59
32 -0.03
33 0.87
34 0.55
35 -0.56
36 0.79
37 0.68
38 0.24
39 0.52
40 -0.93
41 -0.37
42 -1.03
43 0.82
44 0.75
45 1.17
46 1.19
47 1.13
48 0.97
49 -1.45
50 -0.51
51 -0.58
52 -0.19
53 -0.74
54 -0.90
55 -0.36
56 -0.68
57 0.00
58 -0.04
59 -0.23
60 1.05
61 -0.88
62 0.81
63 -0.02
64 -1.16
65 -0.66
66 -0.22
67 -0.73
68 -0.85
69 0.10
70 -0.17
71 0.23
72 0.31
73 -0.10
74 0.18
75 -1.77
76 0.10
77 0.27
78 0.07
79 -0.33
80 0.08
81 -0.13
82 -0.30
83 -1.58
84 -1.16
85 0.13
86 -0.26
87 0.54
88 -0.77
89 -0.78
90 -0.01
91 -1.34
92 0.09
93 0.14
94 -0.16
95 -0.30
96 -0.87
97 -0.04
98 -0.76
99 -1.37
100 -0.58
101 0.92
102 0.21
103 -0.74
104 0.53
105 -0.97
106 -2.07
107 -0.89
108 0.52
110 -0.70
111 -1.72
112 1.14
113 1.03
114 0.18
115 -0.88
116 -0.66
117 -2.26
118 -0.68
119 -1.75
120 -0.77
121 -0.98
122 -0.94
123 -0.33
124 0.37
125 -0.04
126 0.82
127 -0.52
128 -0.22
129 -0.53
130 -5.13
131 0.60
132 -1.35
133 -1.39
134 0.71
135 0.82
136 1.15
137 -0.15
139 0.00
140 0.07
141 -2.46
142 -1.15
143 -3.49
144 0.42
145 0.39
146 0.88
147 0.82
148 -0.27
149 0.27
150 0.11
151 0.47
152 0.39
153 -0.13
154 0.60
155 0.42
156 0.52
157 1.05
158 0.17
159 -1.43
160 0.14
161 -0.19
162 0.76
173 0.00
174 -0.04
175 0.66
176 -0.86
177 0.70
178 -0.03
179 0.58
180 -0.04
181 0.23
182 -0.08
183 -0.47
184 -1.29
185 -2.24
186 -0.59
187 -1.28
188 -0.39
189 -0.39
190 -5.13
191 -1.30
192 -0.43
193 -1.08
194 -0.38
195 -0.42
196 0.34
197 -0.01
198 0.32
199 0.53
200 -0.18
201 0.78
202 0.55
203 -0.46
204 0.83
205 0.68
206 0.24
207 0.52
208 -0.93
209 -0.42
210 -1.03
211 0.75
212 0.75
213 1.17
214 1.19
215 1.13
216 0.97
217 -1.45
218 -0.51
219 -0.44
220 -0.19
221 -0.89
222 -0.90
223 -0.28
224 -0.79
225 0.00
226 -0.58
227 -0.23
228 1.05
229 -0.59
230 0.81
231 0.15
232 -1.26
233 -0.62
234 -0.22
235 -0.66
236 -0.85
237 0.10
238 -0.17
239 0.23
240 0.31
241 -0.10
242 0.18
243 -1.77
244 0.10
245 0.27
246 0.07
247 -0.33
248 0.08
249 -0.13
250 -0.30
251 -1.58
252 -1.16
253 0.13
254 -0.26
255 0.54
256 -0.77
257 -0.78
258 -0.01
259 -1.34
260 -0.37
261 -0.33
262 -0.26
263 -1.17
264 -0.87
265 -0.04
266 -0.76
267 -1.86
268 -1.57
269 1.04
270 0.21
271 -0.74
272 0.53
273 -0.97
274 -2.07
275 -0.89
276 0.52
278 -0.70
279 -1.72
280 1.14
281 1.03
282 0.18
283 -0.88
284 -0.66
285 -2.26
286 -0.68
287 -1.75
288 -0.77
289 -0.98
290 -0.94
291 -0.33
292 0.37
293 -0.04
294 0.82
295 -0.52
296 -0.22
297 -0.65
298 -5.13
299 0.60
300 -1.11
301 -1.39
302 0.71
303 0.82
304 1.15
305 -0.15
307 0.00
308 0.07
309 -2.46
310 -1.15
311 -3.49
312 0.42
313 0.39
314 0.75
315 0.82
316 -0.41
317 0.27
318 0.11
319 0.47
320 0.39
321 0.02
322 0.60
323 0.42
324 0.52
325 0.77
326 0.22
327 -1.37
328 0.14
329 0.02
330 -0.57
341 0.00
342 -0.04
343 0.66
344 -0.93
345 0.28
346 -0.03
347 0.78
348 -0.04
349 0.23
350 -0.08
351 -0.47
352 -1.29
353 -2.24
354 -0.35
355 -1.28
356 -0.39
357 -0.68
358 -5.13
359 -1.30
360 -0.43
361 -1.08
362 -0.38
363 -0.42
364 0.30
365 0.36
366 0.32
367 0.59
368 -0.03
369 0.78
370 0.55
371 -0.56
372 0.83
373 0.68
374 0.24
375 0.52
376 -0.73
377 -0.37
378 -1.03
379 0.79
380 0.75
381 1.17
382 1.19
383 1.13
384 0.97
385 -1.45
386 -0.51
387 -0.58
388 -0.19
389 -0.89
390 -0.90
391 -0.36
392 -0.79
393 -0.36
394 0.10
395 -0.23
396 1.05
397 -0.88
398 0.81
399 0.22
400 -1.26
401 -0.62
402 -0.22
403 -0.73
404 -0.85
405 -0.03
406 -0.17
407 0.23
408 0.31
409 -0.10
410 0.18
411 -1.77
412 0.10
413 0.27
414 0.07
415 -0.33
416 0.08
417 -0.15
418 -0.30
419 -1.58
420 -1.16
421 0.13
422 -0.26
423 0.54
424 -0.77
425 -0.78
426 -0.01
427 -1.34
428 -0.37
429 -0.15
430 -0.26
431 -1.17
432 -0.87
433 -0.04
434 -1.08
435 -1.86
436 -1.57
437 0.92
438 0.21
439 -0.74
440 0.53
441 -0.97
442 -2.07
443 -0.89
444 0.52
446 -0.70
447 -1.72
448 1.14
449 1.03
450 0.18
451 -0.88
452 -0.66
453 -2.26
454 -0.68
455 -1.75
456 -0.77
457 -0.98
458 -0.94
459 -0.33
460 0.37
461 -0.04
462 0.82
463 -0.52
464 -0.22
465 -0.47
466 -5.13
467 0.60
468 -1.35
469 -1.43
470 0.71
471 0.82
472 1.15
473 -0.15
475 0.00
476 0.07
477 -2.46
478 -1.15
479 -3.49
480 0.42
481 0.39
482 0.75
483 0.82
484 -0.41
485 0.27
486 0.11
487 0.47
488 0.39
489 0.02
490 0.60
491 0.23
492 0.52
493 0.77
494 0.29
495 -1.37
496 0.14
497 0.02
498 -0.49
509 0.00
510 0.13
511 0.66
512 -0.93
513 0.70
514 -0.18
515 0.58
516 -0.04
517 0.23
518 -0.08
519 -0.47
520 -1.29
521 -2.24
522 -0.69
523 -1.28
524 -0.39
525 -0.39
526 -5.13
527 -1.30
528 -0.43
529 -1.08
530 -0.38
531 -0.42
532 0.45
533 0.31
534 0.32
535 0.53
536 -0.18
537 0.87
538 0.55
539 -0.56
540 0.83
541 0.68
542 0.08
543 0.52
544 -0.93
545 -0.37
546 -1.03
547 0.79
548 0.75
549 1.17
550 1.19
551 1.20
552 0.97
553 -1.45
554 -0.51
555 -0.44
556 -0.19
557 -0.89
558 -0.90
559 -0.36
560 -0.79
561 0.00
562 -0.11
563 -0.23
564 1.05
565 -0.59
566 0.81
567 -0.02
568 -1.26
569 -0.66
570 -0.22
571 -0.66
572 -0.85
573 -0.03
574 -0.17
575 0.23
576 0.31
577 -0.10
578 0.18
579 -1.77
580 0.10
581 0.27
582 0.07
583 -0.33
584 0.08
585 -0.13
586 -0.30
587 -1.58
588 -1.16
589 0.13
590 -0.26
591 0.54
592 -0.77
593 -0.54
594 -0.01
595 -1.34
596 0.09
597 0.20
598 -0.26
599 -0.30
600 -0.87
601 -0.04
602 -0.76
603 -1.86
604 -1.57
605 0.92
606 0.21
607 -0.74
608 0.53
609 -0.97
610 -2.07
611 -0.89
612 0.52
614 -0.70
615 -1.72
616 1.14
617 1.03
618 0.18
619 -0.88
620 -0.66
621 -2.26
622 -0.68
623 -1.75
624 -0.77
625 -0.98
626 -0.94
627 -0.33
628 0.64
629 -0.04
630 0.82
631 -0.52
632 -0.22
633 -0.53
634 -5.13
635 0.60
636 -1.35
637 -1.39
638 0.71
639 0.82
640 1.15
641 -0.15
643 0.00
644 0.07
645 -2.46
646 -0.96
647 -3.49
648 0.42
649 0.39
650 0.75
651 0.82
652 -0.41
653 0.27
654 0.11
655 0.47
656 0.39
657 0.02
658 0.65
659 0.42
660 0.52
661 0.77
662 0.22
663 -1.37
664 0.14
665 0.02
666 0.76
#Reported_Model_Average -0.298
#Overall_Average_Reported -0.298
Output from Verify3D
Verify3D Score over a window of $winsize_s residues
JPEG image for Verify3D Score
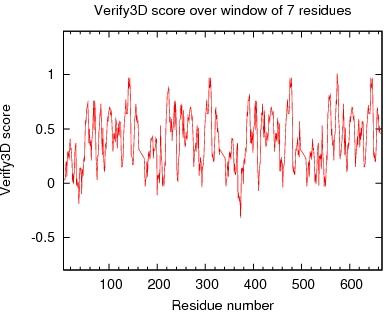
Table of Verify3D scores for ordered residues across all models
#verify3d
#Residue\Model only_model
5 0.14
6 0.08
7 0.24
8 0.24
9 -0.74
10 -0.09
11 0.34
12 0.28
13 0.93
14 0.44
15 -0.74
16 0.29
17 0.47
18 0.08
19 0.51
20 0.49
21 1.10
22 -0.07
23 -0.41
24 0.51
25 -0.41
26 0.04
27 -0.68
28 1.62
29 -0.74
30 0.25
31 0.71
32 0.77
33 0.47
34 -0.46
35 -1.13
36 -0.43
37 -0.03
38 0.59
39 1.06
40 -0.94
41 0.24
42 0.52
43 -0.74
44 0.28
45 0.51
46 -0.56
47 0.47
48 0.41
49 0.14
50 0.23
51 0.41
52 0.34
53 1.12
54 1.40
55 0.24
56 1.06
57 0.28
58 0.59
59 0.51
60 -0.10
61 0.04
62 1.10
63 0.08
64 0.24
65 0.96
66 -0.84
67 1.10
68 0.47
69 1.29
70 1.12
71 1.25
72 -0.94
73 1.04
74 0.51
75 -0.68
76 -0.68
77 0.47
78 0.52
79 0.28
80 1.40
81 0.51
82 0.93
83 -0.59
84 1.40
85 0.51
86 0.93
87 0.44
88 -0.94
89 0.08
90 1.14
91 -0.11
92 0.08
93 0.08
94 0.49
95 0.59
96 0.28
97 0.93
98 0.49
99 1.00
100 0.44
101 0.28
102 1.06
103 0.51
104 1.10
105 0.47
106 0.55
107 0.49
108 0.47
109 0.23
110 -0.43
111 0.24
112 1.10
113 1.10
114 0.08
115 0.93
116 -0.35
117 1.06
118 0.08
119 0.23
120 0.20
121 1.40
122 0.47
123 0.44
124 0.29
125 0.96
126 0.14
127 0.24
128 -0.26
129 -0.74
130 0.44
131 0.47
132 1.40
133 1.10
134 1.06
135 0.49
136 -0.61
137 1.06
138 0.91
139 0.49
140 1.06
141 1.10
142 1.06
143 1.10
144 0.44
145 0.96
146 1.06
147 0.14
148 -0.74
149 -1.13
150 0.93
151 0.44
152 0.51
153 1.06
154 0.81
155 0.25
156 0.08
157 1.10
158 1.00
159 0.81
160 -0.03
161 1.04
162 -0.10
173 0.14
174 0.08
175 0.24
176 0.24
177 -0.74
178 -0.09
179 0.34
180 0.28
181 0.93
182 0.44
183 -0.74
184 0.29
185 0.47
186 0.08
187 0.51
188 0.49
189 1.10
190 -0.07
191 0.24
192 0.51
193 0.24
194 0.28
195 -0.68
196 1.62
197 -0.09
198 0.25
199 0.71
200 0.77
201 0.08
202 -0.46
203 -2.01
204 1.14
205 0.25
206 0.59
207 1.06
208 -0.54
209 0.71
210 0.52
211 -0.40
212 -0.46
213 0.51
214 -0.56
215 0.08
216 0.51
217 0.14
218 0.23
219 0.41
220 0.34
221 1.62
222 1.40
223 0.71
224 1.06
225 0.28
226 0.59
227 0.51
228 0.47
229 0.28
230 1.10
231 0.08
232 0.71
233 0.96
234 -0.84
235 1.10
236 0.47
237 1.29
238 1.12
239 1.25
240 -0.94
241 1.04
242 0.23
243 -1.14
244 -0.68
245 0.47
246 1.14
247 0.28
248 1.40
249 0.51
250 0.93
251 -0.59
252 1.40
253 0.51
254 0.93
255 0.44
256 -0.28
257 0.55
258 1.14
259 -0.11
260 0.08
261 0.55
262 0.49
263 0.59
264 0.28
265 0.93
266 0.49
267 1.00
268 0.44
269 0.28
270 1.06
271 0.51
272 1.10
273 0.47
274 0.55
275 0.14
276 0.47
277 0.23
278 -0.43
279 -0.41
280 1.10
281 1.10
282 0.08
283 0.93
284 -0.35
285 0.77
286 0.08
287 0.51
288 0.20
289 1.40
290 -2.12
291 0.44
292 0.29
293 0.96
294 0.14
295 0.24
296 -0.26
297 -0.74
298 0.44
299 0.47
300 1.40
301 1.10
302 1.06
303 0.49
304 -0.61
305 1.06
306 0.91
307 0.49
308 1.06
309 1.10
310 1.06
311 1.10
312 0.44
313 0.96
314 1.06
315 0.14
316 -0.74
317 -1.13
318 0.93
319 0.64
320 0.51
321 1.06
322 0.81
323 0.25
324 0.47
325 1.10
326 -0.09
327 0.81
328 -0.03
329 1.04
330 -0.10
341 0.14
342 0.08
343 0.24
344 0.24
345 -0.74
346 -0.09
347 0.34
348 0.28
349 0.93
350 0.44
351 -0.74
352 0.29
353 0.47
354 0.08
355 0.51
356 0.49
357 1.10
358 -0.07
359 0.24
360 0.51
361 -0.41
362 0.04
363 -0.68
364 1.62
365 -0.74
366 0.25
367 0.71
368 0.77
369 0.47
370 -0.46
371 -2.01
372 -0.43
373 -0.03
374 0.59
375 1.06
376 -0.94
377 0.24
378 0.52
379 -0.74
380 0.28
381 0.51
382 0.09
383 0.47
384 0.41
385 0.14
386 0.23
387 0.41
388 0.34
389 1.62
390 1.40
391 0.24
392 1.06
393 0.28
394 0.59
395 0.51
396 -0.10
397 0.04
398 1.10
399 0.08
400 0.71
401 0.96
402 -0.84
403 1.10
404 0.47
405 1.29
406 1.12
407 1.25
408 -0.94
409 1.04
410 0.23
411 -1.14
412 -0.68
413 0.47
414 0.52
415 0.28
416 1.40
417 0.51
418 0.93
419 -0.59
420 1.40
421 0.51
422 0.93
423 0.44
424 -0.28
425 0.55
426 1.14
427 -0.11
428 0.08
429 0.55
430 0.49
431 0.59
432 0.28
433 0.93
434 0.49
435 1.00
436 0.44
437 0.28
438 1.06
439 0.51
440 1.10
441 0.47
442 0.55
443 0.49
444 0.47
445 0.23
446 -0.43
447 0.24
448 1.10
449 1.10
450 0.08
451 0.93
452 -0.35
453 1.06
454 0.08
455 0.51
456 0.20
457 1.40
458 -2.12
459 0.44
460 0.29
461 0.96
462 0.14
463 0.24
464 -0.26
465 -0.74
466 0.44
467 0.47
468 1.40
469 1.10
470 1.06
471 0.49
472 -0.61
473 1.06
474 0.91
475 0.49
476 1.06
477 1.10
478 1.06
479 1.10
480 0.44
481 0.96
482 1.06
483 0.14
484 -0.74
485 -2.01
486 0.93
487 0.64
488 0.51
489 1.06
490 0.81
491 0.25
492 -2.12
493 1.10
494 1.00
495 0.81
496 -0.03
497 1.04
498 -0.10
509 0.14
510 0.08
511 0.24
512 0.24
513 -0.74
514 -0.09
515 0.34
516 0.28
517 0.93
518 0.44
519 -0.74
520 0.29
521 0.47
522 0.08
523 0.51
524 0.49
525 1.10
526 -0.07
527 -0.41
528 0.51
529 -0.41
530 0.04
531 -0.68
532 1.62
533 -0.74
534 0.25
535 0.71
536 0.77
537 0.47
538 -0.46
539 -1.13
540 1.14
541 0.25
542 0.59
543 1.06
544 -0.54
545 0.71
546 0.52
547 -0.40
548 -0.46
549 0.51
550 -0.56
551 0.08
552 0.51
553 0.14
554 0.51
555 0.51
556 0.34
557 1.12
558 1.40
559 0.71
560 1.06
561 0.28
562 0.59
563 0.51
564 -0.10
565 0.04
566 1.10
567 0.08
568 0.24
569 0.96
570 -0.84
571 1.10
572 0.47
573 1.29
574 1.12
575 1.25
576 0.81
577 1.04
578 0.51
579 0.29
580 -0.68
581 0.47
582 0.52
583 -0.59
584 1.40
585 0.51
586 0.93
587 -0.59
588 1.40
589 0.51
590 0.93
591 0.44
592 -0.94
593 0.08
594 1.14
595 -0.11
596 0.08
597 0.08
598 0.49
599 0.59
600 0.28
601 0.93
602 0.49
603 1.00
604 0.44
605 0.28
606 1.06
607 0.51
608 1.10
609 0.47
610 0.55
611 0.14
612 0.47
613 0.23
614 -0.43
615 -0.41
616 1.10
617 1.10
618 0.47
619 0.93
620 -0.35
621 0.77
622 0.08
623 0.23
624 0.20
625 1.40
626 0.47
627 0.44
628 0.29
629 0.96
630 0.14
631 0.24
632 -0.26
633 -0.74
634 0.44
635 0.47
636 1.04
637 1.10
638 1.06
639 0.49
640 -0.61
641 1.06
642 0.91
643 0.49
644 1.06
645 1.10
646 1.06
647 1.10
648 0.44
649 0.96
650 1.06
651 0.14
652 -0.74
653 -1.13
654 0.93
655 0.64
656 0.51
657 1.06
658 0.81
659 0.25
660 0.47
661 1.10
662 -0.09
663 0.81
664 -0.03
665 1.04
666 -0.10
#Reported_Model_Average 0.394
#Overall_Average_Reported 0.394
Output from ProsaII
ProsaII Score over a window of $winsize_s residues
JPEG image for ProsaII Score
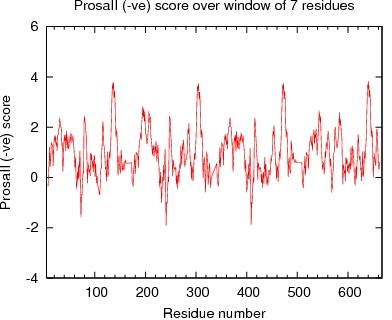
Table of Verify3D scores for ordered residues across all models
#verify3d
#Residue\Model only_model
5 0.14
6 0.08
7 0.24
8 0.24
9 -0.74
10 -0.09
11 0.34
12 0.28
13 0.93
14 0.44
15 -0.74
16 0.29
17 0.47
18 0.08
19 0.51
20 0.49
21 1.10
22 -0.07
23 -0.41
24 0.51
25 -0.41
26 0.04
27 -0.68
28 1.62
29 -0.74
30 0.25
31 0.71
32 0.77
33 0.47
34 -0.46
35 -1.13
36 -0.43
37 -0.03
38 0.59
39 1.06
40 -0.94
41 0.24
42 0.52
43 -0.74
44 0.28
45 0.51
46 -0.56
47 0.47
48 0.41
49 0.14
50 0.23
51 0.41
52 0.34
53 1.12
54 1.40
55 0.24
56 1.06
57 0.28
58 0.59
59 0.51
60 -0.10
61 0.04
62 1.10
63 0.08
64 0.24
65 0.96
66 -0.84
67 1.10
68 0.47
69 1.29
70 1.12
71 1.25
72 -0.94
73 1.04
74 0.51
75 -0.68
76 -0.68
77 0.47
78 0.52
79 0.28
80 1.40
81 0.51
82 0.93
83 -0.59
84 1.40
85 0.51
86 0.93
87 0.44
88 -0.94
89 0.08
90 1.14
91 -0.11
92 0.08
93 0.08
94 0.49
95 0.59
96 0.28
97 0.93
98 0.49
99 1.00
100 0.44
101 0.28
102 1.06
103 0.51
104 1.10
105 0.47
106 0.55
107 0.49
108 0.47
109 0.23
110 -0.43
111 0.24
112 1.10
113 1.10
114 0.08
115 0.93
116 -0.35
117 1.06
118 0.08
119 0.23
120 0.20
121 1.40
122 0.47
123 0.44
124 0.29
125 0.96
126 0.14
127 0.24
128 -0.26
129 -0.74
130 0.44
131 0.47
132 1.40
133 1.10
134 1.06
135 0.49
136 -0.61
137 1.06
138 0.91
139 0.49
140 1.06
141 1.10
142 1.06
143 1.10
144 0.44
145 0.96
146 1.06
147 0.14
148 -0.74
149 -1.13
150 0.93
151 0.44
152 0.51
153 1.06
154 0.81
155 0.25
156 0.08
157 1.10
158 1.00
159 0.81
160 -0.03
161 1.04
162 -0.10
173 0.14
174 0.08
175 0.24
176 0.24
177 -0.74
178 -0.09
179 0.34
180 0.28
181 0.93
182 0.44
183 -0.74
184 0.29
185 0.47
186 0.08
187 0.51
188 0.49
189 1.10
190 -0.07
191 0.24
192 0.51
193 0.24
194 0.28
195 -0.68
196 1.62
197 -0.09
198 0.25
199 0.71
200 0.77
201 0.08
202 -0.46
203 -2.01
204 1.14
205 0.25
206 0.59
207 1.06
208 -0.54
209 0.71
210 0.52
211 -0.40
212 -0.46
213 0.51
214 -0.56
215 0.08
216 0.51
217 0.14
218 0.23
219 0.41
220 0.34
221 1.62
222 1.40
223 0.71
224 1.06
225 0.28
226 0.59
227 0.51
228 0.47
229 0.28
230 1.10
231 0.08
232 0.71
233 0.96
234 -0.84
235 1.10
236 0.47
237 1.29
238 1.12
239 1.25
240 -0.94
241 1.04
242 0.23
243 -1.14
244 -0.68
245 0.47
246 1.14
247 0.28
248 1.40
249 0.51
250 0.93
251 -0.59
252 1.40
253 0.51
254 0.93
255 0.44
256 -0.28
257 0.55
258 1.14
259 -0.11
260 0.08
261 0.55
262 0.49
263 0.59
264 0.28
265 0.93
266 0.49
267 1.00
268 0.44
269 0.28
270 1.06
271 0.51
272 1.10
273 0.47
274 0.55
275 0.14
276 0.47
277 0.23
278 -0.43
279 -0.41
280 1.10
281 1.10
282 0.08
283 0.93
284 -0.35
285 0.77
286 0.08
287 0.51
288 0.20
289 1.40
290 -2.12
291 0.44
292 0.29
293 0.96
294 0.14
295 0.24
296 -0.26
297 -0.74
298 0.44
299 0.47
300 1.40
301 1.10
302 1.06
303 0.49
304 -0.61
305 1.06
306 0.91
307 0.49
308 1.06
309 1.10
310 1.06
311 1.10
312 0.44
313 0.96
314 1.06
315 0.14
316 -0.74
317 -1.13
318 0.93
319 0.64
320 0.51
321 1.06
322 0.81
323 0.25
324 0.47
325 1.10
326 -0.09
327 0.81
328 -0.03
329 1.04
330 -0.10
341 0.14
342 0.08
343 0.24
344 0.24
345 -0.74
346 -0.09
347 0.34
348 0.28
349 0.93
350 0.44
351 -0.74
352 0.29
353 0.47
354 0.08
355 0.51
356 0.49
357 1.10
358 -0.07
359 0.24
360 0.51
361 -0.41
362 0.04
363 -0.68
364 1.62
365 -0.74
366 0.25
367 0.71
368 0.77
369 0.47
370 -0.46
371 -2.01
372 -0.43
373 -0.03
374 0.59
375 1.06
376 -0.94
377 0.24
378 0.52
379 -0.74
380 0.28
381 0.51
382 0.09
383 0.47
384 0.41
385 0.14
386 0.23
387 0.41
388 0.34
389 1.62
390 1.40
391 0.24
392 1.06
393 0.28
394 0.59
395 0.51
396 -0.10
397 0.04
398 1.10
399 0.08
400 0.71
401 0.96
402 -0.84
403 1.10
404 0.47
405 1.29
406 1.12
407 1.25
408 -0.94
409 1.04
410 0.23
411 -1.14
412 -0.68
413 0.47
414 0.52
415 0.28
416 1.40
417 0.51
418 0.93
419 -0.59
420 1.40
421 0.51
422 0.93
423 0.44
424 -0.28
425 0.55
426 1.14
427 -0.11
428 0.08
429 0.55
430 0.49
431 0.59
432 0.28
433 0.93
434 0.49
435 1.00
436 0.44
437 0.28
438 1.06
439 0.51
440 1.10
441 0.47
442 0.55
443 0.49
444 0.47
445 0.23
446 -0.43
447 0.24
448 1.10
449 1.10
450 0.08
451 0.93
452 -0.35
453 1.06
454 0.08
455 0.51
456 0.20
457 1.40
458 -2.12
459 0.44
460 0.29
461 0.96
462 0.14
463 0.24
464 -0.26
465 -0.74
466 0.44
467 0.47
468 1.40
469 1.10
470 1.06
471 0.49
472 -0.61
473 1.06
474 0.91
475 0.49
476 1.06
477 1.10
478 1.06
479 1.10
480 0.44
481 0.96
482 1.06
483 0.14
484 -0.74
485 -2.01
486 0.93
487 0.64
488 0.51
489 1.06
490 0.81
491 0.25
492 -2.12
493 1.10
494 1.00
495 0.81
496 -0.03
497 1.04
498 -0.10
509 0.14
510 0.08
511 0.24
512 0.24
513 -0.74
514 -0.09
515 0.34
516 0.28
517 0.93
518 0.44
519 -0.74
520 0.29
521 0.47
522 0.08
523 0.51
524 0.49
525 1.10
526 -0.07
527 -0.41
528 0.51
529 -0.41
530 0.04
531 -0.68
532 1.62
533 -0.74
534 0.25
535 0.71
536 0.77
537 0.47
538 -0.46
539 -1.13
540 1.14
541 0.25
542 0.59
543 1.06
544 -0.54
545 0.71
546 0.52
547 -0.40
548 -0.46
549 0.51
550 -0.56
551 0.08
552 0.51
553 0.14
554 0.51
555 0.51
556 0.34
557 1.12
558 1.40
559 0.71
560 1.06
561 0.28
562 0.59
563 0.51
564 -0.10
565 0.04
566 1.10
567 0.08
568 0.24
569 0.96
570 -0.84
571 1.10
572 0.47
573 1.29
574 1.12
575 1.25
576 0.81
577 1.04
578 0.51
579 0.29
580 -0.68
581 0.47
582 0.52
583 -0.59
584 1.40
585 0.51
586 0.93
587 -0.59
588 1.40
589 0.51
590 0.93
591 0.44
592 -0.94
593 0.08
594 1.14
595 -0.11
596 0.08
597 0.08
598 0.49
599 0.59
600 0.28
601 0.93
602 0.49
603 1.00
604 0.44
605 0.28
606 1.06
607 0.51
608 1.10
609 0.47
610 0.55
611 0.14
612 0.47
613 0.23
614 -0.43
615 -0.41
616 1.10
617 1.10
618 0.47
619 0.93
620 -0.35
621 0.77
622 0.08
623 0.23
624 0.20
625 1.40
626 0.47
627 0.44
628 0.29
629 0.96
630 0.14
631 0.24
632 -0.26
633 -0.74
634 0.44
635 0.47
636 1.04
637 1.10
638 1.06
639 0.49
640 -0.61
641 1.06
642 0.91
643 0.49
644 1.06
645 1.10
646 1.06
647 1.10
648 0.44
649 0.96
650 1.06
651 0.14
652 -0.74
653 -1.13
654 0.93
655 0.64
656 0.51
657 1.06
658 0.81
659 0.25
660 0.47
661 1.10
662 -0.09
663 0.81
664 -0.03
665 1.04
666 -0.10
#Reported_Model_Average 0.394
#Overall_Average_Reported 0.394
Output from MolProbity
VdW violations from MAGE
JPEG image for MAGE VdW violation
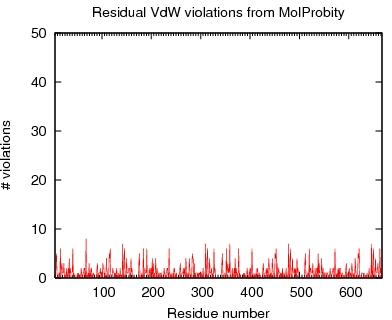
Table of MAGE VdW violations for ordered residues across all models
#mage_clash
#Residue\Model only_model
5.000 0
6.000 0
7.000 3
8.000 5
9.000 1
10.000 0
11.000 0
12.000 0
13.000 1
14.000 1
15.000 0
16.000 6
17.000 0
18.000 2
19.000 3
20.000 1
21.000 1
22.000 1
23.000 3
24.000 0
25.000 0
26.000 0
27.000 0
28.000 2
29.000 0
30.000 0
31.000 1
32.000 2
33.000 0
34.000 4
35.000 0
36.000 2
37.000 1
38.000 1
39.000 1
40.000 1
41.000 6
42.000 0
43.000 1
44.000 0
45.000 0
46.000 0
47.000 0
48.000 0
49.000 0
50.000 0
51.000 1
52.000 0
53.000 1
54.000 0
55.000 0
56.000 0
57.000 0
58.000 2
59.000 1
60.000 0
61.000 0
62.000 1
63.000 1
64.000 2
65.000 2
66.000 0
67.000 0
68.000 8
69.000 1
70.000 0
71.000 0
72.000 1
73.000 0
74.000 3
75.000 0
76.000 1
77.000 0
78.000 2
79.000 2
80.000 0
81.000 0
82.000 1
83.000 1
84.000 0
85.000 0
86.000 0
87.000 0
88.000 0
89.000 0
90.000 2
91.000 3
92.000 0
93.000 0
94.000 1
95.000 1
96.000 1
97.000 2
98.000 0
99.000 1
100.000 1
101.000 0
102.000 3
103.000 2
104.000 2
105.000 0
106.000 1
107.000 0
108.000 0
109.000 4
110.000 2
111.000 1
112.000 0
113.000 0
114.000 1
115.000 5
116.000 4
117.000 6
118.000 0
119.000 0
120.000 0
121.000 1
122.000 2
123.000 1
124.000 1
125.000 1
126.000 0
127.000 1
128.000 0
129.000 1
130.000 2
131.000 0
132.000 0
133.000 1
134.000 0
135.000 0
136.000 0
137.000 2
138.000 0
139.000 0
140.000 0
141.000 0
142.000 7
143.000 0
144.000 0
145.000 4
146.000 6
147.000 0
148.000 0
149.000 1
150.000 4
151.000 1
152.000 1
153.000 1
154.000 2
155.000 2
156.000 2
157.000 0
158.000 0
159.000 4
160.000 1
161.000 2
162.000 0
163.000 0
164.000 0
165.000 0
166.000 0
167.000 0
168.000 0
169.000 0
170.000 0
171.000 0
172.000 0
173.000 0
174.000 0
175.000 3
176.000 5
177.000 1
178.000 0
179.000 0
180.000 0
181.000 1
182.000 0
183.000 0
184.000 6
185.000 0
186.000 1
187.000 2
188.000 1
189.000 1
190.000 1
191.000 6
192.000 0
193.000 0
194.000 0
195.000 0
196.000 2
197.000 1
198.000 0
199.000 2
200.000 2
201.000 0
202.000 4
203.000 0
204.000 3
205.000 1
206.000 0
207.000 1
208.000 1
209.000 4
210.000 0
211.000 1
212.000 2
213.000 0
214.000 0
215.000 1
216.000 1
217.000 0
218.000 0
219.000 1
220.000 1
221.000 1
222.000 0
223.000 0
224.000 0
225.000 0
226.000 2
227.000 1
228.000 0
229.000 0
230.000 1
231.000 1
232.000 2
233.000 2
234.000 0
235.000 0
236.000 6
237.000 1
238.000 0
239.000 0
240.000 1
241.000 0
242.000 0
243.000 0
244.000 1
245.000 0
246.000 2
247.000 2
248.000 0
249.000 0
250.000 1
251.000 1
252.000 0
253.000 0
254.000 0
255.000 0
256.000 0
257.000 0
258.000 2
259.000 3
260.000 0
261.000 0
262.000 1
263.000 1
264.000 1
265.000 2
266.000 0
267.000 2
268.000 1
269.000 0
270.000 4
271.000 2
272.000 0
273.000 0
274.000 1
275.000 0
276.000 0
277.000 4
278.000 2
279.000 0
280.000 0
281.000 0
282.000 1
283.000 5
284.000 4
285.000 4
286.000 0
287.000 3
288.000 0
289.000 0
290.000 1
291.000 1
292.000 1
293.000 1
294.000 0
295.000 1
296.000 0
297.000 1
298.000 1
299.000 0
300.000 0
301.000 1
302.000 0
303.000 0
304.000 0
305.000 2
306.000 0
307.000 0
308.000 0
309.000 0
310.000 7
311.000 0
312.000 0
313.000 4
314.000 6
315.000 0
316.000 0
317.000 1
318.000 4
319.000 1
320.000 1
321.000 1
322.000 2
323.000 0
324.000 1
325.000 0
326.000 0
327.000 6
328.000 1
329.000 2
330.000 0
331.000 0
332.000 0
333.000 0
334.000 0
335.000 0
336.000 0
337.000 0
338.000 0
339.000 0
340.000 0
341.000 0
342.000 0
343.000 3
344.000 5
345.000 1
346.000 0
347.000 0
348.000 0
349.000 1
350.000 0
351.000 0
352.000 6
353.000 0
354.000 2
355.000 3
356.000 1
357.000 2
358.000 1
359.000 7
360.000 0
361.000 0
362.000 0
363.000 0
364.000 2
365.000 0
366.000 0
367.000 1
368.000 2
369.000 0
370.000 4
371.000 0
372.000 2
373.000 1
374.000 0
375.000 1
376.000 1
377.000 6
378.000 0
379.000 1
380.000 0
381.000 0
382.000 0
383.000 0
384.000 0
385.000 0
386.000 0
387.000 1
388.000 0
389.000 1
390.000 0
391.000 0
392.000 0
393.000 0
394.000 2
395.000 1
396.000 0
397.000 0
398.000 1
399.000 1
400.000 2
401.000 2
402.000 0
403.000 0
404.000 6
405.000 1
406.000 0
407.000 0
408.000 1
409.000 0
410.000 0
411.000 0
412.000 1
413.000 0
414.000 2
415.000 2
416.000 0
417.000 0
418.000 1
419.000 1
420.000 0
421.000 0
422.000 0
423.000 0
424.000 0
425.000 0
426.000 2
427.000 3
428.000 0
429.000 0
430.000 1
431.000 1
432.000 1
433.000 2
434.000 0
435.000 2
436.000 1
437.000 0
438.000 4
439.000 2
440.000 2
441.000 0
442.000 1
443.000 0
444.000 0
445.000 4
446.000 2
447.000 1
448.000 0
449.000 0
450.000 1
451.000 5
452.000 4
453.000 6
454.000 0
455.000 3
456.000 0
457.000 1
458.000 2
459.000 1
460.000 1
461.000 1
462.000 0
463.000 1
464.000 0
465.000 1
466.000 2
467.000 0
468.000 0
469.000 1
470.000 0
471.000 0
472.000 0
473.000 2
474.000 0
475.000 0
476.000 0
477.000 0
478.000 7
479.000 0
480.000 0
481.000 4
482.000 6
483.000 0
484.000 0
485.000 1
486.000 4
487.000 1
488.000 1
489.000 1
490.000 2
491.000 0
492.000 2
493.000 0
494.000 0
495.000 4
496.000 1
497.000 2
498.000 0
499.000 0
500.000 0
501.000 0
502.000 0
503.000 0
504.000 0
505.000 0
506.000 0
507.000 0
508.000 0
509.000 0
510.000 0
511.000 3
512.000 5
513.000 1
514.000 0
515.000 0
516.000 0
517.000 1
518.000 1
519.000 0
520.000 6
521.000 0
522.000 2
523.000 2
524.000 1
525.000 1
526.000 1
527.000 3
528.000 0
529.000 0
530.000 0
531.000 0
532.000 2
533.000 0
534.000 0
535.000 2
536.000 2
537.000 0
538.000 5
539.000 0
540.000 3
541.000 1
542.000 1
543.000 1
544.000 1
545.000 4
546.000 0
547.000 1
548.000 2
549.000 0
550.000 0
551.000 1
552.000 1
553.000 0
554.000 0
555.000 1
556.000 1
557.000 1
558.000 0
559.000 0
560.000 0
561.000 0
562.000 2
563.000 1
564.000 0
565.000 0
566.000 1
567.000 1
568.000 2
569.000 2
570.000 0
571.000 0
572.000 6
573.000 1
574.000 0
575.000 0
576.000 1
577.000 0
578.000 2
579.000 0
580.000 2
581.000 0
582.000 2
583.000 2
584.000 0
585.000 0
586.000 1
587.000 1
588.000 0
589.000 0
590.000 0
591.000 0
592.000 0
593.000 0
594.000 2
595.000 3
596.000 0
597.000 0
598.000 1
599.000 1
600.000 1
601.000 2
602.000 0
603.000 1
604.000 1
605.000 0
606.000 3
607.000 2
608.000 0
609.000 0
610.000 1
611.000 0
612.000 0
613.000 4
614.000 2
615.000 0
616.000 0
617.000 0
618.000 1
619.000 5
620.000 4
621.000 6
622.000 0
623.000 0
624.000 0
625.000 0
626.000 1
627.000 1
628.000 1
629.000 1
630.000 0
631.000 1
632.000 0
633.000 0
634.000 0
635.000 0
636.000 0
637.000 1
638.000 0
639.000 0
640.000 0
641.000 2
642.000 0
643.000 0
644.000 0
645.000 0
646.000 7
647.000 0
648.000 0
649.000 4
650.000 6
651.000 0
652.000 0
653.000 1
654.000 4
655.000 1
656.000 1
657.000 1
658.000 2
659.000 3
660.000 1
661.000 0
662.000 0
663.000 6
664.000 1
665.000 2
666.000 0
#Reported_Model_Average 1.048
#Overall_Average_Reported 1.048
List of bad contacts calculated by MAGE
/farm/software/bin/probe
: 10404:M 74 ASP HA :M 659 GLN 2HE2 : -0.981: 12
: 10404:M 74 ASP HA :M 659 GLN NE2 : -0.761: 12
: 10404:M 659 GLN 2HE2 :M 74 ASP CA : -0.412: 12
: 10404:M 344 ARG 2HH1 :M 343 ARG 1HD : -0.959: 75
: 10404:M 343 ARG 1HD :M 344 ARG NH1 : -0.572: 75
: 10404:M 344 ARG HA :M 344 ARG NE : -0.516: 78
: 10404:M 343 ARG 2HB :M 344 ARG NH1 : -0.459: 73
: 10404:M 175 ARG 1HD :M 176 ARG 2HH1 : -0.948: 73
: 10404:M 175 ARG 1HD :M 176 ARG NH1 : -0.568: 73
: 10404:M 176 ARG HA :M 176 ARG NE : -0.497: 77
: 10404:M 176 ARG NH1 :M 175 ARG 2HB : -0.436: 73
: 10404:M 155 GLN 2HE2 :M 578 ASP HA : -0.942: 19
: 10404:M 578 ASP HA :M 155 GLN NE2 : -0.752: 19
: 10404:M 7 ARG 1HD :M 8 ARG 2HH1 : -0.925: 78
: 10404:M 7 ARG 1HD :M 8 ARG NH1 : -0.542: 78
: 10404:M 8 ARG NE :M 8 ARG HA : -0.498: 78
: 10404:M 7 ARG 2HB :M 8 ARG NH1 : -0.463: 73
: 10404:M 142 LEU HG :M 97 ILE 1HG2 : -0.910: 26
: 10404:M 95 PRO 1HD :M 137 LEU 2HD2 : -0.752: 19
: 10404:M 159 ILE 3HD1 :M 159 ILE C : -0.619: 41
: 10404:M 146 LEU HG :M 142 LEU 2HD2 : -0.571: 20
: 10404:M 82 ILE 1HG2 :M 142 LEU 1HD1 : -0.566: 24
: 10404:M 154 ILE 1HG1 :M 159 ILE 3HG2 : -0.556: 33
: 10404:M 160 GLN N :M 159 ILE 3HD1 : -0.535: 41
: 10404:M 97 ILE 1HD1 :M 137 LEU HG : -0.503: 36
: 10404:M 69 CYS SG :M 146 LEU 1HD1 : -0.475: 22
: 10404:M 146 LEU O :M 150 ILE 1HG1 : -0.469: 23
: 10404:M 150 ILE 1HD1 :M 146 LEU CD2 : -0.469: 18
: 10404:M 146 LEU CD1 :M 142 LEU 2HD2 : -0.456: 20
: 10404:M 154 ILE 2HG1 :M 150 ILE O : -0.429: 13
: 10404:M 142 LEU 2HD2 :M 146 LEU CG : -0.422: 20
: 10404:M 151 PRO CD :M 150 ILE HB : -0.415: 19
: 10404:M 142 LEU 3HD2 :M 142 LEU HA : -0.409: 20
: 10404:M 352 LEU 2HD1 :M 370 GLU 2HB : -0.908: 42
: 10404:M 370 GLU 2HB :M 352 LEU CD1 : -0.560: 42
: 10404:M 354 THR O :M 367 ARG 2HD : -0.518: 34
: 10404:M 370 GLU OE2 :M 354 THR OG1 : -0.517: 34
: 10404:M 352 LEU N :M 352 LEU 2HD2 : -0.482: 46
: 10404:M 352 LEU 2HD1 :M 370 GLU CB : -0.461: 42
: 10404:M 469 GLY 2HA :M 352 LEU 1HD2 : -0.429: 46
: 10404:M 646 LEU HG :M 601 ILE 1HG2 : -0.903: 27
: 10404:M 599 PRO 1HD :M 641 LEU 2HD2 : -0.760: 21
: 10404:M 663 ILE C :M 663 ILE 3HD1 : -0.613: 50
: 10404:M 586 ILE 1HG2 :M 646 LEU 1HD1 : -0.568: 26
: 10404:M 650 LEU HG :M 646 LEU 2HD2 : -0.558: 21
: 10404:M 664 GLN N :M 663 ILE 3HD1 : -0.554: 50
: 10404:M 601 ILE 1HD1 :M 641 LEU HG : -0.522: 43
: 10404:M 658 ILE 1HG1 :M 663 ILE 3HG2 : -0.512: 42
: 10404:M 573 CYS SG :M 650 LEU 1HD1 : -0.509: 21
: 10404:M 650 LEU CD2 :M 654 ILE 1HD1 : -0.462: 19
: 10404:M 646 LEU 2HD2 :M 650 LEU CD1 : -0.458: 21
: 10404:M 654 ILE 1HG1 :M 650 LEU O : -0.451: 28
: 10404:M 654 ILE O :M 658 ILE 2HG1 : -0.437: 13
: 10404:M 646 LEU 2HD2 :M 650 LEU CG : -0.426: 21
: 10404:M 655 PRO CD :M 654 ILE HB : -0.415: 19
: 10404:M 663 ILE C :M 663 ILE CD1 : -0.414: 50
: 10404:M 646 LEU HA :M 646 LEU 3HD2 : -0.402: 21
: 10404:M 34 GLU 2HB :M 16 LEU 2HD1 : -0.901: 44
: 10404:M 31 ARG 2HD :M 18 THR O : -0.535: 32
: 10404:M 34 GLU 2HB :M 16 LEU CD1 : -0.535: 44
: 10404:M 16 LEU 2HD2 :M 16 LEU N : -0.498: 49
: 10404:M 34 GLU OE2 :M 18 THR OG1 : -0.484: 39
: 10404:M 16 LEU 2HD1 :M 34 GLU CB : -0.467: 44
: 10404:M 16 LEU 1HD2 :M 133 GLY 2HA : -0.448: 49
: 10404:M 512 ARG 2HH1 :M 511 ARG 1HD : -0.895: 72
: 10404:M 512 ARG HA :M 512 ARG NE : -0.512: 74
: 10404:M 511 ARG 1HD :M 512 ARG NH1 : -0.509: 72
: 10404:M 512 ARG NH1 :M 511 ARG 2HB : -0.425: 72
: 10404:M 455 ASP 2HB :M 191 ARG 2HH2 : -0.893: 58
: 10404:M 191 ARG NH2 :M 455 ASP 2HB : -0.729: 58
: 10404:M 191 ARG H :M 189 GLY C : -0.526: 28
: 10404:M 191 ARG 2HH1 :M 455 ASP CG : -0.504: 61
: 10404:M 191 ARG 1HG :M 191 ARG 1HH2 : -0.404: 53
: 10404:M 433 ILE 1HG2 :M 478 LEU HG : -0.891: 31
: 10404:M 473 LEU 2HD2 :M 431 PRO 1HD : -0.777: 23
: 10404:M 495 ILE C :M 495 ILE 3HD1 : -0.605: 46
: 10404:M 418 ILE 1HG2 :M 478 LEU 1HD1 : -0.592: 22
: 10404:M 482 LEU HG :M 478 LEU 2HD2 : -0.565: 25
: 10404:M 490 ILE 1HG1 :M 495 ILE 3HG2 : -0.549: 34
: 10404:M 495 ILE 3HD1 :M 496 GLN N : -0.530: 46
: 10404:M 473 LEU HG :M 433 ILE 1HD1 : -0.521: 38
: 10404:M 486 ILE 1HD1 :M 482 LEU CD2 : -0.478: 15
: 10404:M 482 LEU CD1 :M 478 LEU 2HD2 : -0.469: 25
: 10404:M 486 ILE 1HG1 :M 482 LEU O : -0.459: 24
: 10404:M 405 CYS SG :M 482 LEU 1HD1 : -0.445: 24
: 10404:M 478 LEU 2HD2 :M 482 LEU CG : -0.436: 25
: 10404:M 478 LEU 3HD2 :M 478 LEU HA : -0.428: 25
: 10404:M 486 ILE HB :M 487 PRO CD : -0.415: 15
: 10404:M 490 ILE 2HG1 :M 486 ILE O : -0.412: 14
: 10404:M 265 ILE 1HG2 :M 310 LEU HG : -0.884: 21
: 10404:M 305 LEU 2HD2 :M 263 PRO 1HD : -0.760: 18
: 10404:M 327 ILE C :M 327 ILE 3HD1 : -0.613: 48
: 10404:M 250 ILE 1HG2 :M 310 LEU 1HD1 : -0.574: 22
: 10404:M 314 LEU HG :M 310 LEU 2HD2 : -0.573: 25
: 10404:M 328 GLN N :M 327 ILE 3HD1 : -0.561: 48
: 10404:M 327 ILE 3HG2 :M 322 ILE 1HG1 : -0.532: 42
: 10404:M 305 LEU HG :M 265 ILE 1HD1 : -0.510: 36
: 10404:M 237 CYS SG :M 314 LEU 1HD1 : -0.490: 22
: 10404:M 310 LEU 2HD2 :M 314 LEU CD1 : -0.466: 25
: 10404:M 318 ILE 1HD1 :M 314 LEU CD2 : -0.465: 18
: 10404:M 314 LEU O :M 318 ILE 1HG1 : -0.464: 21
: 10404:M 314 LEU CG :M 310 LEU 2HD2 : -0.445: 25
: 10404:M 322 ILE 2HG1 :M 318 ILE O : -0.422: 15
: 10404:M 310 LEU HA :M 310 LEU 3HD2 : -0.419: 25
: 10404:M 318 ILE HB :M 319 PRO CD : -0.411: 16
: 10404:M 327 ILE C :M 327 ILE CD1 : -0.406: 48
: 10404:M 520 LEU 2HD1 :M 538 GLU 2HB : -0.869: 43
: 10404:M 538 GLU 2HB :M 520 LEU CD1 : -0.529: 43
: 10404:M 535 ARG 2HD :M 522 THR O : -0.522: 34
: 10404:M 520 LEU 2HD2 :M 520 LEU N : -0.503: 39
: 10404:M 538 GLU OE2 :M 522 THR OG1 : -0.462: 26
: 10404:M 520 LEU 2HD1 :M 538 GLU CB : -0.426: 43
: 10404:M 520 LEU 1HD2 :M 637 GLY 2HA : -0.420: 39
: 10404:M 535 ARG O :M 538 GLU 1HB : -0.409: 33
: 10404:M 202 GLU 2HB :M 184 LEU 2HD1 : -0.864: 36
: 10404:M 202 GLU 2HB :M 184 LEU CD1 : -0.544: 36
: 10404:M 199 ARG 2HD :M 186 THR O : -0.536: 34
: 10404:M 184 LEU 2HD2 :M 184 LEU N : -0.524: 35
: 10404:M 184 LEU 2HD1 :M 202 GLU CB : -0.458: 36
: 10404:M 184 LEU 1HD2 :M 301 GLY 2HA : -0.415: 35
: 10404:M 202 GLU 1HB :M 199 ARG O : -0.400: 31
: 10404:M 359 ARG 2HH2 :M 287 ASP 2HB : -0.807: 59
: 10404:M 359 ARG NH2 :M 287 ASP 2HB : -0.690: 59
: 10404:M 359 ARG H :M 357 GLY C : -0.538: 32
: 10404:M 359 ARG 2HH1 :M 287 ASP CG : -0.496: 66
: 10404:M 359 ARG 1HG :M 359 ARG 1HH2 : -0.418: 59
: 10404:M 357 GLY O :M 359 ARG N : -0.406: 32
: 10404:M 400 ARG 1HD :M 399 THR OG1 : -0.768: 43
: 10404:M 400 ARG 2HD :M 395 ASN ND2 : -0.563: 43
: 10404:M 568 ARG 1HD :M 567 THR OG1 : -0.755: 46
: 10404:M 568 ARG 2HD :M 563 ASN ND2 : -0.578: 46
: 10404:M 231 THR OG1 :M 232 ARG 1HD : -0.743: 47
: 10404:M 232 ARG 2HD :M 227 ASN ND2 : -0.558: 47
: 10404:M 32 LEU HG :M 36 TYR CE2 : -0.738: 17
: 10404:M 36 TYR HE2 :M 32 LEU HG : -0.586: 17
: 10404:M 540 TYR CE2 :M 536 LEU HG : -0.731: 19
: 10404:M 540 TYR HE2 :M 536 LEU HG : -0.571: 19
: 10404:M 488 ASP OD1 :M 492 LYS 2HE : -0.452: 11
: 10404:M 540 TYR 2HB :M 492 LYS 2HD : -0.420: 6
: 10404:M 200 LEU HG :M 204 TYR CE2 : -0.727: 21
: 10404:M 204 TYR HE2 :M 200 LEU HG : -0.546: 21
: 10404:M 156 LYS 2HE :M 152 ASP OD1 : -0.469: 11
: 10404:M 204 TYR 2HB :M 156 LYS 2HD : -0.424: 14
: 10404:M 372 TYR CE2 :M 368 LEU HG : -0.720: 15
: 10404:M 372 TYR HE2 :M 368 LEU HG : -0.554: 15
: 10404:M 64 ARG 1HD :M 63 THR OG1 : -0.715: 43
: 10404:M 64 ARG 2HD :M 59 ASN ND2 : -0.573: 43
: 10404:M 442 THR HB :M 489 LEU 1HD2 : -0.677: 32
: 10404:M 68 LYS 2HB :M 68 LYS NZ : -0.677: 42
: 10404:M 68 LYS 2HD :M 79 GLU OE1 : -0.608: 59
: 10404:M 68 LYS 2HB :M 68 LYS 2HZ : -0.456: 42
: 10404:M 68 LYS CD :M 79 GLU OE1 : -0.448: 59
: 10404:M 68 LYS 2HB :M 68 LYS 3HZ : -0.407: 42
: 10404:M 100 PRO 1HG :M 83 GLU OE2 : -0.676: 37
: 10404:M 274 THR HB :M 321 LEU 1HD2 : -0.671: 34
: 10404:M 419 GLU OE2 :M 436 PRO 1HG : -0.667: 40
: 10404:M 106 THR HB :M 153 LEU 1HD2 : -0.665: 29
: 10404:M 251 GLU OE2 :M 268 PRO 1HG : -0.662: 30
: 10404:M 657 LEU 1HD2 :M 610 THR HB : -0.657: 34
: 10404:M 427 PRO HA :M 426 TYR CD1 : -0.651: 29
: 10404:M 427 PRO HA :M 426 TYR CG : -0.565: 30
: 10404:M 427 PRO 2HD :M 356 ALA O : -0.536: 26
: 10404:M 91 PRO HA :M 90 TYR CD1 : -0.649: 28
: 10404:M 91 PRO HA :M 90 TYR CG : -0.556: 31
: 10404:M 91 PRO 2HD :M 20 ALA O : -0.544: 27
: 10404:M 607 ASP HA :M 613 MET 1HE : -0.646: 53
: 10404:M 613 MET 1HE :M 607 ASP CA : -0.587: 53
: 10404:M 606 LEU 2HD1 :M 603 VAL 1HG1 : -0.543: 31
: 10404:M 613 MET 1HE :M 606 LEU O : -0.467: 53
: 10404:M 606 LEU C :M 613 MET 1HE : -0.447: 53
: 10404:M 572 LYS NZ :M 572 LYS 2HB : -0.643: 46
: 10404:M 572 LYS 2HD :M 583 GLU OE1 : -0.580: 56
: 10404:M 572 LYS 2HB :M 572 LYS 3HZ : -0.490: 46
: 10404:M 572 LYS CD :M 583 GLU OE1 : -0.428: 56
: 10404:M 258 TYR CD1 :M 259 PRO HA : -0.641: 30
: 10404:M 604 PRO 1HG :M 587 GLU OE2 : -0.641: 32
: 10404:M 259 PRO HA :M 258 TYR CG : -0.577: 32
: 10404:M 188 ALA O :M 259 PRO 2HD : -0.568: 22
: 10404:M 594 TYR CD1 :M 595 PRO HA : -0.639: 27
: 10404:M 594 TYR CG :M 595 PRO HA : -0.568: 30
: 10404:M 524 ALA O :M 595 PRO 2HD : -0.525: 20
: 10404:M 404 LYS 2HB :M 404 LYS NZ : -0.634: 39
: 10404:M 404 LYS 2HD :M 415 GLU OE1 : -0.599: 58
: 10404:M 415 GLU OE1 :M 404 LYS CD : -0.434: 58
: 10404:M 404 LYS 2HB :M 404 LYS 2HZ : -0.411: 39
: 10404:M 580 LEU 3HD2 :M 197 VAL 1HG2 : -0.632: 12
: 10404:M 580 LEU O :M 576 ILE HA : -0.596: 22
: 10404:M 552 ASN HA :M 447 ARG NH1 : -0.622: 54
: 10404:M 103 ASP HA :M 109 MET 3HE : -0.620: 51
: 10404:M 109 MET 3HE :M 103 ASP CA : -0.566: 51
: 10404:M 99 VAL 1HG1 :M 102 LEU 2HD1 : -0.553: 28
: 10404:M 109 MET 3HE :M 102 LEU O : -0.487: 51
: 10404:M 102 LEU C :M 109 MET 3HE : -0.457: 51
: 10404:M 277 MET 3HE :M 271 ASP HA : -0.615: 49
: 10404:M 267 VAL 1HG1 :M 270 LEU 2HD1 : -0.565: 31
: 10404:M 271 ASP CA :M 277 MET 3HE : -0.564: 49
: 10404:M 277 MET 3HE :M 270 LEU O : -0.508: 49
: 10404:M 277 MET 3HE :M 270 LEU C : -0.472: 49
: 10404:M 270 LEU 2HD1 :M 267 VAL CG1 : -0.408: 31
: 10404:M 111 ARG NH1 :M 216 ASN HA : -0.614: 56
: 10404:M 58 SER 1HB :M 65 TRP CE3 : -0.612: 29
: 10404:M 58 SER 1HB :M 65 TRP CD2 : -0.486: 29
: 10404:M 236 LYS 2HB :M 236 LYS NZ : -0.611: 38
: 10404:M 247 GLU OE1 :M 236 LYS 2HD : -0.575: 57
: 10404:M 247 GLU OE1 :M 236 LYS CD : -0.436: 57
: 10404:M 236 LYS 2HZ :M 236 LYS 2HB : -0.434: 38
: 10404:M 394 SER 1HB :M 401 TRP CE3 : -0.609: 30
: 10404:M 394 SER 1HB :M 401 TRP CD2 : -0.493: 30
: 10404:M 161 HIS 1HB :M 78 TYR CE1 : -0.608: 33
: 10404:M 78 TYR CZ :M 161 HIS 1HB : -0.500: 33
: 10404:M 562 SER 1HB :M 569 TRP CE3 : -0.607: 23
: 10404:M 226 SER 1HB :M 233 TRP CE3 : -0.607: 24
: 10404:M 562 SER 1HB :M 569 TRP CD2 : -0.505: 23
: 10404:M 226 SER 1HB :M 233 TRP CD2 : -0.499: 24
: 10404:M 72 ILE HA :M 76 LEU O : -0.605: 20
: 10404:M 377 ARG 1HG :M 377 ARG 1HH1 : -0.601: 55
: 10404:M 329 HIS 1HB :M 246 TYR CE1 : -0.601: 39
: 10404:M 329 HIS 1HB :M 246 TYR CZ : -0.485: 39
: 10404:M 377 ARG HA :M 377 ARG NH1 : -0.455: 55
: 10404:M 377 ARG 1HH1 :M 377 ARG CG : -0.406: 55
: 10404:M 497 HIS 1HB :M 414 TYR CE1 : -0.597: 34
: 10404:M 497 HIS 1HB :M 414 TYR CZ : -0.476: 34
: 10404:M 379 VAL 3HG2 :M 375 LEU O : -0.594: 22
: 10404:M 117 LEU 2HD2 :M 117 LEU H : -0.591: 48
: 10404:M 665 HIS 1HB :M 582 TYR CE1 : -0.591: 37
: 10404:M 117 LEU 2HD2 :M 117 LEU N : -0.500: 48
: 10404:M 582 TYR CZ :M 665 HIS 1HB : -0.479: 37
: 10404:M 117 LEU H :M 117 LEU CD2 : -0.418: 48
: 10404:M 28 TRP CD2 :M 22 PRO HA : -0.588: 33
: 10404:M 62 GLY O :M 28 TRP HZ2 : -0.413: 36
: 10404:M 627 PRO 2HD :M 626 LYS 2HB : -0.587: 37
: 10404:M 545 ARG 1HG :M 545 ARG 1HH1 : -0.587: 47
: 10404:M 545 ARG HA :M 545 ARG NH1 : -0.440: 47
: 10404:M 373 GLN O :M 376 ILE 2HG1 : -0.586: 37
: 10404:M 412 LEU O :M 408 ILE HA : -0.586: 27
: 10404:M 458 LYS 2HB :M 459 PRO 2HD : -0.584: 32
: 10404:M 457 PHE O :M 458 LYS C : -0.414: 39
: 10404:M 435 VAL 1HG1 :M 438 LEU 2HD1 : -0.582: 26
: 10404:M 439 ASP HA :M 445 MET 1HE : -0.582: 51
: 10404:M 445 MET 1HE :M 439 ASP CA : -0.549: 51
: 10404:M 445 MET 1HE :M 438 LEU O : -0.534: 51
: 10404:M 445 MET 1HE :M 438 LEU C : -0.489: 51
: 10404:M 438 LEU 2HD1 :M 435 VAL CG1 : -0.426: 26
: 10404:M 40 ILE 2HG1 :M 37 GLN O : -0.581: 40
: 10404:M 240 ILE HA :M 244 LEU O : -0.580: 21
: 10404:M 621 LEU H :M 621 LEU 2HD2 : -0.579: 46
: 10404:M 291 PRO 2HD :M 290 LYS 2HB : -0.579: 34
: 10404:M 621 LEU 2HD2 :M 621 LEU N : -0.496: 46
: 10404:M 621 LEU H :M 621 LEU CD2 : -0.411: 46
: 10404:M 104 GLY 1HA :M 212 GLU OE1 : -0.577: 39
: 10404:M 43 VAL 3HG2 :M 39 LEU O : -0.577: 19
: 10404:M 104 GLY 1HA :M 212 GLU CD : -0.541: 42
: 10404:M 41 ARG 1HG :M 41 ARG 1HH1 : -0.576: 58
: 10404:M 41 ARG NH1 :M 41 ARG HA : -0.455: 58
: 10404:M 41 ARG 1HH1 :M 41 ARG CG : -0.421: 58
: 10404:M 453 LEU 2HD2 :M 453 LEU H : -0.575: 49
: 10404:M 285 LEU H :M 285 LEU 2HD2 : -0.575: 46
: 10404:M 453 LEU 2HD2 :M 453 LEU N : -0.524: 49
: 10404:M 285 LEU N :M 285 LEU 2HD2 : -0.519: 46
: 10404:M 453 LEU H :M 453 LEU CD2 : -0.427: 49
: 10404:M 209 ARG 1HH1 :M 209 ARG 1HG : -0.570: 49
: 10404:M 123 PRO 2HD :M 122 LYS 2HB : -0.570: 34
: 10404:M 209 ARG NH1 :M 209 ARG HA : -0.447: 49
: 10404:M 121 PHE O :M 122 LYS C : -0.411: 40
: 10404:M 190 PRO HA :M 196 TRP CD2 : -0.569: 32
: 10404:M 196 TRP HZ2 :M 230 GLY O : -0.417: 28
: 10404:M 358 PRO HA :M 364 TRP CD2 : -0.567: 30
: 10404:M 548 GLU OE1 :M 440 GLY 1HA : -0.567: 39
: 10404:M 548 GLU CD :M 440 GLY 1HA : -0.531: 36
: 10404:M 398 GLY O :M 364 TRP HZ2 : -0.417: 30
: 10404:M 541 GLN O :M 544 ILE 2HG1 : -0.557: 40
: 10404:M 526 PRO HA :M 532 TRP CD2 : -0.556: 30
: 10404:M 532 TRP HZ2 :M 566 GLY O : -0.420: 27
: 10404:M 205 GLN O :M 208 ILE 2HG1 : -0.553: 40
: 10404:M 207 LEU O :M 211 VAL 3HG2 : -0.542: 23
: 10404:M 543 LEU O :M 547 VAL 3HG2 : -0.542: 27
: 10404:M 115 ILE 2HD1 :M 145 TRP CE2 : -0.541: 30
: 10404:M 115 ILE 3HG2 :M 145 TRP CD1 : -0.511: 37
: 10404:M 110 TYR CD1 :M 116 CYS 2HB : -0.486: 50
: 10404:M 116 CYS N :M 115 ILE 2HG2 : -0.465: 37
: 10404:M 116 CYS N :M 115 ILE CG2 : -0.455: 37
: 10404:M 145 TRP CD2 :M 115 ILE 2HD1 : -0.432: 30
: 10404:M 110 TYR CE1 :M 116 CYS 2HB : -0.431: 51
: 10404:M 145 TRP CZ2 :M 149 GLU 2HG : -0.408: 32
: 10404:M 283 ILE 2HD1 :M 313 TRP CE2 : -0.538: 38
: 10404:M 451 ILE 2HD1 :M 481 TRP CE2 : -0.538: 36
: 10404:M 313 TRP CD1 :M 283 ILE 3HG2 : -0.521: 39
: 10404:M 481 TRP CD1 :M 451 ILE 3HG2 : -0.502: 40
: 10404:M 284 CYS 2HB :M 278 TYR CD1 : -0.483: 49
: 10404:M 451 ILE 2HG2 :M 452 CYS N : -0.476: 40
: 10404:M 446 TYR CD1 :M 452 CYS 2HB : -0.474: 51
: 10404:M 284 CYS N :M 283 ILE 2HG2 : -0.467: 39
: 10404:M 452 CYS N :M 451 ILE CG2 : -0.460: 40
: 10404:M 283 ILE 2HD1 :M 313 TRP CD2 : -0.452: 38
: 10404:M 481 TRP CD2 :M 451 ILE 2HD1 : -0.452: 36
: 10404:M 284 CYS N :M 283 ILE CG2 : -0.447: 39
: 10404:M 284 CYS 2HB :M 278 TYR CE1 : -0.430: 51
: 10404:M 446 TYR CE1 :M 452 CYS 2HB : -0.412: 51
: 10404:M 313 TRP CZ2 :M 317 GLU 2HG : -0.410: 30
: 10404:M 481 TRP CZ2 :M 485 GLU 2HG : -0.409: 32
: 10404:M 187 ASN 2HD2 :M 187 ASN C : -0.537: 34
: 10404:M 355 ASN 2HD2 :M 355 ASN C : -0.536: 34
: 10404:M 466 PRO HA :M 465 VAL HA : -0.424: 29
: 10404:M 355 ASN 1HB :M 466 PRO 2HG : -0.411: 35
: 10404:M 619 ILE 2HD1 :M 649 TRP CE2 : -0.535: 34
: 10404:M 619 ILE 3HG2 :M 649 TRP CD1 : -0.510: 37
: 10404:M 620 CYS 2HB :M 614 TYR CD1 : -0.490: 48
: 10404:M 620 CYS N :M 619 ILE 2HG2 : -0.459: 37
: 10404:M 649 TRP CD2 :M 619 ILE 2HD1 : -0.450: 34
: 10404:M 620 CYS 2HB :M 614 TYR CE1 : -0.444: 51
: 10404:M 620 CYS N :M 619 ILE CG2 : -0.437: 37
: 10404:M 649 TRP CZ2 :M 653 GLU 2HG : -0.410: 33
: 10404:M 557 TRP HD1 :M 555 ASN O : -0.533: 21
: 10404:M 21 GLY C :M 23 ARG H : -0.529: 36
: 10404:M 23 ARG 1HH2 :M 23 ARG 1HG : -0.446: 57
: 10404:M 19 ASN 2HD2 :M 19 ASN C : -0.526: 30
: 10404:M 129 VAL HA :M 130 PRO HA : -0.416: 29
: 10404:M 19 ASN 1HB :M 130 PRO 2HG : -0.401: 35
: 10404:M 523 ASN 2HD2 :M 523 ASN C : -0.525: 29
: 10404:M 525 GLY C :M 527 ARG H : -0.524: 28
: 10404:M 527 ARG 1HG :M 527 ARG 1HH2 : -0.432: 57
: 10404:M 53 TRP HD1 :M 51 ASN O : -0.519: 19
: 10404:M 387 ASN O :M 389 TRP HD1 : -0.510: 22
: 10404:M 114 LYS 1HD :M 96 GLU 2HB : -0.509: 55
: 10404:M 221 TRP HD1 :M 219 ASN O : -0.506: 23
: 10404:M 600 GLU 2HB :M 618 LYS 1HD : -0.503: 54
: 10404:M 432 GLU 2HB :M 450 LYS 1HD : -0.501: 54
: 10404:M 320 ASP OD1 :M 324 LYS 2HE : -0.497: 17
: 10404:M 282 LYS 1HD :M 264 GLU 2HB : -0.492: 56
: 10404:M 656 ASP OD1 :M 660 LYS 2HE : -0.474: 17
: 10404:M 298 PRO HA :M 297 VAL HA : -0.458: 27
: 10404:M 9 VAL O :M 13 ILE 2HG1 : -0.458: 52
: 10404:M 345 VAL O :M 349 ILE 2HG1 : -0.450: 47
: 10404:M 124 LEU HA :M 127 ARG NE : -0.441: 47
: 10404:M 181 ILE 2HG1 :M 177 VAL O : -0.437: 44
: 10404:M 463 ARG NE :M 460 LEU HA : -0.436: 50
: 10404:M 513 VAL O :M 517 ILE 2HG1 : -0.432: 50
: 10404:M 295 ARG NE :M 292 LEU HA : -0.428: 44
: 10404:M 628 LEU HA :M 631 ARG NE : -0.426: 44
: 10404:M 598 ALA 2HB :M 629 TRP CE2 : -0.415: 31
: 10404:M 293 TRP CE2 :M 262 ALA 2HB : -0.413: 26
: 10404:M 94 ALA 2HB :M 125 TRP CE2 : -0.409: 30
: 10404:M 220 ASP 1HB :M 215 LYS 1HG : -0.409: 20
: 10404:M 430 ALA 2HB :M 461 TRP CE2 : -0.408: 29
: 10404:M 518 PRO 1HG :M 542 SER HA : -0.405: 23
: 10404:M 556 ASP 1HB :M 551 LYS 1HG : -0.405: 23
: 10404:M 14 PRO 1HG :M 38 SER HA : -0.401: 20
#sum2 ::33.35 clashscore : 24.75 clashscore B<40
#summary::10404 atoms:8485 atoms B<40:1157503 potential dots:72340.0 A^2:347 bumps:210 bumps B<40:2417 score
Output from PDB validation software
Summary from PDB validation
May. 10, 07:51:48 2013
Greetings,
[ Text modified to reflect that this was run under PSVS - Aneerban Bhattacharya: Dec 2005 ]
The following checks were made on :
-----------------------------------------
DISTANCES AND ANGLES
We have checked your intra and intermolecular distances and angles with the
procedures currently in place at PDB:
==> The following solvent molecules are further away than 3.5 Angstroms from
macromolecule atoms which are available for hydrogen bonding in the
asymmetric unit.
none
The coordinates for water molecules which could be translated back into
the asymmetric unit are listed. If you do not indicate otherwise we will
replace the solvent coordinates in the entry with the ones below:
none
==> Close contacts in same asymmetric unit. Distances smaller than 2.2
Angstroms are considered as close contacts.
none
==> Close contacts based on crystal symmetry. Distances smaller than 2.2
Angstroms are considered as close contacts.
none
==> Bond and angle checks are performed by first computing the average rms
error for all bonds and angles relative to standard values for nucleotide
units [L. Clowney et al., Geometric Parameters in Nucleic Acids: Nitrogenous
Bases, J.Am.Chem.Soc. 1996, 118, 509-518; A. Gelbin et al., Geometric
Parameters in Nucleic Acids: Sugar and Phosphate Constituents, J.Am.Chem.Soc.
1996, 118, 519-529] and amino acid units [R.A. Engh and R. Huber, Accurate
Bond and Angle Parameters for X-ray protein structure refinement, Acta
Crystallogr. 1991, A47, 392-400]. Any bond or angle which deviates from the
dictionary values by more than six times this computed rms error is
identified as an outlier.
*** Covalent Bond Lengths:
The RMS deviation for covalent bonds relative to the standard
dictionary is 0.008 Angstroms
The following table contains a list of the covalent bonds
greater than 6.0*RMSD.
Deviation Residue Chain Sequence AT1 - AT2 Bond Dictionary
Name ID Number Distance Value
------------------------------------------------------------------------
0.054 VAL A 99 CA - CB 1.594 1.540
0.057 VAL B 99 CA - CB 1.597 1.540
0.051 VAL C 99 CA - CB 1.591 1.540
0.055 VAL D 99 CA - CB 1.595 1.540
*** Covalent Angle Values:
The RMS deviation for covalent angles relative to the standard
dictionary is 1.2 degrees.
The following table contains a list of the covalent bond angles
greater than 6.0*RMSD.
Deviation Residue Chain Sequence AT1 - AT2 - AT3 Bond Dictionary
Name ID Number Angle Value
--------------------------------------------------------------------------------
7.4 PRO A 91 C - N - CA 130.0 122.6
7.9 GLU A 149 N - CA - C 119.1 111.2
7.5 PRO B 91 C - N - CA 130.1 122.6
7.6 GLU B 149 N - CA - C 118.8 111.2
7.5 GLU C 149 N - CA - C 118.7 111.2
7.6 PRO D 91 C - N - CA 130.2 122.6
8.0 GLU D 149 N - CA - C 119.2 111.2
TORSION ANGLES
The torsion angle distributions have been checked. The postscript file of the
conformation rings showing the torsion angle distributions will be sent in a
separate E-mail message.
CHIRALITY
The chirality has been checked and there are no incorrect carbon chiral centers.
Some of O1P and O2P atoms do not follow the convention defined in the standard
IUBMB nomenclature (Liebecq, C. Compendium of Biochemical Nomenclature and Related
Documents, 2nd ed.; Portland Press: London and Chapel Hill, 1992). If you do not
indicate otherwise, we will switch the labels of O1P and O2P as shown below.
OTHER IMPORTANT ISSUES
==> Please check carefully REMARKS 3 and 200 and fill in the parameters as
appropriate.
















































































